Endy:Translation demand
Contact
Overview
Motivation
Our ability to reliably engineer biological systems is very limited. One reason for this is because we have a crude understanding of the interaction between an engineered biological system and the cell that hosts it, here called a cellular chassis. It is my goal to elucidate and quantify the interaction between engineered biological system and cellular chassis.
Demand & chassis response
An engineered biological system places many different demands on its cellular chassis. For example, an engineered biological system will compete with chassis systems for machinery such as polymerases and ribosomes and for "raw materials" such as nucleotides and amino acids. When these demands are placed on a cellular chassis, it may lead to changes in the physiology of the chassis (e.g. growth rates, protein synthesis rates etc.). I would like to examine the relationship between applied demand and chassis response.
Goal
We expect that as translation demand increases, the chassis response will become more pronounced (decreasing growth rate, triggering of stress response pathways etc.). I am interested in examining what range of translational demands can be applied to a cellular chassis before the chassis response to that demand adversely affects the performance of the engineered biological system.
You can read about the current status of this project here.
Approach
Getting at these issues requires me to achieve two intermediate goals. Firstly, I need to develop methods to place a specified range of demands on the chassis. Secondly, I need to develop relevant measures of the chassis response to that applied demand.
Placing a Range of Demands on the Chassis
In order to develop a quantitative relationship between protein synthesis rate and cellular growth rate, I have to place a range of demands on the cell. I'm employing two methods in this experiment. First, I am using both a low and high copy vector. pSB4A3, the low copy vector, produces approximately 10-12 copies for cell. The high copy vector, pSB1A3, produces 100-300 copies per cell. This is a means of roughly but dramatically increasing the demand on a cell. In order to gain more intermediate steps in demand, I am also utilizing several different ribosome binding sites chosen to represent a range of strengths. I would expect that a stronger RBS would place a greater demand on the cell. The strenth of the RBS has been determined by the ranking system established by Ron Weiss, data taken from experiment conducted with the T7 Bacteriophage, and data from Heather Keller's work.
My experiment will utilize fluoroescent proteins to measure protein synthesis rate. By measuring GFP and Mcherry Counts with respect to time and determining the slope at selected points, I can determine the net synthesis rate of protein. By then taking optical density measurements, I can determine the number of cells at these selected times and determine rate of protein synthesis per cell. I will also use the OD vs. time data to determine the growth rate of the cells in culture. These analyses will allow me to directly compare the rate of protein synthesis and growth rate for a cell.
Measuring relevant chassis responses to an applied demand
I need to determine the appropriate metrics of the chassis response to an applied demand. For example, if the application of a demand on the chassis reduces chassis growth rate, then that will affect the behavior of the engineered biological system and hence growth rate would be a relevant chassis response. Conversely, if an applied demand leads to an increase in the synthesis of some heat shock proteins but these merely serve to ensure that protein synthesis is unaffected by the increase in demand, then those heat shock proteins would not be a very relevant chassis response.
Materials & Methods
Demand Constructs
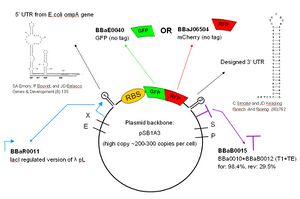
The GFP and Mcherry scaffolds I'm using were constructed by Heather Keller of the Endy Lab. They include a LacI regulated version of the lambda pL promoter (BBaR0011), two hair pins on either side of the coding sequence to increase stability, GFP (BBa0040) or Mcherry (BBaJ06504), and an RBS that I intend to vary. As I stated earlier, I'm using the Biobricks vectors pSB1A3 and pSB4A3. For those unfamiliar with Biobricks and the Registry for Standard Biological Parts, they are both very successful efforts to make biological engineering more standardized and modular by making and recording various interchangable parts to be used in the construction of DNA. It is this modularity that allows me to place Heather's scaffolds on both the high and low copy vectors and switch RBS's without having to syntheisize each construct from scratch.
Bacterial strains
My initial characterization work will be done in the E. coli strain MG1655 (No, I did not discover the strain. I just happen to share my initials with a bacterium), though DB 3.1 will be used in an intermediary step to construct the recombinant plasmids.
X90/C86
Media & growth conditions
As I build my constructs and grow cells, I'll use LB with Ampicillin for both cultures and plates. In preparation for the plate reader, however, I'll innoculate culture in Neidhardt EZ Rich Defined and induce with IPTG. The plate will have samples of each RBS on the high and low copy vectors, and as controls I will have "empty" vectors (just the vector without the scaffold), RBSes constructed to be insignificantly weak, the untransformed strain MG1655, and blanks of the EZ media with both Ampicillin and IPTG.
Future work
The major thrust of this project is to better characterize the relationship between recombinant load and growth rate and eventually tease out the point at which the strain MG1655 takes a significant growth hit. A natural progression of this project would be to examine the variables that determine this point. Perhaps MG1655 and a host of other strains could be "refactored" simliar to T7 Rebuilding T7 for optimum production capacity and reliable function.
Data
All raw data and data processing scripts can be found here.