20.109(F15):Evaluate mutations and site-directed mutagenesis (Day1)
Introduction
The process of scientific inquiry encompasses much more than the collection and interpretation of data. A key part of the process is design – specifically of experiments that address a hypothesis and of new materials or technologies. Moreover, any design is subject to continued revision. You might redesign an experiment or tool based on your own research, or you might consult the vast body of scientific literature for other perspectives. As the old graduate student saying (sarcastically) goes, “A month in the lab might save you a day in the library!” In other words, although the process of combining the literature can be arduous or even tedious at times, it beats wasting a month of your time repeating experiments already proven not to work or reinventing the wheel.
During this module, you will generate and test a new version of inverse pericam (IPC). Today, we will refer to a few primary research articles to familiarize ourselves with this recombinant protein and its constituent parts. The fluorescent component of IPC is an enhanced yellow fluorescent protein (abbreviated EYFP), one of the many derivatives of green fluorescent protein (GFP). GFP is naturally produced by jellyfish and was cloned into other organisms in the early 1990’s. It has since been exploited as a genetically encodable reporter and mutagenized to vary its excitation and emission spectra. The other key component of inverse pericam is the protein calmodulin (CaM), a natural calcium sensor that is present in all eukaryotes (and briefly reviewed here). Calmodulin has many ligands that it binds only in the presence of calcium ion, including the peptide fragment M13. This conditional specificity for M13 binding is enabled by the change in CaM’s conformation when it binds calcium.
Within inverse pericam, M13 and CaM are located at opposite ends, surrounding a permuted (i.e., rearranged) version of EYFP. In the absence of calcium, this EYFP exhibits strong fluorescence. However, when enough calcium is added to a solution of inverse pericam, CaM and M13 interact, disrupting the conformation and, as a result, the fluorescence of EYFP. The transition from bright to dim fluorescence occurs over a particular concentration range of calcium. Your goal today is to propose a mutation that will shift the concentration range over which IPC fluorescence decreases. Specifically, you will modify the calcium sensor portion of inverse pericam in a manner that is likely to increase or decrease its affinity for calcium ion.
To make reasonable modifications to inverse pericam, we will use several protein analysis tools. Proteins are modular materials that may be described and examined at multiple levels of a structural hierarchy (from primary to quaternary in the classical paradigm). Primary structure refers to a protein’s amino acid sequence, which might reveal a cluster of charged residues, say, or a pattern of alternating polar and nonpolar residues. One cannot predict off-hand the conformation of a protein merely from its linear sequence; however, due to rotational flexibility of bonds and non-covalent interactions between non-adjacent amino acids (as well as covalent disulfide bonds) some structural characteristics can be inferred.
Physical methods used to interrogate 3D protein structure include X-ray diffraction (XRD), electron microscopy (EM), and nuclear magnetic resonance (NMR) spectroscopy. The paper by Zhang et al. that you will refer to today describes the decoding of calmodulin’s structure using NMR, which depends on subjecting molecules to electromagnetic fields and analyzing the resulting energy absorption spectra of their nuclei. Scientists who elucidate protein structures, in addition to publishing their results, will often add them to public databases such as the Protein Data Bank (PDB). Because many proteins have structural motifs in common (e.g., alpha helices and beta sheets at the secondary level, or leucine-rich repeats at the tertiary level), which ultimately arise from their amino acid sequences, such databases can be useful for making predictions about proteins with known amino acid sequences but unknown structures. Today we will use a computer program that harnesses information in the Protein Data Bank to display interactive 3D models.

After examining both two- and three-dimensional protein information, you will select primers that incorporate a mutation to the wild-type inverse pericam protein and use site-directed mutagenesis to incorporate the corresponding base pair change. The site-directed mutagenesis (SDM) strategy you will use shares some features with the polymerase chain reaction (PCR) for DNA amplification. Recall from Module 1 that PCR amplification involves multiple cycles of melting, annealing, and extending. To create one or more base-pair mutations in the product DNA, primers that have a slight mismatch to the original template can be used. At a low enough annealing temperature (~25 °C below the primer melting temperature as defined for mutagenesis), these nearly-complementary primers will still anneal to the template DNA, but the copies created during the extension phase will contain the mutation.
