BIOL368/F14:Nicole Anguiano Week 10
From OpenWetWare
Jump to navigationJump to search
HIV Structure Redux
- To get more information on the individual subjects and clones studied, see Week 9. The presentation this information is coming from can be found here.
- Note: Upon further investigation, it appears that I studied the actual beta sheet proteins instead of the protein in the beta sheet listed by PsiPred. As a result, instead of the information coming from the sequence "FYT", as PsiPred called for, I found the information for "RAF", which is the actual beta sheet in the V3 tip.
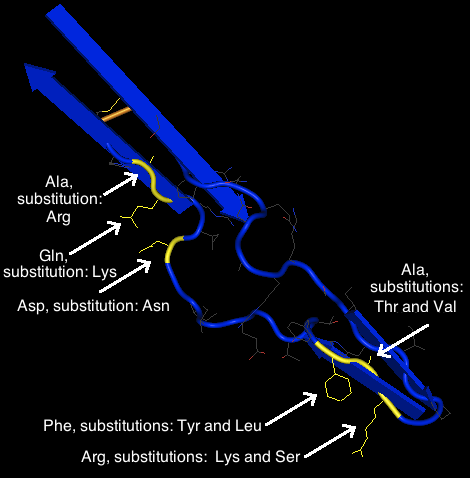
- It is likely that the majority of the amino acids that switch out may not be necessarily relevant to the function of the gp120 protein. Due to the fact that all of the substitutions existed in viable clones, all of the substitutions create a functional protein that is capable of infection. Beginning on the right hand side at the bottom near the V3 tip, the substitution of Arginine for Lysine would likely had no effect. However, the substitution of Arginine for Serine would likely have an effect due to the dramatic change in size. The substitution of the Alanine for Threonine or Valine would likely cause some change in function, as Alanine is a very small molecule and Threonine and Valine are much larger. Phenylalanine, Tyrosine, and Leucine are all large, bulky molecules. The substitutions of Phenylalanine for Tyrosine would likely cause no effect, as they are both benzene rings. A substitution with Leucine would also likely have no effect due to its large size and lack of polarity, both traits that it shared with phenylalanine. The substitution of Aspartic Acid for Asparagine, due to its location in the already disordered random coil of the returning strand, likely would have little no effect. The substitution of Glutamine with Lysine, due to the large size, would also likely have little effect. Lastly, the substitution of alanine with arginine would likely have a dramatic effect. Alanine is a small, non-polar molecule and Arginine is an extremely large, basic molecule. As a result, it is highly likely that a change in function would occur as a result of this substitution.
Introduction to DNA Microarrays
- Questions are adapted from Campbell, A.M. and Heyer, L.J. (2003), “Chapter 4: Basic Research with DNA Microarrays”, in Discovering Genomics, Proteomics, and Bioinformatics, Cold Spring Harbor Laboratory Press, pp. 107-124.
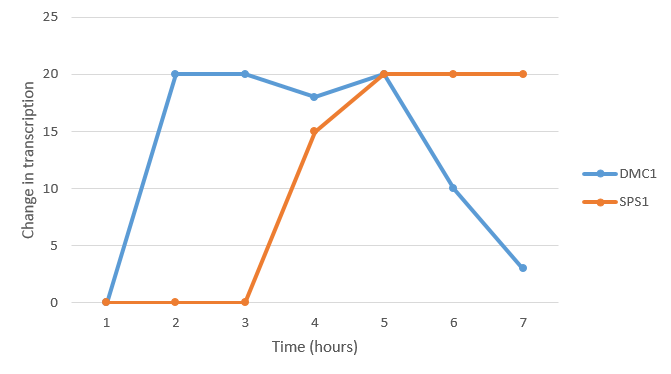
- (Question 5, p. 110) Choose two genes from Figure 4.6b (PDF of figures on MyLMUConnect) and draw a graph to represent the change in transcription over time.
- (Question 6b, p. 110) Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6, determine the color for each ratio in Figure 4.7b. (Use the nomenclature "bright green", "medium green", "dim green", "black", "dim red", "medium red", or "bright red" for your answers.)
-
1 Hour 3 Hour 5 Hour 9 Hour gene X 1.0: Black 2.2: Dim red 1.0: Black 0.15: Medium Green gene Y 1.0: Black 4.5: Dim red 0.95: Black 0.05: Bright Green gene Z 1.0: Black 1.5: Dim red 2.0: Dim red 2.0: Dim red
-
- (Question 7, p. 110) Were any of the genes in Figure 4.7b transcribed similarly? If so, which ones were transcribed similarly to which ones?
