20.109(F15):Homework
Module 1: DNA engineering
Due M1D1
- Review the lab orientation exercises to prepare for the lab practical that you and your partner will take together next time.
- Complete the required EHS Training on-line.
- There are two web-based training modules required for 20.109. They are Chemical Hygiene Training and Managing Hazardous Waste. Chemical Hygiene includes 7 sections and 6 quizzes with an estimated completion time of 1 hour, while Managing Hazardous Waste has one quiz and should take somewhat less time to complete. Both courses can be accessed through MIT's Environmental Health and Safety page, from any computer that has your MIT certificate on it.
- If you have completed EHS training in a UROP or in another lab class, you do not need to repeat the training but you do need to print out your training record to hand in.
- From the EHS training page select the second button labeled “I have EHS training requirements for an academic subject.”
- Your summary page (“My EHS Training") should show Chemical Hygiene and Managing Hazardous Waste as requirements for 20.109. Click on the purple button “Go to Web Classes” right above the training requirements section. You may stop and start the web-based courses as many times as you need to complete them; the software keeps track of your progress in the course.
- Print the certificates of completion (or your training record) to turn in next time.
- Open an Evernote account at this link. We recommend downloading Evernote to your computer versus operating from the web based application (although both work). You can also download Evernote to your smart phone for ease of uploading pictures and data. You should make a notebook that will be utilized as your lab notebook. Give it a name with the following convention: 20.109(F15)_Name and share that notebook with your lab instructor and Module 1 TA.
- Prepare for the first day of Module 1 by reading the module overview and day one introduction.
Due M1D3
- Sketch the expected product from the PCR you performed today.
- You may work on paper or electronically. Either way, prepare a schematic rather than detailing each base.
- Clearly indicate the 5' and 3' end of each DNA strand.
- Be sure to reflect every new feature that you have introduced (e.g., restriction site) or deleted.
- What is the expected size of the PCR product?
- Following the directions in Part 4 of M1D2, prepare a plasmid map in ApE of the clone you are trying to create in lab. Print the graphic map with all singly present restriction sites shown.
- Hint: You may choose to show fewer restriction sites in your Module 1 summary.
- Using your map, calculate the fragment sizes expected for each double digest below. Please show your work.
- EcoRV and XbaI
- BamHI and XhoI
- Note: You should complete problem #2 after the M1D2 lab.
- In Module 2, you will document your experiments in a written methods section that will be part of a larger report. To help you prepare, as well as give you feedback early on, you will draft portions of the Module 1 methods. For this assignment, write a draft of the Methods: on PCR and DNA digestion. Be sure to read the Materials and Methods section guidelines at this link before you begin; doing so may save you some effort.
Due M1D4
- You will document the work you do during Module 1 in a DNA engineering summary. To help you pace your work, as well as give you feedback early on, you will draft small portions of the summary as homework assignments. For M1D4, you should document your experimental outcomes.
- Begin by reading, under Logistics, Guidelines on Formatting and Length. Next, skim the Content Guidelines, paying particular attention to those relevant for M1D3.
- You should present both the purification gel and the recovery gel, each in a well-labeled figure with an appropriate figure caption. Be sure to follow our written guidelines as well as the suggestions presented by your writing instructor during class.
- Below the figures, interpret the outcomes in concise language. Consider reviewing slides 30 and 31 in the Writing Lab talk titled EBICS REU Poster Best Practices DMC SWO 1 to learn how to turn bulky sentences into succinct bullets. It's also okay to use complete sentences when they more clearly make your point.
- Please see this example for a suggested layout.
- To prepare for the lab session, calculate the amounts of DNA you will use in the ligation by following the steps below.
- Separately calculate the concentration of backbone and of insert based on the recovery gel posted on today's Talk page. Refer to the NEB marker definitions to estimate the ng of DNA in each lane, and refer to your notebook/protocol for the relevant volume basis. Note that the ng listed are for 10 μL of ladder.
- You may convert the mass concentration to a molar concentration, using the fact that a typical DNA base is 500 g/mol. This conversion will mostly cancel out between the insert and the backbone, except for the difference in number of bases. Feel free to either omit steps that will cancel if you are comfortable doing so, or to keep them if you follow the lab math better that way.
- Ideally, you will use 50-100 ng of backbone in the upcoming ligation. Referring to the mass concentration, what volume of DNA will this amount require?
- Ideally, you will use a 4:1 molar ratio of insert to backbone. Referring to the molar concentrations, how much insert do you need per μL of backbone?