You will combine the mutagenic primers of your choice with plasmid DNA encoding wild-type inverse pericam. These will be acted upon by a DNA polymerase to generate a plasmid that carries the inverse pericam gene. Even more copies of the mutant plasmid can be made by introducing it into bacteria in a process called transformation, which you are familiar with from Module 1. Remember that there is still parental -- that is, non-mutant -- DNA present in your SDM reaction mixture. In order to propagate only the mutant plasmid upon introduction into bacteria, the parental DNA is selectively digested using the DpnI enzyme prior to bacterial transformation. The underlying selective property is that DpnI only digests methylated DNA. Thus, the synthetically made (and thus non-methylated) mutant DNA is not digested, while the parental DNA is digested due to methylation by the host bacterial strain originally used to amplify it. The resulting small linear pieces of parental DNA are simply degraded by the bacteria, whereas the largely intact (but nicked) mutant DNA is actually repaired by these very same bacteria.
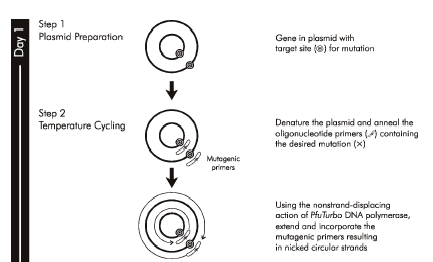
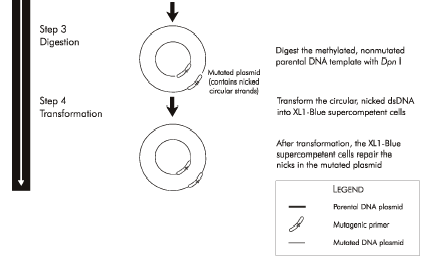
Now might be a good time to mention why we care about measuring intracellular calcium in the first place. Calcium is involved in many signal transduction cascades, which regulate everything from immune cell activation to muscle contraction, from adhesion to apoptosis - see for example this review by David Clapham in Cell, or this one by Ernesto Carafoli in PNAS. Intracellular calcium (Ca2+) is normally maintained at ~100 nM, orders of magnitude less than the ~mM concentration outside the cell. ATPase pumps act to keep the basal concentration of cytoplasmic calcium low. Often calcium acts as a secondary messenger, i.e., it relays a message from the cell surface to its cytoplasm. For example, a particular ligand may bind a cell surface receptor, causing a flood of calcium ions to be released from the intracellular compartments in which they are usually sequestered. These free ions in turn may promote phosphorylation or other downstream signaling.
The proteins that bind calcium do so with a great variety of affinities, and have roles ranging from sequestration to sensing. Some calcium responses may have long-term effects, particularly in the case of transcription factors that can bind calcium. As discussed in lecture, calmodulin works as a calcium sensor by undergoing a conformational change upon calcium binding. Your goal today is to prepare mutant calmodulin (in the context of inverse pericam) DNA, in order to alter the affinity of the resulting protein for calcium.
After today's lab session, the teaching faculty will transform your mutated plasmids into cells that are able to generate multiple copies. When you return you will receive your purified (and hopefully mutated) plasmid. The details will be discussed further during prelab.
Protocols
Part 1: Protein backbone
Perhaps nothing is so conducive to a feeling of intimate familiarity with a protein as studying it at the amino acid level (primary structure). For the first part of lab today, you will examine a two-dimensional representation of inverse pericam.
