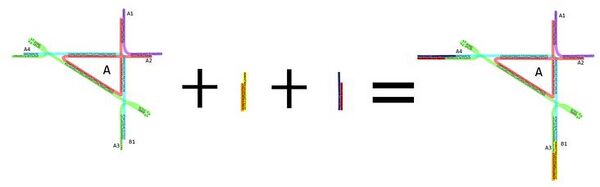Biomod/2013/StJohns/approaches
<html>
<head>
<link href='http://fonts.googleapis.com/css?family=Open+Sans' rel='stylesheet' type='text/css'>
</head>
<style>
body {
font-family: 'Trebuchet MS', 'Open Sans', sans-serif; overflow-y: auto;
}
.container {
background-color: #ffffff; margin-top:0px
} .OWWNBcpCurrentDateFilled { display: none; }
h5 {
font-family: 'Trebuchet MS', 'Open Sans', sans-serif; font-size: 11px; font-style: normal; text-align: center; margin:0px; padding:0px;
}
- column-content
{
width: 0px; float: left; margin: 0 0 0 0; padding: 0;
} .firstHeading {
display:none; width:0px;
}
- column-one
{
display:none; width:0px; background-color: #ffffff;
}
- globalWrapper
{
width: 1024px; background-color: #ffffff; margin-left: auto; margin-right: auto
}
- content
{
margin: 0 0 0 0; align: center; padding: 12px 12px 12px 12px; width: 1000px; background-color: #ffffff; border: 0;
}
- bodyContent
{
width: 974px; align: justify; background-color: #fffffff;
}
- column-content
{
width: 1024px; background-color: #ffffff;
}
- footer
{
position: absolute; bottom: 0; width: 1024px;
}
- menu
{
position: fixed; float: left; width: 180px; padding: 10px; background-color: #FFFFFF;
}
- resolutionwarning
{
display: none; text-align: center;
}
- pagecontent
{
text-align: justify; float: right; width: 744px; margin-left: 300px; min-height: 400px
}
- toc { display: none; }
@media all {
body { background: #ffffff 0 0 no-repeat;}
}
/*Expanding list*/ ul { list-style: none; }
- exp li ul { display: none; }
- exp li:hover ul { display: block; }
- exp li a:active ul { display: block; }
a:link {color:#007788;} /*unvisited link*/ a:visited {color:#007788;} /* visited link */ a:hover {color:#00ccff;} /* mouse over link */ a:active {color:#00ccff;} /* selected link */
</style> </html>
<html><style>
- exp li ul { display: inherit; }
</style></html>
Claw
The central aspect of the project is, as described, the synthesis of a DO claw capable of binding with high affinity to specific target substrates and displaying a significant conformational change in response to this binding. Our initial approach uses a modified bacteriophage MS2 capsid as the target substrate, with complementary pieces of ssDNA attached to the claw and capsid. Thus, binding is driven by Watson-Crick base-pairing and the hydrophobic effect, the same effects that allow DNA origami to be reliably designed in the first place.
Specifically, we aim to synthesize an unmodified claw structure, a modified version with the ssDNA binding elements ("sticky" claw), modified versions with fluorescent tags attached at particular points on the structure for FRET analysis, and combinations of these two modifications. Further, we wish to characterize each of these structures by gel electrophoresis, atomic force microscopy, dynamic light scattering, and FRET analysis to demonstrate the expected shift in size and shape caused by the binding interaction. Lastly, we wish to find binding conditions which optimize for strong binding between the functionalized capsid and origami with minimal nonspecific binding between other elements.
Selection
Our eventual goal regarding selection is to create “libraries” of virus-binding claws that have different geometries of virus-binding sites, selecting the most avid one for further study. However, the very minute gradations between each claw in this “library” makes selecting between them extremely complicated, so to ensure the best eventual results we are first attempting to perfect our selection protocol.
To this end, we have constructed a model selection using two types of origami that are different to a much greater extent than the claws in our “library” will be from each other. In this approach, we functionalize agarose resin with the same strands displayed on modified MS2 capsids. We then create a selection mixture containing both our claw of interest—nicknamed “sticky claw” for the strands it displays—and another type of origami nicknamed “blunt triangle” for its triangular shape and its lack of complementary strands. Our current goal is to select the sticky claw alone from this mixture.
The strands on the sticky claw are complementary to the strands displayed by the agarose resin, subsequently sticking to the resin. The blunt triangle, however, lacks those complementary strands and is washed out during a series of buffer washes. The linker that attaches the strands to the resin is photocleavable, so after selection ultraviolet light is used to release the active claw.