- A 15 μL scale ligation should not include more than 13.5 μL of DNA. If your backbone and insert volumes total to greater than this amount, you must (1) scale down both DNA amounts, using less than 50 ng backbone and/or (2) stray from the ideal 4:1 molar ratio. You may ask the teaching faculty for advice during class if you are unsure what choice is best, but make and submit an initial ligation plan for now.
Due M1D5
- In addition to reporting your results in the DNA engineering summary, you will need to introduce your project. For this assignment, write topic sentences for the paragraphs that will be included in the background and motivation section of your data summary for module 1. Your topic sentences should include references that you find to validate and support the statement. At this point the references can be in any format you choose. Please include between 3 and 7 topic sentences.
- Review the general guidelines for writing up an introduction to your research.
- In addition to the topic sentences, submit a list of your references. You should include the title of the referenced article and a brief summary of the article that includes why you chose that reference to support your topic sentence.
- Prepare a schematic diagram that describes your study of homologous recombination -- the diagram should include information about the construction of your plasmid system and (at this point) a rough idea of how you will use the system to study DNA damage. See the schematic overview on the Module 1 homepage, but make sure that your diagram is presents your own thoughts and ideas (i.e. do not just copy that diagram!). Note: schematic diagrams require figure captions.
Due M1D6
- Revise your earlier draft of the Methods section, just through M1D2, applying the feedback you received.
- Prepare the rest of your Methods section (through M1D5) in outline form. Start by considering what methods may be logically grouped together. Please submit:
- sub-section titles
- topic sentences for each sub-section
- bulleted lists of the individual methods that will be included in each section
- Prepare the rest of your Methods section (through M1D5) in outline form. Start by considering what methods may be logically grouped together. Please submit:
- Review the tissue culture guidelines in preparation for M1D6, when you will set up the culminating experiment of this module!
- Lastly, consider the following questions concerning the transformation experiment:
- Imagine that your EGFP positive control turns up 200 colonies. What is the transformation efficiency under these conditions, in colony-forming units (CFU) per μg of DNA? Be sure to carefully consider how much DNA ended up on the plate.
- If you found an equal number of colonies on your "bkb + ligase" plate and your "bkb + insert + ligase" plate, what DNA would you predominantly expect to isolate from the "bkb + insert + ligase" colonies, plasmid with insert or plasmid without insert? In 2-3 sentences, explain how plasmid without insert could end up being recovered here -- which experimental step(s) failed, and how?
- Note: These questions are not to be turned in but they should help prepare you to interpret the results you will collect and to analyze and frame your data for the DNA engineering summary.
Due M1D7
- In addition to the DNA engineering summary, each student will individually complete a mini-presentation that is focused on the data generated in Module 1. The mini-presentation (or 'elevator pitch') is your opportunity to practice your oral communication skills before the Journal Club presentation in Module 2. For this assignment, prepare an outline for your mini-presentation. Be sure to include the following:
- topic sentence that introduces the big picture
- key results
- take-home message
Wrapping up M1
- The DNA engineering summary draft is due by 5 pm on Tuesday, October 13 for both sections.
- The DNA engineering mini-presentation is due by 5 pm on Saturday, October 17 for both sections.
- The DNA engineering summary revision is due by 5 pm on Saturday, October 24 for both sections.
All assignments are to be submitted on Stellar.
- Don't forget to contribute your thoughts, comments and ideas to the 20.109 class blog, remember that you are required to post within 24 hr of submitting your DNA engineering summary revision.
Module 2: Protein engineering
Due M2D1
- Prepare for the first day of Module 2 by reading the module overview and day one introduction.
Due M2D2
- Toward the end of class, we will discuss an article from the primary literature. Specifically, we will discuss the construction and analysis of the inverse pericam (IPC) multi-component calcium sensor. To prepare for this discussion, you should do a close reading of the paper by Nagai et al that introduced IPC. For context, you are also encouraged to read the short paper by Heim, Prasher, and Tsien, in which the very first attempt to mutagenize GFP is described, and which is a fine introduction to some of the concepts and methods used in this module. Though you should read the entire paper, be prepared to specifically discuss the section assigned to your group on the M2D2 page.
- Create a diagram that depicts the aspects of your plasmid, which contains your mutated inverse pericam. Be sure to highlight the following:
- inverse pericam protein coding sequence
- primer site
- location of both mutations
- location of the introduced restriction enzyme recognition sequence
- Note: Use the diagrams below as a guide.
- The following assignment will prepare you to set up your digestion reactions. On the whole, Day 2 will likely run long, so reading ahead and completing as many calculations as possible in advance will be of help in making the day run smoothly.
