Chris Rhodes Week 5
Methods
- To create ClustalW alignments and phylogenetic trees for the chosen subjects(3,7,8,11,12, and 13) each subject's sequence datawas uploaded onto Workbench each as a separate session.
- For each subject's clones a ClustalW alignment was run using default parameters to generate both the rooted and unrooted phylogenetic trees. Using the alignments the S value was also calculated. This S value was normalized across the subjects by correcting for the number of clones of each subject (c) and the number of visits of each subject (v) using the equation Adjusted S= (S/cv). The values for each Subject are given in the table below:
Subject # S Adjusted S 3 42 0.216 7 54 0.252 8 68 0.199 11 34 0.385 12 42 0.191 13 25 0.193
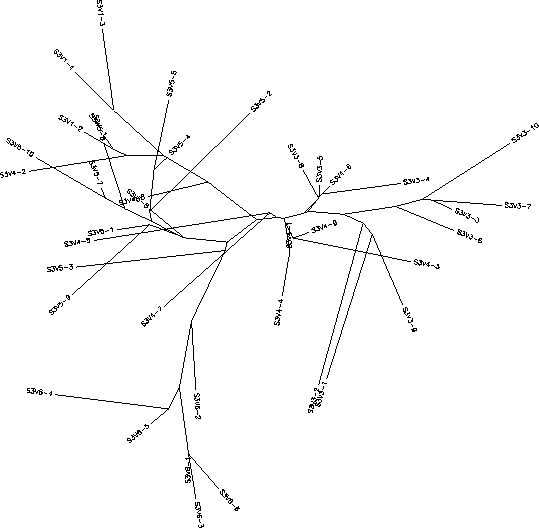
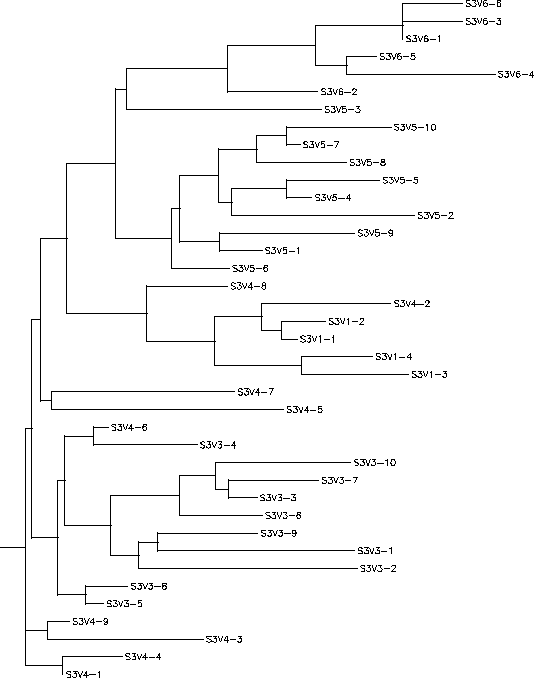
Subject 3
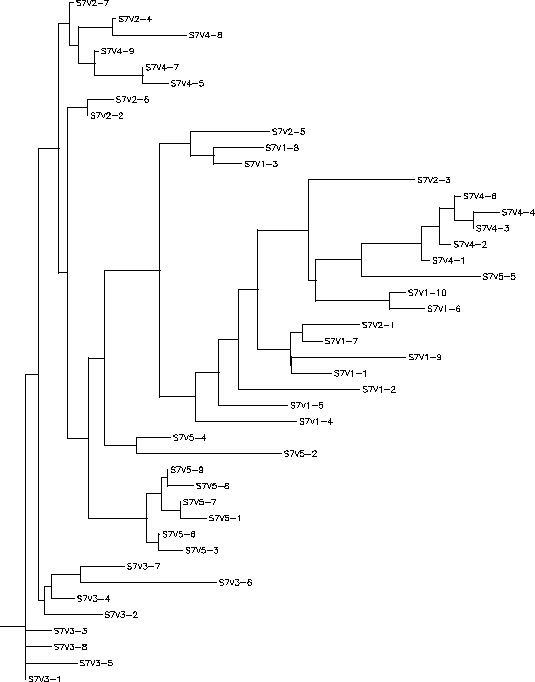
The S value for Subject 3 was 25, the adjusted S value was .216. Based on both the rooted and unrooted trees you can see that Subject 3 had very high diversity and divergence. The rooted tree shows a large vertical distance between its two ends and a large amount of branching with no branch being prominent. The clones seem to cluster together based on their visit number to create the various branches and show little similarity with clones outside of their own visit. The long branches between the clones show an amount of intravisit diversity as well. The only exceptions to this pattern are some of the clones of visit 4 who share greater similarities to visit 1 or 3 clones rather than with other visit 4 clones. However, this overall pattern is indicative of an immune response in which there is high selection happening against a lack of change in the amino acid sequence which results in large divergence between every visit.
