Quantifying sequence libraries with qPCR
Curator
Abstract
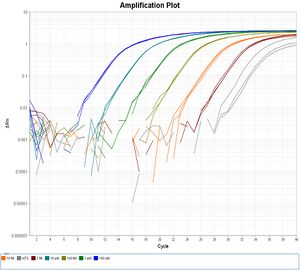
Compared to other methods like the Nanodrop, Qubit/PicoGreen, and Bioanalyzer/TapeStation, qPCR is the most accurate way to quantify sequencing libraries. This is because it only detects molecules that have complete sequencing adapters at both ends, which are the ones that will successfully generate sequence reads. The purpose-made Kapa Library Quantification Kits include specific primers and standards as well as qPCR reagents. However, these kits are based on SYBR Green I, a dye that nonspecifically binds all DNA: the fluorescence signal is proportional to molecule lengths, therefore you must normalize all your estimated molarities according to their average molecule lengths, which are variable in most kinds of sequencing libraries. For this application, SYBR Green I qPCR can be inconvenient and depends on the accuracy of a secondary measurement like the Bioanalyzer, which may be unreliable if the libraries have unusual length distributions or contain unsequenceable byproducts.
This protocol, based on Quail et al. 2008, describes qPCR using dual-labeled hydrolysis probes (TaqMan). In this approach, each sequenceable DNA molecule only hybridizes with a single reporter probe, so the fluorescence signal is insensitive to molecule lengths and no normalization is required. This protocol is less expensive than the Kapa kit as well (~$50 per full 96-well plate vs. ~$60 per full plate at current US list prices).
Materials
Reagents and consumables
- Hydrolysis-probe qPCR mix (containing polymerase with 5′→3′ exonuclease activity and no SYBR dye), e.g. Thermo Fisher #444456
- Molecular biology–grade water
- Basic buffer for DNA dilution and storage, ideally with mild nonionic detergent, e.g. Tris+Tween or TE+Tween (note relatively large volume required)
- Compatible library of known molarity, e.g. Illumina #FC-110-3001
- Pipet tips and tubes/plate, low-retention recommended, e.g. Eppendorf Dualfilter T.I.P.S.® LoRetention,
- qPCR plate
Oligonucleotides (Illumina-specific)
P5: 5′-AAT GAT ACG GCG ACC ACC GA-3′
P7: 5′-CAA GCA GAA GAC GGC ATA CGA-3′
Probe: fluorophore-5′-CCC TAC ACG ACG CTC TTC CGA TCT-3′-quencher
HPLC purification is required for the probe; no special purification is required for the primers.
Warning: these sequences are compatible with the standard TruSeq adapters, but may not work with some others such as Nextera; see the Customer Sequence Letter to make sure.
Purchasing: select the cheapest purification option (e.g. "Standard desalting") for the primers and HPLC purification for the probe. A good combination of fluorophore and quencher is FAM and IDT's Iowa Black FQ (PrimeTime 5' 6-FAM/ZEN/3' IBFQ).
Oligonucleotide sequences © 2016 Illumina, Inc. All rights reserved. Derivative works created by Illumina customers are authorized for use with Illumina instruments and products only. All other uses are strictly prohibited.
Procedure
Preparing oligonucleotide stocks
- Resuspend lyophilized oligonucleotides at ~100 μM, in suitable DNA storage buffer, according to manufacturer's stated yields.
- (Optional) Measure actual concentrations by UV spectrophotometry.
- Prepare 10 μM dilution of each oligonucleotide, or a mix of all three (at appropriate concentrations) to save time.
- Store at −20 °C in the dark.
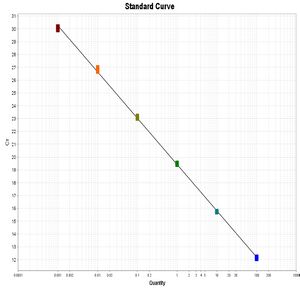
Preparing standard dilutions
- Dilute library of known molarity to 100 pM, e.g. 2 μL Illumina PhiX Control (10 nM) + 198 μL DNA dilution buffer. Mix very well.
- From this, make serial 1/10 dilutions down to 10 fM (5 total concentrations), e.g. 20 μL previous dilution + 180 μL DNA dilution buffer.
- Separate the dilutions into single-use aliquots.
- Store the aliquots at −20 °C and avoid thawing them until use.
![]() Pipet very carefully when preparing your standards! Reverse-pipetting and low-retention plastics may improve your accuracy.
Pipet very carefully when preparing your standards! Reverse-pipetting and low-retention plastics may improve your accuracy.
Diluting sample libraries
- Dilute each library stock 1/100 in DNA dilution buffer (use the same one as you did with the standards), e.g. 1 μL library + 99 μL buffer. Mix very well.
- Redilute each of these dilutions 1/100, e.g. 1 μL first dilution + 99 μL buffer. Mix very well.
The final dilution, ready for qPCR, is 1/10,000.
A 96-well plate and multichannel pipet are helpful to increase the speed and reproducibility of the sample preparation. Reverse-pipetting and low-retention plastics may improve your accuracy.
Example reaction mix (per plate well)
| Probe qPCR mix, 2X | 5.0 μL |
| Illumina P5 qPCR primer, 10 μM | 0.4 μL |
| Illumina P7 qPCR primer, 10 μM | 0.4 μL |
| Illumina qPCR probe, 10 μM | 0.2 μL |
| molecular biology–grade water | 2.0 μL |
| diluted library or standard, 1 fM to 100 pM | 2.0 μL |
Prepare a master mix with all reagents except the library/standard. Use three replicates per sample. Include the standards and a no-template control on every plate.
Example program
| UDG treatment | 50 °C | 2:00 | (only if master mix contains UDG; see its user guide) |
| Initial denaturation | 95 °C | 3:00 | (one time only) |
| Denaturation | 95 °C | 0:03 | 40 cycles |
| Annealing + extension + acquisition | 60 °C | 1:00 |
This program requires a qPCR mix capable of "fast" two-step cycling.
![]() The extension time is long in order to accomodate longer molecules than are normally used in qPCR.
The extension time is long in order to accomodate longer molecules than are normally used in qPCR.