BISC219/F11: RNAi Lab 9: Difference between revisions
No edit summary |
No edit summary |
||
| Line 58: | Line 58: | ||
== Assignment == | == Assignment == | ||
Remember to check the Assignment section of the wiki for instructions about the graded assignment due in the next lab and check the Weekly Calendar for other work to accomplish before the next lab. | Remember to check the Assignment section of the wiki for instructions about the graded assignment due in the next lab and check the Weekly Calendar for other work to accomplish before the next lab. | ||
<div class=noprint> | |||
== Outline of Experimental Design for REVERSE Genetics Project == | |||
'''Where are you now in this process?''' (What have you done so far? What's next?)<BR> | |||
'''A.''' Make the feeder strain of bacteria<BR> | |||
# Amplify gene of interest by PCR <BR> | |||
# Restriction Enzyme digestion of amplified DNA to create "sticky ends" for ligation<BR> | |||
# Clean up DNA (remove enzymes) <BR> | |||
# Cloning: ligate gene into vector plasmid with amp resistance gene <BR> | |||
# Transform competent bacterial cells | |||
# Select for transformants on media with ampicillin<BR> | |||
# Perform colony pcr on several transformants to be sure to find one colony containing a vector plasmid with the gene of interest | |||
# Culture the selected colony from colony pcr to create a lot of copies of these bacteria | |||
# Isolate the cloned plasmid DNA from that cultured colony by miniprep<BR> | |||
# Retransform isolated plasmids (with gene interest) into HT115 (DE3)cells genetically modified to have impaired ability to degrade RNA<BR> | |||
# Select for transformants on media with ampicillin | |||
# Choose an isolated colony to culture and make lots of feeder strain bacteria <br> | |||
# Induce expression of ''C. elegans'' gene dsRNA from the pL4440 vector in the bacteria by IPTG induction <br> | |||
# Seed NM lite worm growth media plates with feeder strain produced as described <BR> | |||
'''B.''' Plate wild type ''C. elegans'' worms (N2 and ''rrf-3'' strains) on feeder plates made as described (containing bacteria expressing dsRNA of our gene of interest). <BR> | |||
'''C.''' Observe phenotype change in progeny caused by RNAi silencing or knockdown of the gene of interest compared to control worms of same strains that were NOT fed feeder strain bacteria.<BR> | |||
</div> | |||
Revision as of 16:59, 17 May 2011
Lab 9: Series 3 Reverse Genetics-Induction of HT115(DE3) feeding strains for RNAi
Through reverse genetics we will deduce the function of a gene starting with its sequence and working back to its phenotype. There are many genes in the genome whose phenotype when mutated is lethal; therefore, it's impossible (or very difficult) to tie function to a particular gene in the traditional forward genetics manner of creating random mutations, looking for phenotype changes, and then finding the defective gene responsible for that function.
In our reverse genetics study of some interesting C. elegans genes, two different strains of worms, wild-type and rrf-3 (RNAi enhanced), are fed bacteria expressing dsRNA specific to a particular worm gene. Ingesting dsRNA initiates cascade of events that leads to the destruction of the mRNA of the target gene. An altered phenotype in the progeny of RNAi-treated worms indicates what happens when the normal function of this gene is lost or significantly downregulated.
Double stranded RNA (dsRNA) can be introduced to the C. elegans cells in many different ways including: feeding, injection and soaking. Each of these methods has positives and negatives. We are using the feeding method - where we use genetically modified bacteria as dsRNA factories.
To begin to investigate the power of reverse genetics, you will need to grow your own induced bacteria to seed your plates for RNAi feeding.
Done on the night before this lab:
You and your partner made an overnight broth culture of your selected colony. This process created a sub-culture of many identical copies of the plasmid carrying the construct that will induce RNAi to silence or downregulate the gene that you want to study.
On the morning of lab:
Your instructor or the lab staff will come in early in the morning and sub-culture your bacterial overnight. The cells will be in stationary phase in the morning and successful induction requires log phase growth.
To create the subculture of bacteria your cultures will be diluted 1:5 (1 mL of culture into 4 ml of LB + amp). These cultures will be allowed to grow until lab time - approximately 3-4 hours.
When you come in to lab you will induce your cultures to make lots of dsRNA by adding IPTG to the culture and letting it continue to incubate for a few hours so the cell is full of dsRNA. The IPTG will compete with the repressors on the lac o promoter and remove them and allow the gene for T7 RNA polymerase to be transcribed and then translated into the RNA polymerase protein. The T7 RNA polymerase then binds to the T7 promoters on the pL4440 plasmid and transcribes our C. elegans DNA into RNA!