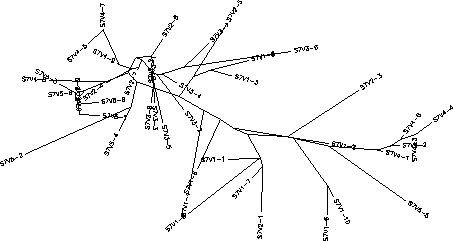
Subject 7
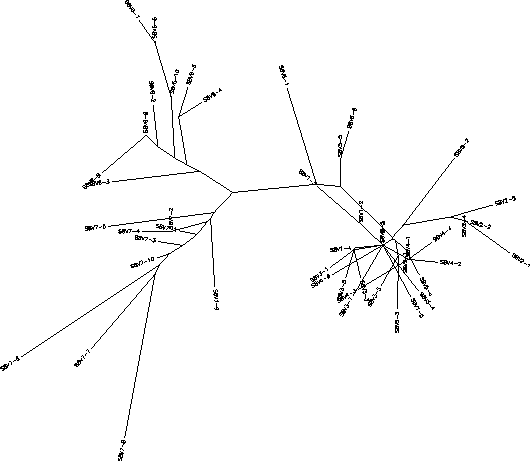
The S value for Subject 7 was 54, the adjusted S value was .252. Based on the rooted tree Subject 7 has a very large amount of genetic diversity between its clones and its and its visits. Like Subject 3, Subject 7's clones show a good amount of clustering based on visit number however unlike Subject 3, Subject 7 shows distinct intravisits clusters meaning that there is more than 1 cluster per visit. Subsequently Subject 7 has a significant amount of intravisit diversity especially within visits 2 and 4. Subject 7 also shows a horizontally prominent branch which contains clones from visits 1, 2, and 4. Visit 3 represents a large divergence from visits 1 and 2 but does not have any subsequent visit clones on its branch indicating that the visit 3 strains may developed a mutation that increased replication resulting in a high concentration of the visit 3 strains, which were then severely cut down by the immune system. A best fit theory, where the most well-suited virus dominates and there is strong selection against change, would explain the initial diversity of visits 1 and 2 along with the apparent decline of diversity as the virus progresses. This theory of a best fit virus is also supported by the large divergence and sudden disappearance of the visit 3 strains. It seems that while there is selection against change it's degree varies with the amount of change occurring.The immune system allows for diversity within a certain genetic proximity to the best fit sequences but selects strongly against drastically different mutations. For example, visit 3 clones showed a drastic change from visits 1 and 2 and were quickly snuffed out were as visits 2 and 4 clones which change less drastically from the best fit sequence are able to survive. Visit 5 however seems to behave similarly to visit 3 in that it clusters and diverges greatly from visit 4, which mainly represents the two best fit sequences. Had there been a visit 6 I suspect that the visit 5 strains would be cleared by the immune system and visit 6 clones would represent the progression of the 2 best fit branches being more similar to visit 2 and 4 clones.