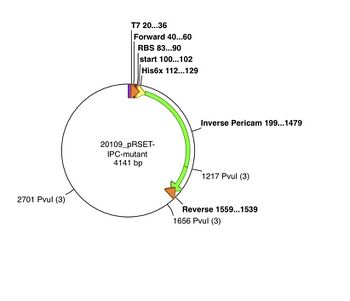
The pRSET plasmid with inverse pericam insert, or pRSET-IPC, is 4141 basepairs long and its GenBank file is linked here. According to the cutters list, restriction site PvuI occurs at ~1655 bp, and again at ~2700 bp into pRSET-IPC. Thus, digesting this parental plasmid with the PvuI enzyme should result in two linear fragments of DNA, with about 1045 and 3095 bp sizes.
As you learned in lab, a silent mutation can be introduced that results in a new restriction enzyme site. For example, let's consider what would happen if we used mutation primers to introduce a new PvuI site at the 341st-342nd residues of inverse pericam (ATT → ATC and TAC → GAC) , or approximately the 1020th basepair of IPC. When IPC is inserted into pRSET, its starting point is ~200 bp into the pRSET plasmid. Thus, if the mutated pRSET-IPC plasmid is digested with PvuI, three linear fragments of DNA are the result: 440, 1045, and 2655 bp. To understand these calculations, see also the plasmid maps above. Make sure you can reproduce the numbers in this example before proceeding with your own samples. Plasmid maps can be made using the Enzyme Selector and Graphic Map functions in ApE.
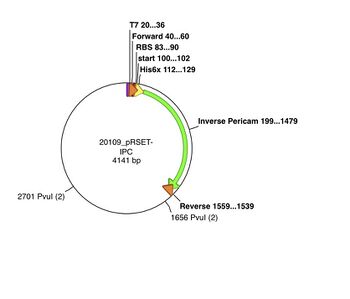

For this assignment, you should prepare for the restriction enzyme digests that allow you to distinguish parental and mutant pRSET-IPC for your X#Z mutation.
- Use the NEB site to determine the appropriate buffer and temperature for your reaction (this will be dependent on the restriction site introduced by the mutation primers you selected). Searching for your enzyme by name is probably easiest.
- Explicitly show how the digest distinguishes between parent IPC and mutant IPC. In other words, what are the expected band sizes for each plasmid upon digestion?
- Calculate the amount of enzyme and buffer needed for your digestion master mixes. Refer to diagnostic digest protocol on M2D2.
Due M2D3
- The major assessment for this module will be a research report describing your protein design work. For this assignment, you will draft the introduction of your report. Recall that the introduction provides a framework for the story you are about to tell, with respect to both background and motivation. You are encouraged to revisit the scientific writing guidelines before you begin.
- Consider the following questions before you start:
- Of what might the big picture section consist? Will you focus on engineering sensors in general, the utility of sensing calcium specifically, or ... what?
- What background information will the reader need to understand your project?
- What is your hypothesis?
- The first paragraph of your introduction should be written in complete sentences (don't forget your references). The remaining paragraphs can be in outline form, but must include topic sentences and a bulleted list of the supporting information that will be used to craft your paragraphs (again, don't forget your references).
- Note: Though you do not have your data yet, you should indicate where this information will be included.
- Consider the following questions before you start:
- Prepare a schematic diagram that depicts your mutagenesis strategy and write a short caption for it. You might show proposed changes at both the nucleotide and amino acid level for your X#Z mutation.
Due M2D4
- To prepare for Day 4, which has the potential to run long, you should fill in the table under "Advance preparation for PAGE" based on the OD values you measured. Be sure to post these to the Day 3 Talk page.
Due M2D6
- Calculate the approximate protein concentrations for your inverse pericams. First make a standard curve from the BSA data, then perform a linear fit (for example, using the Add Trendline function in Excel). The chart does not have to be especially pretty as it will not go in your report, but please do show your work, starting from the raw data. While for now mg/mL is an acceptable concentration unit, note that for your report you will want to convert to mM, μM, or nM - whichever seems most appropriate.
- At this point, you have a lot of practice crafting figures and captions. One change between Module 1 and Module 2 is that your associated results text will be completely in narrative form, rather than partially in bullet points. For M2D4, prepare a figure and caption depicting your digest gel results, as well as the associated results narrative. Here are some things to keep in mind when drafting your Results section:
- Each sub-section should begin with an overview sentence that motivates and introduces the experiment. What did you do and why did you do it?
- State the results of the experiment, minimizing any interpretation of the data (save that for the discussion!)
- Concluding sentences for each paragraph will transition to the next piece of data when possible -- stick to one topic per paragraph, but each sub-section might have a few paragraphs each.
