Falghane Week 2
Week 2 Journal
Purpose
The purpose of this assignment is to learn about how DNA microarray experiments work. Also to reason through some DNA microarray analysis problems, and to discuss a case where standards of research ethics are violated.
Answers for Discovery Questions from Chapter 4
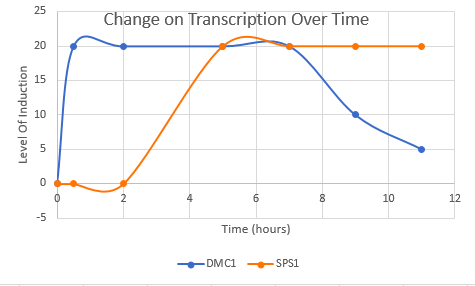
Choose two genes from Figure 4.6b (PDF of figures on Brightspace) and draw a graph to represent the change in transcription over time. You can either create your plot in Excel and put the image up on your wiki page or you can do it by hand and upload a picture or scan
Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6, determine the color for each ratio in Figure 4.7b. (Use the nomenclature "bright green", "medium green", "dim green", "black", "dim red", "medium red", or "bright red" for your answers.)
- Gene X: 1hr; black, 3hrs; dime red, 5hrs; black, 9hrs; dime green
- Gene Y: 1hr; black, 3hrs; dime red, 5hrs; dime green, 9hrs; bright green
- Gene Z: 1hr; black, 3hrs; dime red, 5hrs; dime red, 9hrs; dime red
Were any of the genes in Figure 4.7b transcribed similarly? If so, which ones were transcribed similarly to which ones?
Up to the fifth hour, all the genes were transcribed similarly as all of them were black at hour 1, and dime red at hour 3. After that, each gene started having a different color than the other ones.
Why would most spots be yellow at the first time point? I.e., what is the technical reason that spots show up as yellow - where does the yellow color come from? And, what would be the biological reason that the experiment resulted in most spots being yellow?
The combination fo cDNA labeled red for experimental expression and cDNA labeled green for control produces yellow and since there was no change from the initial or control group the yellow spot appeared.
Go to the Saccharomyces Genome Database and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s change in expression was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
over the course of this experiment, TEF4 was repressed. That is because of the lake of food due to the reduction in glucose, therefore it responded to save energy.
Why would TCA cycle genes be induced if the glucose supply is running out?
a reduction in glucose concentration results in starvation, therefore, the cell induces the energy metabolism genes and repress the production genes. The genes induced shift the pyruvate towards the TCA cycle.
What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
The Guilt by Association method could be used to ensure genes for enzymes in a common pathway are induced or repressed simultaneously.
Consider a microarray experiment where cells deleted for the repressor TUP1 were subjected to the same experiment of a time course of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later time points (glucose depleted) are labeled red. What color would you expect the spots that represented glucose-repressed genes to be in the later time points of this experiment?
I would expect the spots that represented glucose-repressed genes to be red in the later time points of this experiment. That is because there's no repression and therefore there will be more expression of the glucose-repressed genes.
Consider a microarray experiment where cells that overexpress the transcription factor Yap1p were subjected to the same experiment of a time course of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later time points (glucose depleted) are labeled red. What color would you expect the spots that represented Yap1p target genes to be in the later time points of this experiment?
I would expect the color of the spots that represented Yaplp target genes to be red.
Using the microarray data, how could you verify that you had truly deleted TUP1 or overexpressed YAP1 in the experiments described in questions 8 and 9?
To verify that TUP1 was truly deleted, the spots should be red as it indicates a change in gene expression. Similarly the overexpression of YAP1 would be shown by showing a change from control and if it wasn't overexpressed then it would look like the control group.
Acknowledgment
- I worked with my homework partner, Austin Dias over text in this project.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Falghane (talk) 17:58, 30 January 2019 (PST)
References
- Campbell, A.M. & Heyer, L.J. (2003), “Chapter 4: Basic Research with DNA Microarrays”, in Discovering Genomics, Proteomics, and Bioinformatics, Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press, pp. 107-124.
- Dahlquist, K. and Fitzpatrick, B. (2019). BIOL388/S19:Week 2. [online] openwetware.org. Available at:Week 2 Assignment Page [Accessed 30 Jan. 2019].
- News, C. (2012, February 12). Retrieved January 31, 2019, from https://www.youtube.com/watch?v=eV9dcAGaVU8.