IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-7-25
<html><style type='text/css'> .tabs {
font-size:80%;
font-weight:none;
width: 100%;
color: #FFFFFF;
background:#FFFFFF url("/images/5/54/DarkgreenTab-bg.gif") repeat-x bottom;
}
.tabs li {
background:url("/images/3/36/DarkgeenTab-left.gif") no-repeat left top;
}
.tabs a,.tabs strong {
background:url("/images/d/d3/DarkgreenTab-right.gif") no-repeat right top;
color:#FFFFFF;
padding: 3px 10px 3px 4px;
}
.tabs strong{
color:#CCFF00;
background-image:url("/images/b/b1/DarkgreenTab-right_on.gif");
}
.tabs a:hover{
color:#66FF00;
}
</style></html>
Re-innoculation of HH6 from freezer stock
Done at night, 2 re-innoculations off of frozen stock to replenish supply.
Miniprep of reinoculated Hetmann Six
Eluted in 50 uL of H2O. Nanodrop readings:
HH1: 111.5 ng/uL HH2: 47.5 ng/uL HH3: 19.7 ng/uL HH4: 64.0 ng/uL HH5: 54.2 ng/uL HH6: 44.3 ng/uL
Sequencing reactions
We will send out 42 sequencing reactions to GeneWiz tomorrow (6 samples, 7 reactions each). These samples are the Hetmann Six-- Topo + KaiABC (we hope)-- which we have been using in our more recent mutagenesis and splitting attempts.
| Label | Sample | Primer | [Template] ng/uL |
| CY101 | HH1 | seq_A1 | 55.75 |
| CY102 | HH1 | seq_A2 | 55.75 |
| CY103 | HH1 | seq_A3 | 55.75 |
| CY104 | HH1 | seq_A4 | 55.75 |
| CY105 | HH1 | seq_A5 | 55.75 |
| CY106 | HH1 | M13F | 55.75 |
| CY107 | HH1 | M13R | 35.63 |
| CY201 | HH2 | seq_A1 | 35.63 |
| CY202 | HH2 | seq_A2 | 35.63 |
| CY203 | HH2 | seq_A3 | 35.63 |
| CY204 | HH2 | seq_A4 | 35.63 |
| CY205 | HH2 | seq_A5 | 35.63 |
| CY206 | HH2 | M13F | 35.63 |
| CY207 | HH2 | M13R | 35.63 |
| CY301 | HH3 | seq_A1 | 14.78 |
| CY302 | HH3 | seq_A2 | 14.78 |
| CY303 | HH3 | seq_A3 | 14.78 |
| CY304 | HH3 | seq_A4 | 14.78 |
| CY305 | HH3 | seq_A5 | 14.78 |
| CY306 | HH3 | M13F | 14.78 |
| CY307 | HH3 | M13R | 14.78 |
| CY401 | HH4 | seq_A1 | 48 |
| CY402 | HH4 | seq_A2 | 48 |
| CY403 | HH4 | seq_A3 | 48 |
| CY404 | HH4 | seq_A4 | 48 |
| CY405 | HH4 | seq_A5 | 48 |
| CY406 | HH4 | M13F | 48 |
| CY407 | HH4 | M13R | 48 |
| CY501 | HH5 | seq_A1 | 40.65 |
| CY502 | HH5 | seq_A2 | 40.65 |
| CY503 | HH5 | seq_A3 | 40.65 |
| CY504 | HH5 | seq_A4 | 40.65 |
| CY505 | HH5 | seq_A5 | 40.65 |
| CY506 | HH5 | M13F | 40.65 |
| CY507 | HH5 | M13R | 40.65 |
| CY601 | HH6 | seq_A1 | 33.23 |
| CY602 | HH6 | seq_A2 | 33.23 |
| CY603 | HH6 | seq_A3 | 33.23 |
| CY604 | HH6 | seq_A4 | 33.23 |
| CY605 | HH6 | seq_A5 | 33.23 |
| CY606 | HH6 | M13F | 33.23 |
| CY607 | HH6 | M13R | 33.23 |
Transformation
Using the newly arrived Zero Blunt Topo cloning kit, we ligated, transformed, and plated the following:
- Topo + LC1PUR1 3kb segment (2 plates)
- Topo + LC1PUR1 400b segment (2 plates)
- Topo + LC2PUR1 3kb segment (2 plates)
- Topo + LC2PUR1 400b segment (2 plates)
- Topo + positive control template
- Topo + H2O (negative control)
4 µL of template and 20 µL of competent cells were used in each transformation.