- Gene X and gene Y transcribed similarly. Both genes underwent a small amount of activation at the third hour, reached equilibrium at hour 5, then underwent a significant amount of repression at the 9th hour. Gene Z had a relatively small amount of activation throughout the entirety of the cycle.
- (Question 9, p. 118) Why would most spots be yellow at the first time point? I.e., what is the technical reason that spots show up as yellow - where does the yellow color come from? And, what would be the biological reason that the experiment resulted in most spots being yellow?
- Most spots would be yellow at the first time point because at the first time point, there has not been enough elapsed time for a activating or repressing effect to take place. If a gene is repressed by another gene, the other gene has to replicate enough to be able to have a visible effect, which would likely not be possible at the first time point. The yellow color comes from the combination of the red and green labeled cDNA. As neither has had the chance to dominate, the combination of the two will result in a yellow color. The biological reason that the experiment resulted in most spots being yellow is that most spots are neither activated nor repressed by other genes. Most of the DNA sequences represented at those points operate independently from the other represented genes. These spots likely code for genes that are not used in anything relating to the metabolism and/or function of glucose.
- (Question 10, p. 118) Go to the Saccharomyces Genome Database and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s change in expression was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
- Over the course of the experiment, TEF4 was repressed. It is possible that reduction in levels of glucose causes the amount of translation to be reduced. Glucose may perform an important role in the cell performing translation, so to remove it from the environment may cause an important piece of the pathway to be removed, thus preventing it from continuing. This would lead to the repression of the genes in the rest of the pathway.
- (Question, 11, p. 120) Why would TCA cycle genes be induced if the glucose supply is running out?
- The TCA cycle genes would be induced if the glucose supply is running out to ensure that the remaining glucose can be adequately metabolized without being turned into acetyaldehyde. The metabolized glucose can then be stored for use during the period in which there is no glucose supply available.
- (Question 12, p. 120) What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
- The genome could have all of the genes for a pathway located one after the other so they are all translated at the same time. If the area in which they are located is induced, all of the genes in the row will be induced as well. However, if the area is repressed, then all of the genes are repressed.
- # (Question 13, p. 121) Consider a microarray experiment where cells deleted for the repressor TUP1 were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented glucose-repressed genes to be in the later time points of this experiment?
- It would be expected that the spots that represented glucose-repressed genes would be red. As the gene responsible for triggering the repression has been deleted, the spots that represent glucose-repressed genes will continue expressing at the same rate. The inducers that would act on the genes, without the repressor to block them, may even increase the rate of gene expression, causing them to express at an accelerated rate.
- (Question 14, p. 121) Consider a microarray experiment where cells that overexpress the transcription factor Yap1p were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented Yap1p target genes to be in the later time points of this experiment?
- As in the case of the experiment in which TUP1 was deleted, it is likely that in the case that Yap1p is overexpressed, the majority of the spots that represent Yap1p target genes will be red. Yap1p confers resistance to environmental stress. The genes that it activates in order to do this will be heavily activated if Yap1p is mutated in order to be overexpressed.
- (Question 15, p. 121) Could the loss of a repressor or the overexpression of a transcription factor result in the repression of a particular gene?
- Theoretically, yes. However, it is unlikely. If the repressor of a gene is lost, then that gene will likely no longer be repressed, and if a transcription factor that induces the gene is overexpressed, then it will likely only increase in expression as a result.
- (Question 16, p. 121) Using the microarray data, how could you verify that you had truly deleted TUP1 or overexpressed YAP1 in the experiments described in questions 8 and 9?
- In the case of the deletion of TUP1, the location coding for TUP1 should be black, as the gene doesn't exist in the genome. If there is a spot at the location, it is likely that the gene has not been deleted, but if there is no spot then that provides some evidence that it has been deleted. In the case of the overexpression of YAP1, the spot where YAP1 is located should be very bright red, and may be larger than the rest of the spots due to being over expressed. If the spot is not bright red, then it is likely not overexpressed.
Finding a Journal Club Article
Links
Nicole Anguiano
BIOL 368, Fall 2014
Assignment Links
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 13 Assignment
- Week 15 Assignment
Individual Journals
- Individual Journal Week 2
- Individual Journal Week 3
- Individual Journal Week 4
- Individual Journal Week 5
- Individual Journal Week 6
- Individual Journal Week 7
- Individual Journal Week 8
- Individual Journal Week 9
- Individual Journal Week 10
- Individual Journal Week 11
- Individual Journal Week 12
- Individual Journal Week 13
- Individual Journal Week 15