A simplified map of the C. elegans RNAi plasmid :
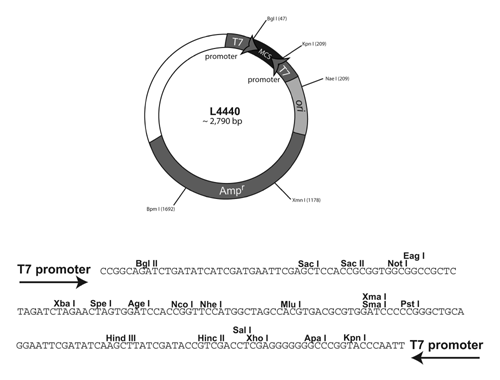
More specifically: The bacteria cells of strain HTll5(DE3) contain the T7 RNA polymerase gene (contained within a stable insertion of a modified lambda prophage λ DE3) under the control of lac operon regulatory elements. This allows expression of T7 polymerase to be controlled by isopropyl-β-D-thiogalactopyranoside (IPTG), a lactose analogue that induces expression of genes under the control of the lac operon o gene. When IPTG is added, the cells will begin to synthesize lots of T7 RNA polymerase. This T7 RNA polymerase can then bind to the T7 promoter sites on the plasmid and begin to synthesize RNA from both T7 RNA polymerase sites. Because the two single strands of RNA are complementary to each other they will form double stranded RNA within the bacterial cell. The IPTG induction allows us to "turn on" and express the plasmid gene of interest only when we want to and it allow us to make much higher levels of RNA for RNA interference than would be made without this induction.
Another useful thing about E coli strain HT 115(DE3)is that this particular strain is deficient for the RNAaseIII enzyme that degrades double stranded RNA (dsRNA) in the bacterial cell. This allows for the accumulation of dsRNA in the cell and, thus, our ability to induce and RNAi effect! This E. coli strain carries a tetracyclin resistance gene so these cells can be selected on media containing tetracyclin, while the plasmid contains an ampicillin resistance gene that allows only transformed cells to grow on media containing ampicillin.
Our goal is to upregulate production of dsRNA of our worm gene of interest in pL4440 vector plasmids containing that gene in HT115(DE3) bacteria.
To induce your cultures:
- Add 5 μL of 0.5 M IPTG to your culture. What is the effective concentration of IPTG in your culture?
- Put your culture back in the 37°C incubator in the spinning wheel for approximately 3-4 hours.
- After the lab introduction, we will head to a more comfortable room to discuss the papers you were assigned.
To do after induction is complete:
- Pour your culture into a 15 ml orange cap centrifuge tube.
- Spin your culture in a table top centrifuge for 5 minutes at 3000 rpm.
- Remove 3.5 ml of the supernatant.
- Resuspend the bacterial pellet in the remaining 1.0 ml of supernatant - you are concentrating your bacteria.
- Pipet a 200 μL aliquot of your induced bacteria onto the center of 4 feeding plates. Be careful not to tilt or jostle your plates so that the bacteria stay in a circle in the center of each plate. These plates contain the same NGM Lite medium used in our mapping series, except that they have been supplemented with 0.4 mM IPTG, 50 μg/mL ampicillin.
- Allow the bacteria to be absorbed into the media before you move them.
- Obtain 2 control plates - these plates contain the same NGM lite medium described above and the bacterial strain on them are identical to your RNAi feeder strain, except that the pL4440 plasmid is only expressing RNA from the vector - it lacks DNA specific to any worm genes.
- Stack all 6 plates carefully (without tilting them) - put them in your lab section box or your project box identified with a labeled piece of your team color tape on top.
- We will allow the bacteria to continue to induce overnight at room temperature.
4 days before next lab:
- Come into lab and find your stack of plates.
- On 2 of the experimental plates add 2 L4 wild type (N2) hermaphrodites
- On 2 of the experimental plates add 2 L4 rrf-3 hermaphrodites
- On 1 of the control plates add 2 L4 wild type (N2) hermaphrodites
- On 1 of the control plates add 2 L4 rrf-3 hermaphrodites
- Wrap the 3 plates of each strain (N2 or rrf-3) with elastics and incubate the N2 worms in your lab day box in the worm incubator set at 23°C. Incubate the 3 plates of rrf-3 strain worms at room temperature in your lab day project box in the stock area. DO NOT incubate the rrf-3 worms at 23C!!!!
You will score your phenotypes in the next lab.
Assignment
Remember to check the Assignment section of the wiki for instructions about the graded assignment due in the next lab and check the Weekly Calendar for other work to accomplish before the next lab.