Site Specific Mutagenesis Attempt 5
Peng f-ed up the primers...
Dave's template was HH5, Peng's was HH6.
Vials , 3-10, 9-8, 6-7, 5-4, -t for each, cat gene as a positive control.
For cat gene, pbc_ks+, kt_b, kt_bb primers - expected size ~1.1kb
New protocol, based on the Silver protocol on OpenWetWare:
5 µL 10x ThermoPol buffer (Vent) 1.0 µL 10 mM dNTPs 2.5 µL 20 µM forward primer 2.5 µL 20 µM reverse primer 1 µL plasmid DNA 1 µL Vent DNA polymerase 37 uL dH20
PCR Time for 3-9, 6-4, 10-7, cat gene (peng), negative for each
#*95 °C for 2 min. (melt) #*95 °C for 0.5 min (melt) #*57 °C for 0.5 min. (anneal) #*74 °C for 1.5 min. (extension) #* Go to step 2 (30x) #*74 °C for 5 min. (final extension; modified from Silver protocol) #* Keep at 4 °C forever
Total time: 82 minutes +/- some = 90 min
PCR Time for 6-7, cat gene (dave), negative for each
#*95 °C for 2 min. (melt) #*95 °C for 0.5 min (melt) #*57 °C for 0.5 min. (anneal) #*74 °C for 2.5 min. (extension) #* Go to step 2 (30x) #*74 °C for 5 min. (final extension; modified from Silver protocol) #* Keep at 4 °C forever
Total time: 111 minutes +/- some = 120min
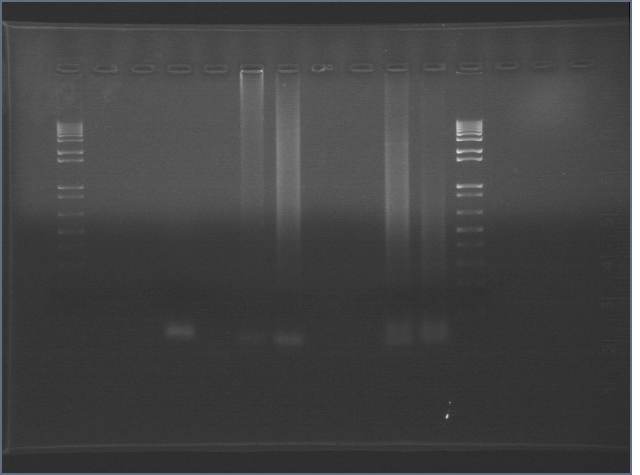
Results
Expected:
3-10: 209bp
9-8: 370bp
6-7: 2402bp
5-4: 80bp
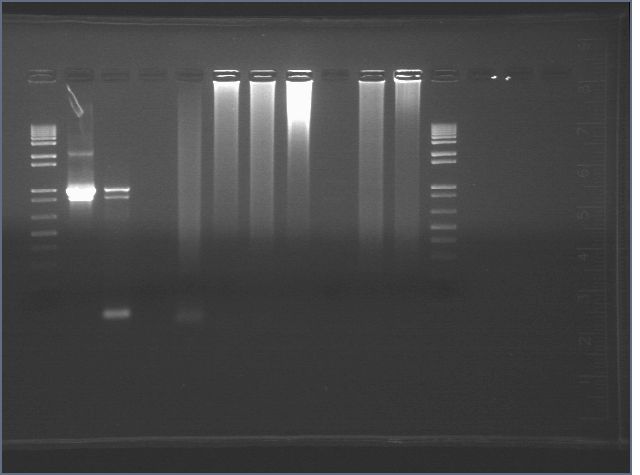
Failed... again...
1: 1 kb+ ladder 2: positive cat dr 3: negative cat dr 4: 6-7 dr 5: 6-7 dr neg 6: 6-7 zs 7: 6-7 zs neg 8: 3-10 dr 9: 3-10 neg dr 10: 3-10 zs 11: 3-10 zs neg 12: ladder
1: 1 kb+ ladder 2: positive cat zs 3: negative cat zs 4: 9-10 dr 5: 9-10 dr neg 6: 9-10 zs 7: 9-10 zs neg 8: 5-4 dr 9: 5-4 neg dr 10: 5-4 zs 11: 5-4 zs neg 12: ladder