Biomod/2011/TeamJapan/Tokyo/Project/Results of DNA Ciliate Body
<html>
<style type="text/css">
/* ====================
主に全体に関わるCSS
==================== */
body {behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc);}
.clear {clear:both;}
body.mediawiki {
font-size: 14px;
background-color:#707070; background-position: center center; background-attachment: fixed; background-repeat: no-repeat; font-family: Calibri, Verdana, helvetica, sans-serif; }
h1 {
padding: 0px 20px 5px 20px;
font-size: 34px;
font-weight: bold;
}
h2 {
padding: 20px 20px 5px 20px;
font-size: 25px;
color: #0083eb;
text-decoration: none;
font-weight: bold;}
border: none;
h2 a {
color: #eb8300;
}
h3 {
padding: 20px 20px 5px 20px;
font-size: 20px;
color: #000;
font-decoration: none;
font-weight: bold;
}
h1.firstHeading {
display: none;
}
p { text-align: justify; } a:link { color: #00a5ea; text-decoration: none } a:visited { color:#00a5ea; text-decoration: none } a:hover {
color: #eb8300;
text-decoration: none } a:active { color:#f29400; text-decoration: none } #bodyContent {
width: 970px;
margin: 0px 0px 0px;
background-color:#ffffff;
border-width: 0px 1px 0px 1px;
border-color: #000000;
}
#content {
padding-left: 0px; width: 970px;
}
table#team_members {
text-align: justify;
border: 0;
}
table#team_members h2, table#team_members h3 {
clear: both;
}
#content * a:hover {
text-decoration:none;
}
#main_wrapper {
position:absolute;
left:0px;
top:20px;
margin-top: 0; width: 969px; height: 221px; align: center; border-style: solid;
border-color: white;
} /* ====================
メニューの画像を変更できる部分 ==================== */
#header {
position:relative;
left:0px;
top:0px;
margin-top: 0; width: 969px; height: 221px; align: left; background-color: #FFFFFF;
background-image: url(http://openwetware.org/images/e/e0/NEW_header.jpg);
} /* ====================
以下、特殊なclassに適用される ==================== */
#navigation { position:absolute;
left:18px; top:155px; width:1200px; height:69px;
z-index:100;
background-color: transparent;
float: left;
color: #0000FF; } #super_main_wrapper {
position:absolute;
left:0px;
top:227px;
width: 975px; align: center; background-color: #ede8e2;
heigth: auto;
}
#SubWrapper {
width: 645px;
padding: 0px;
border-left:4px solid #ede8e2;
float: left;
margin-top: 0px;
background-color: #ede8e2;
}
#SubWrapper * p, #SubWrapper p {
padding: 0 20px;
text-align: justify;
font-size: 12px;
}
#SubWrapper * h3, #SubWrapper h3 {
padding-top: 10px;
font-size: 18px;
}
#news {
width: 322px;
margin-top: 0px;
float: left;
background-color: #d8d5d0;
border-right:4px solid #ede8e2;
}
#news p {
padding: 0 20px 20px 20px;
text-align: justify;
font-size: 12px;
}
#news h3 {
padding: 10px 20px;
font-size: 18px;
}
#mission_box {
width:650px;
float: left;
}
#team_box, #heartbeat_box, #notebook_box, #parts_box, #gallery_box, #sponsors_box, .boxy {
width:215px;
float: left;
padding: 10px 0 0 0;
}
div.tleft {
border-width: 0px;
margin:0;
padding:0;
border-color:transparent;
}
/* ====================
ここからプルダウン周辺のデザイン ==================== */
- menu * {
margin: 0; padding: 0; }
- menu {
behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc); font-family: calibri, verdana, sans-serif;
font-color: #ffffff;
font-size: 19px; background-color: transparent; float:left; padding: 12px 0 0 0; }
- menu ul {
float: left; list-style: none; }
- menu ul li {
background-color:transparent;
position:relative;
float:left; list-style: none; padding: 10px 20px 0 0;
font-weight: bold;
width: auto;
}
- menu a {
color: #FFFFFF; display: inline; text-decoration: none; }
- menu a:visited {
color:#FFFFFF; text-decoration: none }
- menu a:hover {
color: #00a5ea; }
- menu ul li ul {
display: none; position: absolute; left: 0px;
width: 155px;
heigth: 1%;
font-size: 19px; opacity: 0.8; list-style: none;
top: 30px;
padding-top: 20px;
z-index:500;
}
- menu ul li:hover ul {
display: inline;
background-position: bottom;
}
- menu ul li ul li {
width: 100%; list-style: none;
background-color: #000;
margin: -1px;
padding: 0px 0 0 5px;
display: inline;
}
</style>
<script type="text/javascript">
/***********************************************
- CSS Vertical List Menu- by JavaScript Kit (www.javascriptkit.com)
- Menu interface credits: http://www.dynamicdrive.com/style/csslibrary/item/glossy-vertical-menu/
- This notice must stay intact for usage
- Visit JavaScript Kit at http://www.javascriptkit.com/ for this script and 100s more
- /
var menuids=new Array("verticalmenu") //Enter id(s) of UL menus, separated by commas var submenuoffset=-2 //Offset of submenus from main menu. Default is -2 pixels.