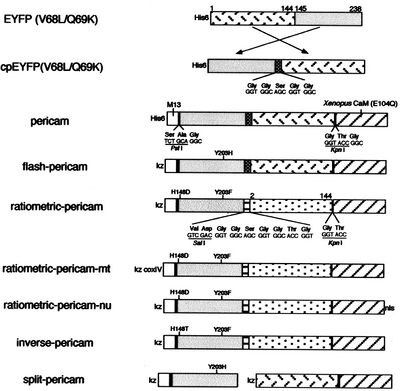
- Begin by downloading this file, which contains the DNA sequence of inverse pericam (IPC) in GenBank format. Open the file in ApE (A plasmid Editor, created by M. Wayne Davis at the University of Utah), which you used during Module 1, and save as a new file called 20109_IPC_YourTeamDay-YourTeamColor.
- In the sequence file, the M13 peptide is highlighted in magenta, the EYFP sequence in yellow, and calmodulin (CaM) in green. Linker sequences connecting these three parts are shown in blue lettering. Refer to Figure 1 of the paper by Nagai et al., which depicts the inverse pericam construct in schematic form, to cement your understanding of how the different components of IPC are connected.
- As you go through the steps below, use the New Feature option on the Features menu to mark the calcium binding sites (and whatever else you deem relevant) in your IPC file. First, you will probably find it helpful to select ORFs → Translate for an amino acid level view of IPC.
- Select the following options for the translation: 1 letter code, line numbers left, DNA below, and copy highlight checked on.
- To help you locate the binding sites for calcium (in the calmodulin portion of IPC), read the following portions of the Zhang paper, along with skimming whatever else you find useful: abstract, first two paragraphs, “Linker and loop flexibility” section.
- In your IPC sequence document, you will mark the amino acid residues that make up the calcium-binding loops in CaM in orange. Begin by looking for the DNA or amino acid sequence of CaM on a site such as NCBI: choose Proteins → Protein Database, search for calmodulin, and scroll down for the amino acid sequence. The CaM sequence is highly conserved across species, so you can refer to almost any sequence and compare it to the one in your file. Are any residues of calmodulin missing in IPC? Why might this be? If you get stuck, use the fact that the CaM within inverse pericam is an E104Q mutant, that is, the 104th residue of calmodulin is Q, to keep yourself oriented.
- Do the four calcium binding loops share any common features? You might imagine that negating or enhancing such features could decrease or increase calcium affinity, respectively.
- If you find other areas of calmodulin that you may be interested in mutagenizing (e.g., hydrophobic pockets), mark these as well. You may find the “Loss of hydrophobic cavities” section in Zhang et al. helpful.
- As you consider sites that may alter calcium binding, keep the following in mind:
- When this module was first debuted, everyone mutated residues directly in the calcium binding loops, and very few groups saw dramatic changes in affinity or cooperativity of calmodulin with respect to calcium. In some years, class-wide results suggested that mutations in the first two binding loops were more likely to have an effect than mutations in the latter two binding loops. Some folks also targeted non-binding structural areas, but results were inconclusive. You may repeat or otherwise build upon prior results as long as you give your own reasoning.
Print out your annotated document and hang on to it for reference. Now let’s put some visuals to all those letters!
Part 2: Higher-order protein features
Unless we are precocious bioengineers indeed, looking at the amino acid sequence alone is unlikely to tell us too much about the protein. We might be left wondering where the binding sites for M13 and for calcium ions are located in calmodulin, for example. In the previous section, you read some primary scientific literature to locate these features. Now you will use a tool called Protein Explorer to visualize them. As you work, you can ask yourself why these stretches of the protein might work the way that they do, and how they might be changed.
- Protein Explorer is a free web-based viewer for biological molecules. To access it, open the Firefox browser and load proteinexplorer.org. Choose “FirstGlance in Jmol” to proceed.
- Structures are organized according to PDB (Protein Data Bank) identification codes, which may be input at the prompt at the top of the page. Begin by looking at the molecule with PDB ID number 1CLL, which is a calcium-bound form of calmodulin. Later you will search for an example of the ligand-free form, also called apo calmodulin.
