Klinke:Research
The Klinke Lab @ West Virginia University
Home
Research
Lab Members
Publications
Talks
Positions
Contact
Summary of Research Program
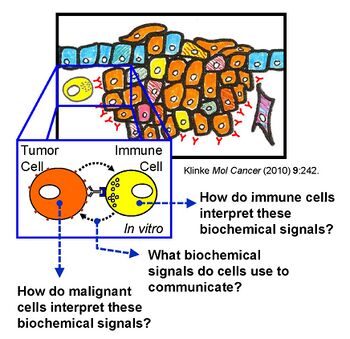
Our vision is to identify and potentially develop novel therapies for treating cancer. To accomplish this vision, we aim to identify alterations in heterocellular communication that locally suppress anti-tumor immunity. We believe that targeting these alterations may provide a therapeutic advantage that distinguishes between heterocellular communication occurring in normal versus malignant tissue. Our approach blends wet experiments, mathematical modeling and simulation, and data analytics to varying degrees. In dissecting heterocellular communication, specific projects may involve probing cellular cross-talk (e.g., Kulkarni et al. Integr Biol (Camb) 2012) or characterizing how a specific cell type may interpret these communicated signals (e.g., Klinke et al. Sci Signal 2012, Klinke et al. Mol Biol Cell 2015), as illustrated in the graphic. A Bayesian perspective permeates throughout the work - a la designing experiments that minimize the influence of a priori bias (e.g., proteomics and high content assays) and analyzing data in a way that maximizes insight into system behavior (e.g., model-based inference).
Model-based inference is the process of encoding our prior knowledge of how cells interpret biochemical signals in the form of a mathematical model. The model is then used to test whether our prior knowledge is consistent with the experimental data. Model-based inference is a key tool as biological systems exhibit intrinsic uncertainty - due to either ethical constraints or due to technical limitations of the available experimental techniques. By combining experimental study with model-based inference, we hope to obtain greater fidelity in understanding how these systems work than could be obtained using either technique in isolation.
Integrative Systems Approach to Identify Local Oncogenic Modulation of the IL12 Axis
Collaborators: J. Michael Ruppert, Yon Rojanasakul, West Virginia University
Funding source: National Cancer Institute
Immunotherapies, like immune checkpoint modulators, are poised to transform the therapeutic landscape for cancer. While these therapies elicit remarkable responses in a subset of patients, a key hurdle for broadening the clinical benefit is identifying collateral targets that relieve tumor-induced changes in the tissue microenvironment that block endogenous anti-tumor signals. To identify these collateral targets, we have developed a phenotypic screen to identify mechanisms that malignant cells use to block immune cell response to cytokines that promote anti-tumor immunity, such as Interleukin-12. The objective of this Bioengineering Research Grant is to identify collateral targets in breast and lung carcinomas by screening a set of transplantable models for these cancers and to validate these targets using pre-clinical mouse models and human data. Given that oncogenesis is an evolutionary process involving repeated mutation and selection, our central hypothesis is that malignant cells evolve to secrete proteins that collectively cross-regulate the response of CD4+ and CD8+ T cells to Interleukin-12 (IL12), a potent adjuvant of anti-tumor immunity. To test our central hypothesis, we will first establish conditions where a T cell response to IL12 can be predicted using a mechanistic math model. Next, we will recreate tumor-induced immunosuppression in vitro, where the mathematical model will be used to define how the T cell response to IL12 is altered upon co-culture with the different transplantable models for lung and breast cancer. Proteins secreted by tumor cells that recreate the observed phenotype will be identified using LC-MS/MS-based proteomics. Immunosuppressive mechanisms identified in vitro will be validated using pre-clinical mouse models and a retrospective cohort study using data obtained from large cancer studies that relate `omics with clinical outcomes. This multidisciplinary approach is projected to yield the following expected outcomes: 1) enable stratifying patients based on mechanistic pathology, 2) guide developing drugs against these collateral targets, and 3) identify patient cohorts that are likely to benefit from specific combination immunotherapies.
CAREER: Interrogating Antagonistic Mechanisms of Signaling Cross-talk in Natural Killer Cells
Collaborators: Jonathan Bramson, McMaster University, Hamilton, ON
Funding source: National Science Foundation

This NSF award by the Biotechnology, Biochemical and Biomass Engineering program supports work to improve our fundamental understanding of how cancer cells escape the cytotoxic action of monoclonal antibodies. Monoclonal antibodies comprise one of the largest classes of cancer drugs that target molecules unique to cancer cells. However, the emergence of resistance to molecular targeted therapies is an increasing, and poorly understood, problem. Without improved understanding of how cancer cells resist the action of molecular targeted therapies, designing effective treatments will remain limited. To improve ultimately the effectiveness of mAbs as cancer drugs, we propose a conceptually novel approach that combines aspects of cellular engineering, immunology, cancer biology, and computationally intensive model-based inference. The research objectives are integrated with educational objectives that aim to promote cross-disciplinary communication among experts and to improve the ability of scientists and engineers to communicate scientific concepts, like how theory and computation are used in scientific practice, effectively with the lay public. It is expected that these aims will have an impact that ranges from local to international. At the local level, the proposed research will provide interdisciplinary training opportunities for graduate and undergraduate students at the interface between multiple disciplines, including biochemical engineering, cancer biology, molecular biology, immunology, and pharmacology. The proposed education aims will also focus outward to create scientists and engineers that can collaborate more effectively across disciplines and, more importantly, that can convey what they do and it's importance to the lay public. Finally, the fundamental fruits of this research may be applied to improve therapies for cancer, a disease that, in developed countries, kills one in three.