Subject 8
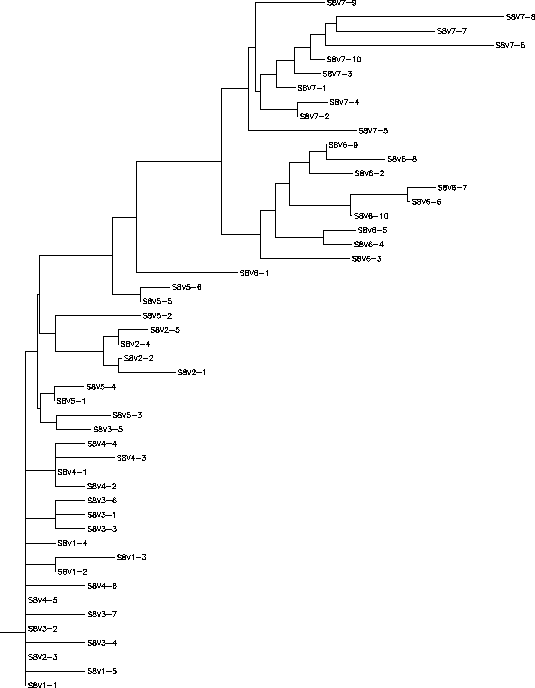
The S value for Subject 8 was 68, the adjusted S value was .199. Based on the rooted and unrooted trees Subject 8 has an overall large amount of diversity and divergence but the rates of diversity and divergence change overtime. There is little intravisit clustering for visits 1-5 with the exception of visit 2, however visits 6 and 7 cluster together based on their visits forming two horizontally prominent branches. From visits 1-5 there seemed to be a fair amount of diversity but little divergence between the different clones and visits, however visits 6 and 7 represent a very large jump in diversity and divergence from the visits 1-5 clones. Visit 1-5 clones also show little branching with the few branches that do exist being short while visit 6 and 7 clones show more frequent and more elongated branches. Visit 7 clones especially have long branches showing an amount of intravisit diversity. Based on these factors it can be hypothesized that the immune system of Subject 8 was originally not providing any selective pressure, resulting in the indiscriminate diversity of visits 1-5, but then suddenly between visits 5 and 6 began selecting for change. This might be explained by an immune system that was unable to detect the protein coats of anything functionally similar to visits 1-5, but which between visits 5 and 6 developed a way of recognizing and destroying proteins coats functional similar to visits 1-5. In this way, the immune system shifts from selecting against change to selecting for change and explains how Subject 8's tree may have developed.
Subject 11
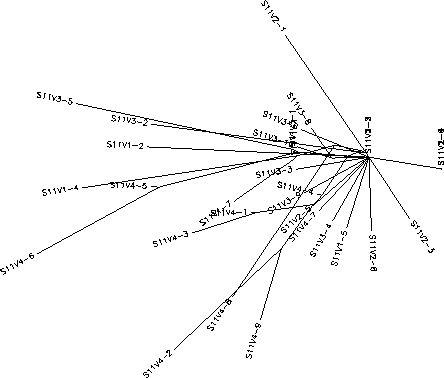
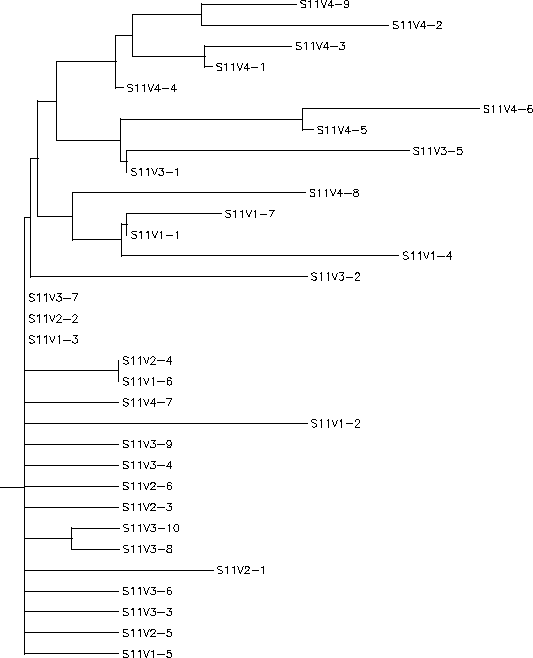
The S value for Subject 11 was 34, the adjusted S value was .385. Based on the rooted and unrooted trees Subject 11 shows a fairly large amount of diversity and divergence. Unlike Subject 8, Subject 11 shows a consistent and large rate of diversity throughout the experiment. There is little clustering based on visit and clones of various visits seem to sort themselves fairly randomly across the tree with visit 1, 2, and 3 clones often intermingling on branches while usually being very far separated from other clones from the same visit. Subject 11 also contains a large amount of individual branches and long chained branches indicating a very diverse population of both intra- and intervisit clones. Visit 4 represents the most divergent population having it's own distinct branch which contains a widespread cluster of about half of the visit 4 population. This type of distinct branching would seem to indicate that this visit 4 strain is being selected for, but upon seeing the positions of the remaining visit 4 clones this doesn't appear to be the case. Although it does seems there was a certain amount of selection for change as the virus progressed, based on the data that 4 out of 9 of the visit 4 clones were variants of visit 1 and 3 clones and diverged highly from the distinct visit 4 branch this selection appears fairly weak. Overall this pattern of high and seemingly indiscriminate diversity indicates an immune system that contains no selection for or against change. This lack of selection is most likely due to an immune response that is either ineffective against the virus or population-specific to one particular strain, assumedly wiped out before the first visit, allowing all other strains to grow without resistance or selection.