- The program will open in FirstView mode for the structure you’ve chosen (ensure that popup blockers are off if the structure fails to load). On the right is the image panel, which shows your protein along with associated ligands (in this case, calcium). Try clicking and dragging on the rotating image to see what happens.
- Now look at the control panel on the upper left: here you can modify the image. Try adding and removing water molecules and ligands see where they interact with the protein.
- As you explore the features of the control panel and image panel, be sure to observe the message frame window on the lower left for any relevant information that may pop up. If you click on an atom in the image panel, its atomic identity will be displayed in the message frame, along with its encompassing amino acid residue and position.
- From the control panel, click on the PDB icon, which leads to detailed information about the publication upon which the model image is based.
- To find further options for modifying how you view the image, or search for particular atoms, click on More Views in the control panel, or on Jmol at the bottom right of the image panel. For example, you can highlight specific amino acids, or change from a backbone trace to a space-filling model. Explore these features. For example, you might use color to highlight all the acidic amino acids in calmodulin.
- Be sure to note any useful information in your notebook as you go. You might ask:
- what method was used to elucidate the structure of this protein?
- how good is the image resolution?
- which species did this protein come from?
- when did the authors publish their results?
- what are the major components of the molecule’s secondary structure?
- what do the calcium binding loops (or other areas of interest you found) look like?
- Once you are satisfied with your understanding of calcium-bound calmodulin, bring up an apo calmodulin structure (or two) for comparison. You might find the structure directly by using PDB, or by using the NCBI Structure database. Write a few sentences in your lab notebook describing the differences between the calcium-bound and apo forms of calmodulin.
Part 3: Primer design for mutagenesis
Mutations
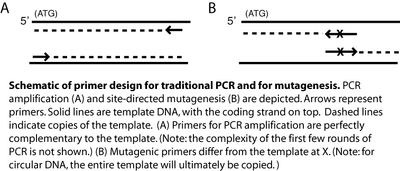
It wouldn’t be very experimentally efficient to somehow pick out and modify a single residue on inverse pericam post-translationally. Instead, researchers genetically encode desired mutations, by making mutated copies of a plasmid originally containing inverse pericam DNA. As you learned in Module 1, DNA polymerases require short initating pieces of DNA (or RNA) called primers in order to copy DNA. Besides non-mutagenic amplification of a specific piece of DNA, synthetic primers can be used for incorporating desired mutations into DNA. For amplification, forward and reverse primers that target the non-coding and coding strands of DNA, respectively, are separated by a distance equal to the length of the DNA to be copied (see figure, part A). In contrast, primer design for site-directed mutagenesis is quite straightforward: both primers are directed at the same location on each strand, and thus will be precisely complementary (see figure, part B). Both direct and mutagenic amplification require cycles of DNA melting, annealing, and extension.
As you know from Module 1, good primers must meet several design criteria in order to promote specificity and efficiency of the desired amplification. Length is one important design feature. Primers that are too short may lack requisite specificity for the desired sequence, and thus amplify an unrelated sequence. The longer a primer is, the more favorable are its energetics for annealing to the template DNA, due to increased hydrogen bonding. On the other hand, longer primers are more likely to form secondary structures such as hairpins, leading to inefficient template priming. Two other important features are G/C content and placement. Having a G or C base at the end of each primer increases priming efficiency, due to the greater energy of a GC pair compared to an AT pair. The latter decrease the stability of the primer-template complex. Overall G/C content should ideally be 50 +/- 10%, because long stretches of G/C or A/T bases are both difficult to copy. The G/C content also affects the melting temperature, which should be high for mutagenesis.
In summary, consider the following design guidelines:
- The desired mutation (1-3 bp) must be present on both strands.
- The mutation should occur approximately in the middle of the primer sequence.
- The primer should be 25-45 bp long.