function createcssmenu(){ for (var i=0; i<menuids.length; i++){
var ultags=document.getElementById(menuids[i]).getElementsByTagName("ul")
for (var t=0; t<ultags.length; t++){
var spanref=document.createElement("span")
spanref.className="arrowdiv" spanref.innerHTML=" " ultags[t].parentNode.getElementsByTagName("a")[0].appendChild(spanref)
ultags[t].parentNode.onmouseover=function(){
this.getElementsByTagName("ul")[0].style.left=this.parentNode.offsetWidth+submenuoffset+"px"
this.getElementsByTagName("ul")[0].style.display="block"
}
ultags[t].parentNode.onmouseout=function(){
this.getElementsByTagName("ul")[0].style.display="none"
}
}
}
}
if (window.addEventListener)
window.addEventListener("load", createcssmenu, false)
else if (window.attachEvent)
window.attachEvent("onload", createcssmenu)
</script>
</html>
Results
DNA ciliate body

- As already mentioned in the project page, we attached DNA strands to polystyrene beads to make DNA ciliate.
- We use two methods to attach DNA to polystyrene beads. In both methods, we bind polystyrene beads' carboxylic acid to amino group of aminated DNA. In first method, we apply reference [1]. In this method, we used linker between polystyrene beads and aminated DNAs. The linker has amino group and carboxylic acid. The linker’s amino group combines with carboxylic acid of polystyrene beads and the linker’s carboxyric acid combines with amino group of aminated DNA, so polystyrene beads combine DNA through the linker. In second method, we use NHS and EDC to alter carboxylic acid to NHS. NHS has very high reactivity, and DNA's amino group reacts with NHS of porystyrene beads. The DNA ciliate body is developed in this process. :
As a motor of DNA ciliate, we used deoxyribozyme which is the enzyme comprised of DNA. Deoxyribozyme cleaves its substrate at an RNA base, if there are 2+ metal ions. Using this reaction, DNA ciliate can move.
Principle and methods
- Two experiments were needed to complete developing DNA ciliate body.
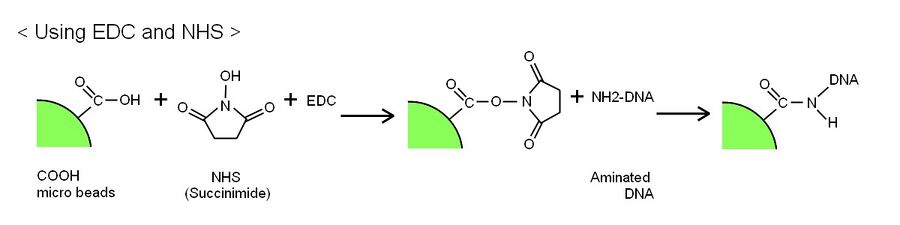
First experiment was creating DNA ciliate by attaching DNAs to polystyrene beads. This process is used the reaction of connecting aminated DNAs’ amino group and polystyrene beads’ carboxylic acid. We took two methods to react. Both methods are used the common reaction, but chemical materials are different. First method is used EDC and NHS. This induces transforming carboxylic acid to succinimide which is been able to react with aminated DNAs and connect.
- Second method is used EDAC. EDAC reacts with both aminated DNAs and polystyrene beads’ carboxylic acid.
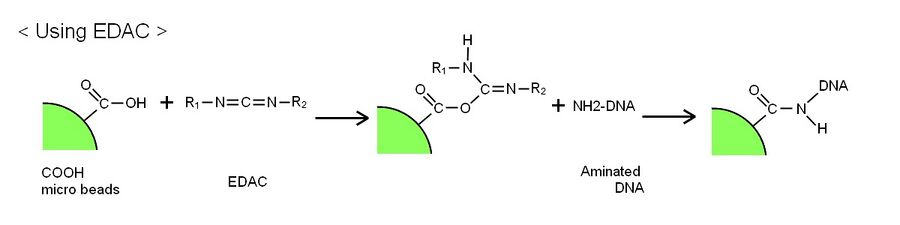
Second experiment is confirming whether deoxyribozyme is attached to polystyrene beads and able to cleave substrate. We confirm deoxyribozyme activity by urea-PAGE. Making mixture of DNA ciliate and substrate and Zn2+ ions. If DNA ciliate has deoxyribozyme activity, substrate is cleaved and the band of cleaved substrate appears as a band.