Subject 12
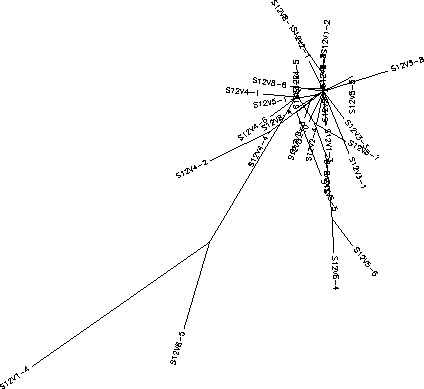
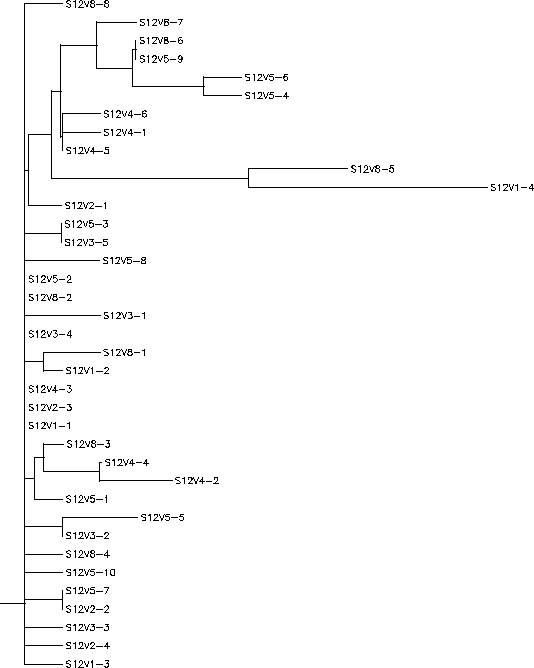
The S value for Subject 12 was 42, the adjusted S value was .19. Based on the rooted and unrooted tree Subject 12 shows a small amount of diversity and little divergence. As with Subject 11, Subject 12 shows little clustering and a large amount of intravisit diversity with clones from each visit being widely dispersed throughout the tree and more similar to clones from other visits than clones in their own visit. While there is a fair amount of intravisit diversity, intervisit diversity is quite low even when comparing over long stretches of time, the most pronounced example being the immense similarity between clones V1-2 and V8-1. This and other similar examples show that diversity is not increasing greatly overtime. Subject 12 contains minimal branching from the root and short horizontal branches indicating high similarity between neighboring clones with 2 notable exceptions. V8-5 and V1-4 are highly divergent from the root and exist on extremely long branches. Due to the relative similarity between V8-5 and V1-4 it could be hypothesized that V8-5 is a much later descendent V1-4 meaning that while it is evident that V1-4 was not favored under selection it was also not completely eradicated and continued to replicate though at a severely diminished population. This, along with the small intervisit diversity and low rate of divergence among the clones points towards an immune system that selected highly against change. As detailed by the Markham paper such an immune response would consist of a broadly effective and non-specific immune system which would cut down any strain whose population went beyond a certain threshold. This means that escape mutations would not be selected for and rapid replication mutations would be harshly selected against creating an environment where only variants similar enough to those already surviving would survive.