- A G/C content of > 40% is desired.
- Both primers should terminate in at least one G or C base.
- The melting temperature should exceed 78°C, according to:
- Tm = 81.5 + 0.41 (%GC) – 675/N - %mismatch
- where N is primer length, and the two percentages should be integers.
To demonstrate primer design, the illustration below uses S101L, which is an uninteresting mutation but is a straightforward teaching example.
Residue 101 of calmodulin is serine, encoded by the AGC codon. This is residue 379 with respect to the entire inverse pericam construct, and we can find it and some flanking code in the DNA sequence from Part 1:
361 (5') GAG GAA ATC CGA GAA GCA TTC CGT GTT TTT GAC AAG GAT GGG AAC GGC TAC ATC AGC GCT
381 (5') GCT CAG TTA CGT CAC GTC ATG ACA AAC CTC GGG GAG AAG TTA ACA GAT GAA GAA GTT GAT
To change from serine to leucine, one might choose TTA, TTG, or CTN (wherer N = T, A, G, or C). Because CTC requires only two mutations (rather than three as for the other options), we choose this codon.
Now we must keep 15-20 bp of sequence on each side in a way that meets all our requirements. To quickly find G/C content and see secondary structures, look at the IDT website. (Note that the Tm listed at this site is not one that is relevant for mutagenesis.)
Ultimately, your primer and its complement might look like the following, which has a Tm of almost 81°C, and a G/C content of ~58%.
5’ GG AAC GGC TAC ATC CTC GCT GCT CAG TTA CGT CAC G
3’ C GTG ACG TAA CTG AGC AGC GAG GAT GTA GCC GTT CC
Silent mutations
It is also possible to incorporate silent mutations using SDM. Here, the purpose of the mutation is not to change the protein, but to create a recognizable DNA tag. Thus, the mutation must be silent, that is, it should not affect the protein code. For example, CCA to CCG is a silent mutation, because both triplets code for the amino acid Proline. You can use the NEB table to find degenerate codons.
One category of useful DNA tags are restriction enzyme sites. Recall from Module 1 that these sequences, usually short and palindromic, are recognized by enzymes that cut the sites in unique and specific ways.
GG AAC GGC TAC ATC CTC GCT GCG CAG TTA CGT CAC G
The underlined codon was changed from AGC (Ser) to CTC (Leu). It is residue 379 of inverse pericam and residue 101 of calmodulin.
The point mutation in bold creates the new restriction site for FspI, TGCGCA, which begins within residue 103 of calmodulin.
Part 4: Primer selection for mutagenesis
You will now integrate the information you learned about inverse pericam (especially calmodulin) at the structural and residue levels. Examine the primer sequences below and consider the mutations that each incorporates into inverse pericam. Note: the mutations are written as X#Z, where X is the original amino acid, Z is the modified amino acid, and # is the residue number with respect to calmodulin (not IPC as a whole). For example, residue 379 of inverse pericam is residue 101 of calmodulin, which happens to be a serine, so a mutation at that site to leucine is written S101L.
- Consider the mutation primer sequences below and compare the mutated residues to those that you identified as potentially interesting in Part 1.
| Mutation (X#Z) | Forward primer (5' - 3') |
| D131G | GAT AAG GGA AGC AGA TAT CGG TGG TGA TGG CCA AGT TAA CTA T |
| D133W | GGG AAG CAG ATA TCG ATG GTT GGG GCC AAG TTA ACT ATG |
| D58H | GAA GTC GAT GCG CAT GGC AAA TGG AAC GAT TTA C |
| D93P | CGA GAA GCT TTC CGT GTT TTT CCC AAG GAT GGG AAC GGC |
| D22W | GCC TTC TCA TTA TTC GAC AAG TGG GGA GAC GGC AAC ATC ACC |
| E31R | ACC ATC ACC ACA AAG AGG CTG GGC ACC GTT ATG AGG |
| M124S | GAA GTT GAT GAA TCG ATA AGG GAA GCA GAT ATC GAT GG |
| L32D | GGC ACC ATC ACC ACA AAG GAA GAT GGT ACC GTT ATC AGG |
| T79P | GAA AAA TGA AGG ACC CAG ACA GCG AAG AGG AAA TCC |
- Choose one modification that you hyphothesize might increase or decrease CaM’s affinity for calcium (or M13), or might affect cooperativity among the four calcium binding sites. Be sure to include your hypothesis for the effect of this mutation in your lab notebook.
