Biomod/2015/StJohns
http://openwetware.org/images/f/f9/Lukemanlab-toehold-conga-nanny-header.jpg
Introduction and Theory: DNA Nanotechnology
Watson and Crick Base pairing allows for a nucleotide sequence to assemble in a predictable manner. Used in combination with the DNA double crossover concepti[1], large macromolecular structures can be assembled into a variety of different patterns and shapesii[2]. All assemblies combine in aqueous solution, and rely on the strands to assemble into their thermodynamically most stable configuration.
Having such high specificity, DNA nanotechnologists are now able to assemble DNA origami (DO) structures to have a set function. This lab sets out to design a DO Claw that would act as a virus sensor allowing for efficient and rapid detection, with a potential future application in diagnostics.
Claw Project
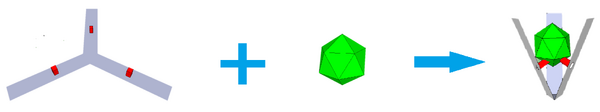
Goal: Design and synthesize a DO Claw which is able to efficiently bind a virus, while undergoing a conformational change.
The DO Claw (Figure 1A) was assembled using a long single strand of DNA (plasmid) and woven into the desired shape, held together by 226 staple strands which are complementary to only very specific areas of the plasmid.
There have been several alterations done to the claw from the original design, which is referred to as Vanilla Claw. New Claw (NClaw) was made in order to test FRET signals upon change in conformation. When successful binding was not demonstrated, Tether Claw (TClaw) was designed to limit the mobility of the three arms of the claw.
FAB'
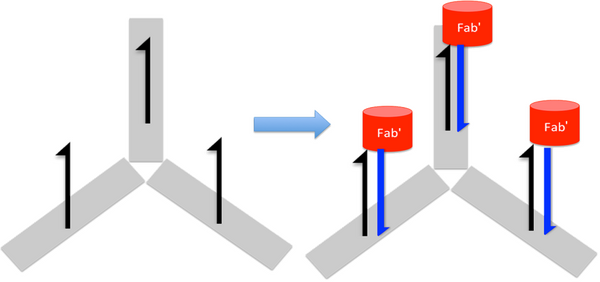
Goal: To modify the DNA claw in such a way that there would be binding sites for the wild-type MS2 virus capsids to bind to.
A Fab’ -DNA conjugate attached to the claw to bind a virus surface was our approach. Fab’ is made from antibodies and has a free, exposed thiol group for conjugation; many virus antibodies are available; thus it should be possible to make these conjugates for a wide variety of virus surfaces.
Near and Future goals
Achieved
<html>
<input type="checkbox" checked="yes"/> Synthesize NClaw
<input type="checkbox" checked="yes"/> Incorporate FRET dyes into NClaw, in appropriate regions
<input type="checkbox" checked="yes"/> Obtain AFM images of NClaw
<input type="checkbox" checked="yes"/> Attach multiple binding sites to claw to increase binding avidity for capsid
<input type="checkbox" checked="yes"/> Modify NClaw to make TClaw (tethered), limiting free mobility of arms
<input type="checkbox" checked="yes"/> Confirm tethers are binding to TClaw via fluorescent strands
<input type="checkbox" checked="yes"/> Conduct control experiment using DO Triangle and MS2 modified capsid
<input type="checkbox" checked="yes"/> Synthesize Fab’
<input type="checkbox" checked="yes"/> Conjugate Fab’ to Maleimide-DNA
<input type="checkbox"/> Illustrate Claw + MS2 modified capsid gel shift, signifying conformational change
<input type="checkbox"/> Demonstrate Claw + MS2 modified capsid binding via TEM imaging
<input type="checkbox"/> Demonstrate Claw + MS2 modified capsid binding via FRET signals
<input type="checkbox"/> Demonstrate Aptamer binding capabilities
<input type="checkbox"/> Obtain Claw + Aptamer conjugate
</html>