Bioanalyzer
From OpenWetWare
Jump to navigationJump to search
| back to protocols | ||
The Bioanalyzer is a chip-based capillary electrophoresis machine to analyse RNA, DNA, and protein. It is produced by Agilent and widely used, among other things, in RNA quality control measurements before downstream experiments like microarrays.
Protocol for Bioanalyzer RNA nano chip
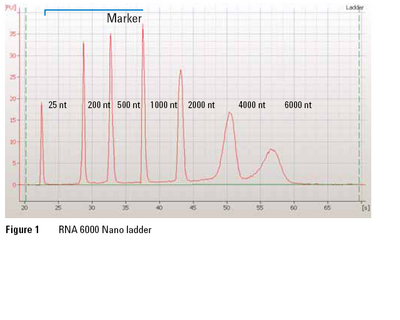
Preparation of material
- Prepare a maximum of 12 samples per chip.
- Maximum concentration recommended is 500ng/µl, 1000ng/µl is okay.
- Denature RNA 70°C 2min, cool on ice.
Cleaning, gel preparation (start 40min before experiment)
- take out filtered gel aliquot and fluorescent dye from fridge next door 30min ahead of time (1 gel aliquot tube enough for 2 chips)
- take out ladder from -80 freezer nextdoor (r145), column 5, nano white box / pico yellow
- vortex dye 10s and spin down
- mix one tube gel aliquot with 1µl of dye
- vortex, centrifuge 13000g +-20% for 10min (12000 rpm on Biofuge pico)
- wash electrodes 1min RNase ZAP, 2x30s water (pipette 500µl into any well, spreads from there)
Prepare chip for measurement
- take out new RNA chip (pico or nano) from drawer below microarray machine and put into station
- fill 9µl gel into dark circle G well (gel reservoir)
- press down plunger and wait 30sec (gel moves through channels)
- after 30sec release silver trigger (should jump up well above 0.3, meaning seal was tight)
- wait 5sec, pull plunger up all the way
- open priming station, add 9µl gel to light circle G wells
- 5µl of marker into all sample wells and ladder well, bottom right (contains small marker to align plots)
- 1µl of sample per well, add 1µl of water or replicates into free wells (6µl required per well for machine to run properly)
- 1µl of ladder into ladder well
- vortex in holder for 1min and put into machine
Start the run
- start 2100 expert software
- choose programme for kit: DNA/RNA pico/RNA nano and material: total RNA/mRNA, prokaryotic/eukaryotic
- start run
Wash, export data
- typical RNA nano run takes just over 20min
- wash 2x30x w water immediately after the run has ended (electrodes in the lid will quickly deteriorate otherwise)
- to export data as PDF you have to print; (PDF option checked will save file, PDF unchecked will print on paper)
Expected results
- expect quantitative range 25–500 ng/μl [claim from the manual]
- expect quantitation accuracy 20%CV (for ladder as sample) (CV = coefficient of variance) [claim from the manual]
Test run with mouse total RNA
- 6 replicas, 378 ng/µl according to nanodrop, Qiagen RNeasy column purified
Bioanalyzer results:
- mean concentration 397 ng/µl ± 18 ng/µl (CV 4%); range 371-417;
- measured RINs: 10, 10, 10, n/a, n/a, 10
Conclusions:
- RNA concentration in good agreement w nanodrop & tighter than expected spread
- automatic graphical analysis of electropherograms failed in every 3rd replica!!
Protocol for Bioanalyzer RNA pico chip
The protocol is very similar for the PICO kit. Similar but reagents adjusted for this kit are used, i.e. a different ladder, marker, and gel.
- quantitative range 50–5000 pg/μl total RNA, 250-5000 pg/µl mRNA
- quantitation accuracy 30%CV (for ladder as sample)
- recommended buffer 50 mM Tris or 50 mM NaCl (assay sensitive to high salt)
Unlike in the nano kit the conditional buffer well has to be filled when using the pico kit.
Stability of reagents
- manual warns that gel-dye mix should be kept at 4°C if stored for more than 1h to prevent degradation
- use gel-dye mix within 4 weeks after preparation
- dye is light sensitive! keep in the dark (but it's not that 2h under electric light in the lab will render your gel-dye mix unusable)
Troubleshooting & tips
A few typical problems occur once in a while:
- small RNA marker is not properly recognised (often the next peak is selected => fragments size mislabelled)
- cleak peak table, a tab at the bottom of the window; lower marker can be manually set here
- 18S, 28S not properly assigned (=> RIN number incorrect)
- peak baseline misplaced resulting in incorrect area calculations for rRNA peaks (adjust by hand)
- obscure "serial port" communication errors occurs frequently but can be ignored and has no impact on the readings
- physical movement of the bioanalyzer can cause errors but small movements like vibrations from a shaker on the same becnh are typically ok
Tips:
- pipette gel-dye mix from the top of the tube after centrifugation to reduce the risk of larger particles obstructing the gel run or of dye agglomerates influencing the fluorescent read
Comments
- The Bioanalyzer is a VERY FUSSY machine.
- - Replicas of the same sample run on the same chip often give different results or none at all;
- - RNA from trizol isolation often does not yield usable data, probably to carry over chemicals from the purification process;
- - RINs are often not calculated for good looking RNA electropherograms due to the recognition software failing. --- JS 05:24, 5 May 2011 (EDT)
- I have sometimes skipped the RNA denaturation steps without negative impact on the results (no detectible difference in electropherogram, RIN of 10). Not denaturing may influence the quantitation (see trouble shooting guide where it's listed as a reason for too high and too low RNA quantitation). --- JS 08:42, 5 May 2011 (EDT)
See also
- RNA Quality Control
- Agilent has a Bioanalyzer Maintenance & Troubleshooting Guide v2001 v2003