- What is the sequence for the reverse primer (in the 5' to 3' direction)?
- Compare the sequence of the forward primer to the native (or wild-type) sequence of inverse pericam. Locate the silent mutation that will be incorporated with this primer. Where does the amino acid substitution occur within the protein? What is the amino acid substitution? What restriction enzyme recognition sequence does the silent mutation create within the mutated sequence?
- Hint: use APE to compare the sequences and identify the restriction enzyme site.
- Lastly, use the design guidelines in Part 3 to examine the primer. Is this a 'good' mutation primer? Use the information you collect to support your decision to use this mutation primer.
- Feel free to select a different mutation primer if you are not satisfied with your first choice at this point.
- Before you leave today, note which primer you chose on today's Talk page. Also, include the information concerning the silent mutation and restriction enzyme recognition site.
Part 5: Site-directed mutagenesis
We will be using the QuickChange® II kit from Stratagene to perform our site-directed mutageneses. Each group will set up one reaction, for their chosen X#Z mutation. Meanwhile, the teaching faculty will set up a single positive control reaction, to ensure that all the reagents are working properly. You should work quickly but carefully, and keep your tube in a chilled container at all times. Please return shared reagents to the ice bucket(s) from which you took them as soon as you are done with each one.
- Read through the following protocol and prepare all calculations before beginning physical manipulations of your samples.
- Get a PCR tube and label the top with your mutation and lab section (write small!). Add 43 μL of "Master Mix" - containing buffer and dNTPs - to your tube. Be sure to use a fresh pipet tip, as several groups will share each aliquot of Master Mix!
- Add 2 μL of template DNA (“IPC plasmid”) to the reaction tube.
- Note: mutagenesis reactions are expected to run smoothly with 5-50 ng of plasmid DNA. You have been given a 1:200 dilution of miniprep DNA.
- Add 5 μL of diluted primer solution (containing both forward and reverse primers) to the tube. The volume of the reaction should be now be 50 μL.
- Finally, add 1 μL of PfuUltra DNA polymerase (do NOT mix the enzyme stock when you take from it) to the reaction using the P20 and extra small tips up front, and then mix the reaction thoroughly by pipetting with your P200 set at 40 μL.
- Once each group is ready, we will begin the thermocycler, under the following conditions:
| Segment | Cycles | Temperature (°C) | Time |
|---|---|---|---|
| 1 | 1 | 95 | 30 sec |
| 2 | 18 | 95 | 30 sec |
| 55 | 1 min | ||
| 68 | 5 min | ||
| 3 | 1 | 4 | indefinite |
- After the cycling is completed, the teaching faculty will add 1 μL of DpnI to the reaction mixture and pipet to mix. Samples will be incubated for one hour at 37 °C then stored at -20 °C.
Reagent list
- QuikChange II Site-Directed Mutagenesis Kit from Stratagene
- 10X Reaction Buffer (100 mM KCl, 100 mM (NH4)2SO4, 200 mM Tris-HCl, 20 mM MgSO4, 1% Triton® X-100, 1 mg/mL BSA)
- PfuUltra® DNA polymerase (2.5 U/μL, 1 μL per rxn)
- dNTP mix (proprietary mix, 1 μL per rxn)
- Dpn I (10 U/μL)
Navigation links
Next day: Prepare expression system
Previous day: M1D7 Data analysis