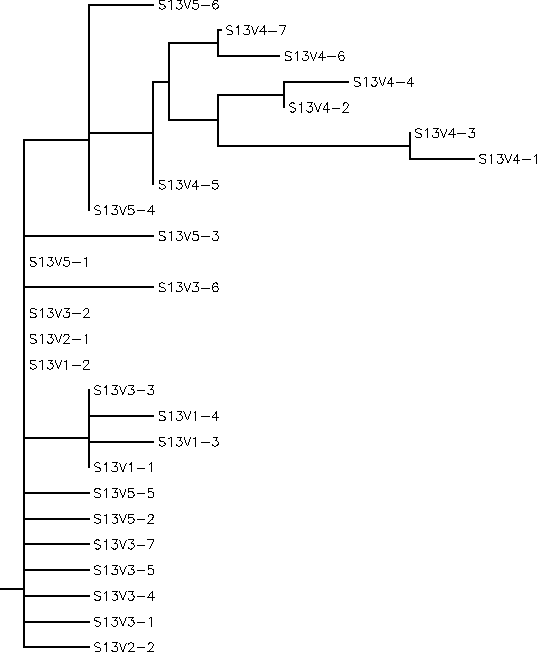
Subject 13
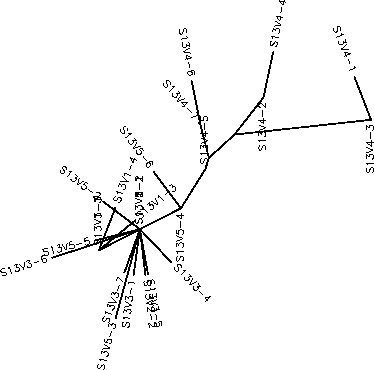
The S value for Subject 13 was 25, the adjusted S value was .193. Based on the unrooted and rooted tree Subject 13 shows very little diversity or divergence. Subject 13 is very similar to Subject 12 in most of it's characteristics. It has a small amount of intravisit clustering, a fair amount of intravisit diversity, a low amount of intervisit diversity, contains minimal branching, and horizontally short branches. Notably, visit 4 clones seem to contradict these patterns having a long distinct branch from the root made up of a long branched cluster of all the visit 4 clones. Since visit 4 represents a very divergent cluster and is a later visit it is interesting that these clones have no descendant strains that diverge even further from the root as one would expect if there was sudden selection for change as in Subject 8. Instead the closest possible descendent strains, V5-6 and V5-4, are actually more similar to the root sequences than any of the visit 4 clones are to the root sequences. It would appear then, that between visits 4 and 5 the most divergent visit 4 clones were cut down by the immune system while any surviving variants may have undergone mutations that "reverted" their sequence to be more similar to the root sequences. This would mean that there is in fact a considerable selection against change in Subject 13 and explain why we see V5-6 and V5-4 phylogentically closer to the root sequences than are the visit 4 clones. As in Subject 12, such an immune response would consist of a broadly effective and non-specific immune system which would cut down any strain whose population went beyond a certain threshold. Therefore, strongly selecting against change that would cause a rapid increase in population such as the visit 4 clones could have possessed.
Reference Paper
Important Conclusions
- Deceleration of HIV sequence evolution occurs in response to decreased immune selection -> Subject 8's increase in diversity could be caused by the opposite phenomenon, where sudden increased selection leads to the jump in evolution rates.
- When the diversifying selection pressure on Env from the immune system weakens new escape mutations are not beneficial, and thus the probability to establish new mutations decreases. -> Subject 7's pattern of genetic progression would seem to indicate a similar theme due to the fact that radically different strains from the seemingly "best fit" strains were quickly cut down and could explain why this occurred.
- One of the selection forces controlling the evolution of env is escape from antibody neutralization -> Subject 11's tree shows data that would seem to indicate a lack of selective pressure. If this is the case, it can be assumed that for whatever reason Subject 11's immune system is not providing the threat of virus neutralizing antibodies.
- Negative rate of evolution, evolution towards the root, may be due to the emergence of less evolved strains from latent reservoirs at later sampling time points. -> Subject 13 contains variants from visit 5 who, though are more similar to the visit 4 branch, are closer to the root than the visit 4 clones. As this paper describes, this negative evolution could be due to an emergence of older variants in later visits. In this case these visit 5 clones could represent the more root similar genetic precursor to the far more divergent visit 4 clones explaining their survival and reemergence.
Presentation Link
Chris Rhodes, Nicki Harmon HIV evolution Presentation
Links
- Chris Rhodes User Page
- Week 2 Journal
- Week 3 Journal
- Week 4 Journal
- Week 5 Journal
- Week 6 Journal
- Week 7 Journal
- Week 8 Journal
- Week 9 Journal
- Week 10 Journal
- Week 11 Journal
- Week 12 Journal
- Week 13 Journal
- Week 14 Journal
- Home Page
- Week 5 Assignment Page
- Week 6 Assignment Page
- Week 7 Assignment Page
- Week 8 Assignment Page
- Week 9 Assignment Page
- Week 10 Assignment Page
- Week 11 Assignment Page
- Week 12 Assignment Page
- Week 13 Assignment Page
- Week 14 Assignment Page