The washes are then visualized through gel electrophoresis, allowing us to determine whether the blunt triangle was removed in our initial resin washes and our sticky claw removed from the functionalized resin after our ultraviolet resin cleave.
Antibodies
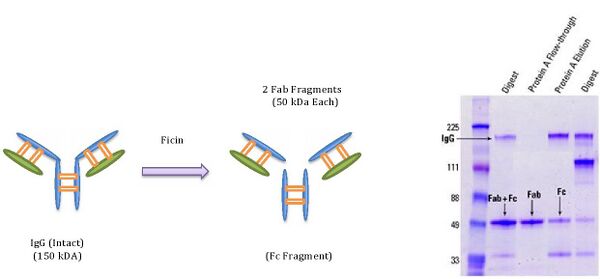
In this approach, the goal is to generate small anti-Flag Fab fragments for potential use as binding elements in future claw designs (visualized as small red protrusions on our models of the claw). These anti-Flag Fab fragments are generated from a larger antibody protein molecule called Immunoglobulin G (IgG).
As can be seen in the figure below, the IgG can be cleaved by an enzyme called ficin. The result yields two Fab fragments per IgG molecule. The sample gel on the right displays the relative mobilities and quantities of these fragments and IgG. Notice that the IgG shows up at around 150 kDa and Fab fragments show up at around 50 kDa.
Ultimately these Fab fragments are meant to be incorporated into the claw's structure. Before we get to that step, we need to make sure that we can attach a DNA strand to the Fab fragment itself. In turn, before working with Fab fragments, we want a model for the conjugation reaction. Preliminary work is being done on attaching a piece of DNA to a ketone molecule, which serves as our model. This small step involves sticking a strand of DNA that is about 20 based long (TCA and TCAT3) onto the ketone molecule. The two conjugated ketone molecules that were studied are shown:
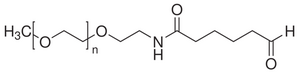
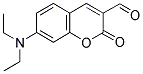
The overall scheme is as follows:
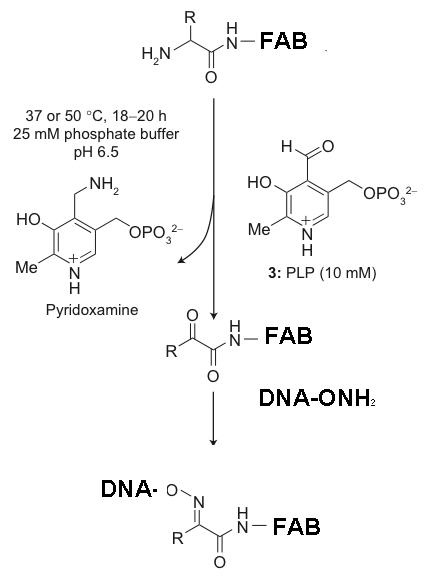
We are testing the DNA-ONH2's ability to cojugate to ketones before using the expensive Fab fragment for the real conjugation reaction
Triangle Tiles
- Note: These small, 10nm triangles are different and unrelated to the control triangles used in the selection approach.
The goal of this aspect project was to create precise DNA right triangles that could potentially serve as hinges for other DNA origami, more specifically DNA claws. These origami complexes can be utilized as hinges through their rigidity. We were successful in making these triangles by combining strands and their complementary strands in solution. After running several gels, we concluded that the solution that we had originally made was a DNA complex with specific configuration by the solid band that it expressed. However, we realized that the triangles were too small to image by themselves by AFM.
This is when we realized that we needed to add DNA to the ends to make them visible. We utilized the triangle’s sticky ends to attach DNA duplexes that would help to make them visible on AFM. We ran gels to see if a new complex was made compared to the triangle. We also added a DNA complex that included the triangle with each group of duplexes on the gel, and managed to successfully show that these different complexes all moved at different mobilities. Further analysis by AFM showed that these complexes with both extenders formed at specific angles. Although several of the angles were not the desired value (90 degrees), many of the angles of the complexes that we measured were precise.
Currently, we are working on creating and imaging origami triangles with modified ‘extenders’. We are conforming plasmid DNA into the modified ‘extenders’ and utilizing staple strands to hold the plasmid DNA in place. The triangle will be made of origami strands and their complementary strands. We expect that these modified extenders will allow us to easily see the angle that is made when the triangle forms.