- In Module 1, we let you omit a methods section from your DNA engineering summary, but this blissful state of affairs cannot last forever! For this assignment, you will take a first stab at writing up the methods for your Protein engineering report. This early draft will include just the content/procedures from Days 1-4 of lab.
- Be sure to read the Methods section guidelines before you begin; doing so may save you some effort.
- As you compose your methods for the first 4 days (site-directed mutagenesis, preparation of expression system, induction of protein expression, and purification of protein), do your best to think ahead about the scope of the experiment and how the individual laboratory days fit into that overall context.
- To maximize the feedback you receive it is important that you (a) complete each homework and (b) put forth your best effort on these assignments.
Due M2D7
- Peer review of methods...
Due M2D8
- Post results to D7 talk page...
Wrapping up M2
- The Protein engineering report is due by 5 pm on Saturday, November 14 for both sections.
Assignment is to be submitted on Stellar.
- Don't forget to contribute your thoughts, comments and ideas to the 20.109 class blog, remember that you are required to post within 24 hr of delivering your Journal Club presentation and within 24 hr of submitting your Protein engineering report.
Module 3: Biomaterials engineering
Due M3D1
- Prepare for the first day of Module 3 by reading the module overview and Day 1 introduction.
Due M3D2
- The primary assignment for this experimental module will be for you to develop a research proposal and present your idea to the class. For next time, please describe five recent findings that could potentially define an interesting research question. You should hand in a 3-5 sentence description of each topic, in your own words, and also formally cite an associated reference from the scientific literature. The topics you pick can be related to any aspect of the class, i.e. DNA, system, or biomaterials engineering. During lab next time, you and your partner will review the topics and narrow your choices, identifying one or perhaps two topics for further research.
- Please note: for this assignment, you do not need to have a novel research idea completely sketched out; you simply have to describe five recent examples of existing work that you find interesting. However, you can start to brainstorm how to build off of those topics into something new if you want to get ahead of the game.
Due M3D3
- Discuss the potential research topics you prepared for the previous assignment with your co-investigator (your lab partner) and write a paragraph concerning the research question you would like to pursue for your research proposal. Please include 2-3 sentences that introduce your topic and a brief discussion of your potential plan. Consider the following as you discuss your potential research topics:
- Your interest in the topic.
- The availability of good background information.
- Your likelihood of successfully advancing current understanding.
- The possibility of advancing foundational technologies or finding practical applications.
- Can your proposal be carried out in a reasonable amount of time and with non-infinite resources?
- Take advantage of downtime in lecture and lab to discuss your research ideas with Angie and the teaching faculty.
- Please note: The idea you submit for this assignment does not have to be the idea you present at the end of Module 3. It is okay if you change directions and decide to pursue other research questions during the process of developing your proposal.
Due M3D4
- Read the introductory material for the battery testing lab, and then, based on the mass of the active electrode material you've made, determine the number of mA to apply in order to fully discharge the battery in 10 hours. Use the loading factor of 51 mA/g, a value that is in accordance with previously tested gold nanowire electrodes, and assume 85% of the mass you measured for the nanowires is active material in the anode.
- Consider the feedback you and your co-investigator received in the peer review exercise and begin to refine your research proposal by making a wiki page to collect your ideas and resources (you can do this on one page with your partner or split the effort and each turn in an individual page). Keep in mind that your presentation to the class will ultimately need:
- a brief project overview
- sufficient background information for everyone to understand your proposal
- a statement of the research problem and goals
- project details and methods
- predicted outcomes if everything goes according to plan and if nothing does
- needed resources to complete the work
You can organize your wiki page along these lines or however you feel is most helpful -- check out the “yeast rebuild” or the “T7.2” wiki pages on OpenWetWare for examples of research ideas in process. For this assignment, please submit a printed copy of your research page, making sure it defines your general topic (background and significance), your specific idea (research gap and general approach), and two or more references you've collected and summarized. Keep in mind that your idea may still change - if you come up with something that you like better later on, that's fine.
Please note: you may also utilize any 'cloud' software (Evernote, Google docs), but it must be something that is accessible to both you and your partner.
Due (after) M3D5
- The Biomaterials engineering mini-report is due by 10 pm on Thursday, December 3 or Friday, December 4, according to which day you have lab.
Wrapping up M3
- The Research proposal presenation slides are due by 1 pm on Tuesday, December 8 or Wednesday, December 9, according to which day you have lab.
Assignment is to be submitted on Stellar.
- Don't forget to contribute your thoughts, comments and ideas to the 20.109 class blog, remember that you are required to post within 24 hr of delivering your Research proposal presentation. In addition, the three optional extra credit blog posts are due by 5 pm on Saturday, December 12.