IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-8-1
<html><style type='text/css'> .tabs {
font-size:80%;
font-weight:none;
width: 100%;
color: #FFFFFF;
background:#FFFFFF url("/images/5/54/DarkgreenTab-bg.gif") repeat-x bottom;
}
.tabs li {
background:url("/images/3/36/DarkgeenTab-left.gif") no-repeat left top;
}
.tabs a,.tabs strong {
background:url("/images/d/d3/DarkgreenTab-right.gif") no-repeat right top;
color:#FFFFFF;
padding: 3px 10px 3px 4px;
}
.tabs strong{
color:#CCFF00;
background-image:url("/images/b/b1/DarkgreenTab-right_on.gif");
}
.tabs a:hover{
color:#66FF00;
}
</style></html>
Gel images from yesterday's mutagenesis PCR #7
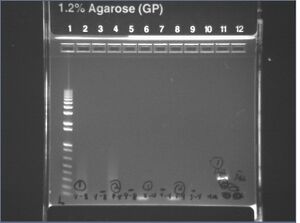
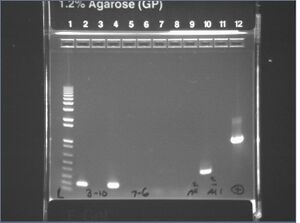
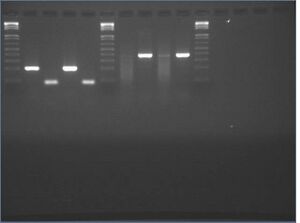
It looks like the 3-10 and 5-4 mutagenesis PCRs succeeded, but the 7-6 and 9-8 did not. Also, we got a strange 400b band from the crossF + crossR reaction instead of the 3kb we predicted. The band is reminescent of the 400b band we saw from genomic extraction #5
NOTE: Raw PCR products of interest were put in .5mL tubes in the freezer in a box labeled with 8.1.
Gel images from yesterday's split-extraction PCR
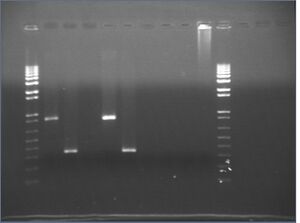
It looks like our KaiA and KaiB extractions succeeded, but the KaiC extraction did not.
Gel purification: done, 4 tubes labeled in yellow, 8.1.06
- The gel extractions were completed on both the mutagenesis cutouts and on the KaiA and KaiB segments
- The weights of each are as follows:
- 0.0946g
- 0.2964g
- 0.4212g
- 0.2055g
- 0.2049g
- 0.2750g
- KaiA (1) 0.1277g
- KaiB (1) 0.1438g
- KaiA (2) 0.2004g
- KaiB (2) 0.1475g
Plan for the next few days
We're going to do BioBrick assembly of the KaiA and KaiB we extracted last night, keep trying to extract KaiC from the cyanobacteria genome, investigate the goofy 400b band, and try to get the other mutagenesis reactions to work.
- Biobrick assembly of KaiA and KaiB
Preparation of backbone- Nick has prepared high-copy plasmids for us to use as backbones (they currently have an RFP insert which must be removed).
Miniprep the cultures that Nick providedDigest the plasmids at X and S.Add phosphatase to the backbone to prevent self-ligation.Run the digested plasmids on a gel to separate the backbone from the insert (and also run some undigested plasmid for comparison).Gel-extract the backbone.
- Ligation of KaiA/B with backbone
Gel purify the KaiA and KaiB bandsDigest KaiA/B at X and S.- Ligate KaiA/B inserts with the backbone prepared above.
- This ligation can produce backwards insertions and tandem insertions (e.g. KaiA+KaiA), but we can screen them out later when we do our X and P digest.
- At this point we can transform and amplify our KaiA/B inserts.
- After we've amplified the plasmid, digest the resulting plasmid at X and P (or E and S).
- Run the result on a gel to screen for our construct.
- Extraction of KaiC
- We need to talk about how to make this work
- Investigation of 400b band
- Hunt for candidate primer binding locations within the KaiABC sequence
- Sequence the 400b band?
- We can try sequencing the band with the crossF primer (primer 3)
- We also have Topo clones of the other 400b band we extracted from genomic extraction #5, but we probably want to digest them first to make sure they have the right length insert before we send them off for sequencing (using the M13 primers).
- Also, we're not 100% sure that our previous 400b band is the same as the one Peng found today
- Continuing mutagenesis
- Gel-purify the succesfull mutations (3-10 and 5-4)
- We're retrying the 9-8 and 6-7 mutations with gradient PCR (see next section)
- We're also trying to amplify the LC1PUR1 and LC2PUR1 3kb segments because we don't have much of them left (see next section)
Extracting longer segments via gradient pcr (mutagenesis #8)
As during the last PCR we were able to get 2 of the 4 segments and we got a 400bp band for "all" which was similiar to the colony PCR band, I hypothesize that a gradient PCR may show us a 3kb "all" band as well as our other products. Hotstartaq as usual.
Vials
- 9-8, 6-7, all, -t for each, cat gene as a positive control, master mix
- 6 for each + negative
- 24 total vials
For cat gene, pbc_ks+, kt_b, kt_bb primers - expected size ~1.1kb
Back to basics: HotStarTaq/Farren Protocol
5 µL 10x buffer (HotStar) 1.0 µL 10 mM dNTPs 1.0 µL 20 µM forward primer 1.0 µL 20 µM reverse primer 1.0 µL plasmid DNA 0.5 µL HotStarTaq DNA polymerase 40.5 uL dH20
PCR Time
#*95 °C for 15 min. (melt) #*95 °C for 0.5 min. (melt) #*GRADIENT °C for 0.5 min. (anneal) #*72 °C for 2.5 min. (extension) #* Go to step 2 (30x) #*72 °C for 10 min. (final extension) #* Keep at 4 °C forever
Total time: 111 minutes +/- some = 120min
Miniprep of RFP-insert BioBrick plasmids
We miniprepped 20 mL of transformant cells carrying a BioBrick plasmid with RFP insert. There are 4 vials sitting in our 4C refrigerator, with the ng/µL concentrations written on the side (eluted in 30 ng/µL).
Gel purification of KaiA/B
6 tubes labeled in red, 8.1.06, nanodrop measurements were all lower than 10ng/uL.