Protocol
- Creating DNA ciliate, we use (1) and (2) protocols.
- (1) The method of using EDC and NHS is here.
- (2) The method of using EDAC…
- Confirming DNA ciliate, we use (3) and (4) protocols
- (3) The method of electrophoresis is here.
- (4) The method of making sample is
Results
- Following is the result of electrophoresis. Result 1 is DNA ciliate using NHS and EDC. Result 2 is DNA ciliate using EDAC. Meanings of the numbers above results are here.
- deoxyribozyme
- substrate
- negative control (Zn(2+) 0 mM)
- positive control (Zn(2+) 10 mM)
- no deoxyribozyme beads (Zn(2+) 0 mM)
- no deoxyribozyme beads (Zn(2+) 10 mM)
- DNA ciliate (Zn(2+) 0 mM)
- DNA ciliate (Zn(2+) 10 mM)
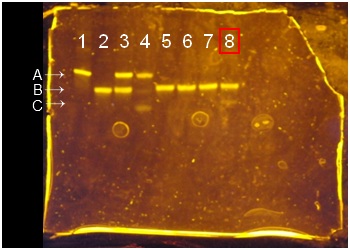
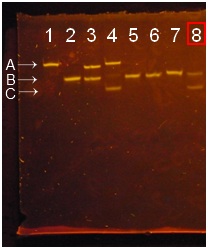
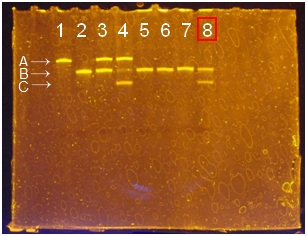
- Three bands were there. From a top, A is the band of deoxyribozyme, B is the band of substrate (didn’t be cleaved), and C is the band of cleaved substrate. If there is cleaved substrate band, it means deoxyribozyme activity is appeared and deoxyribozyme is attached to polystyrene beads successfully.
[Result 1]
- PAGE of φ200 nm polystyrene beads using NHS and EDC.
- PAGE of φ1 um polystyrene beads using NHS and EDC.
[Result 2]
- PAGE of φ200 nm polystyrene beads using EDAC.
Discussions
- First, we explain this experimentation’s appropriateness for checking DNA ciliate body. To create DNA ciliate body, it was necessary to attach deoxyrobozyme to micrometer-sized polystyrene beads firmly. Furthermore, we had to check deoxyribozyme activity of DNA ciliate body. In this method, these two things could be checked. First, the degree of fixation can be checked. DNA ciliate is removed before loading to polyacrylamide gel, so if deoxyribozyme can’t be attached to polystyrene beads firmly, the leg band is appeared. Second, the deoxyribozyme activity of DNA ciliate body can be confirmed. If there is deoxyribozyme activity of DNA ciliate body, the cleaved substrate band is appeared. Based on the above, this method is appropriate for checking DNA ciliate body.
- Second, we explain necessity of lanes. Lanes of 1 to 4 are needed for checking deoxyribozyme activity and the positions of each band of DNAs. Lane 1 and lane 2 are control lanes. The band in lane 1 is deoxyribozyme band, and the band of lane 2 is substrate band. Lane 3 and lane 4 are lanes to check deoxyribozyme activity. The solutions in lane 3 doesn’t contain of Zn(2+), the solution in lane 4 contains of Zn(2+). The cleaved band is appeared in lane 4, but doesn’t be appeared in lane 3. This means normal deoxyribozyme isn’t active in no metal ions solutions. Lane 4’s bottom band means position of cleaved substrate.
- Lanes of 5 to 8 are needed for checking deoxyribozyme activity of DNA ciliate body. Lane 5 and 6 are lanes for checking to polystyrene beads. If polystyrene beads had deoxyribozyme activity, the cleaved band would be appeared. Lanes of 7 and 8 are needed for checking DNA ciliate body’s deoxyribozyme activity. If DNA ciliate has normal deoxyribozyme activity, the cleaved band is appeared in lane 8 because metal ions are needed for deoxyribribozyme activity.
DNA ciliate body

- As already mentioned in the project page, we attached DNA strands to polystyrene beads to make DNA ciliate.
- We use two methods to attach DNA to polystyrene beads. In both methods, we bind polystyrene beads' carboxylic acid to amino group of aminated DNA. In first method, we apply reference [1]. In this method, we used linker between polystyrene beads and aminated DNAs. The linker has amino group and carboxylic acid. The linker’s amino group combines with carboxylic acid of polystyrene beads and the linker’s carboxyric acid combines with amino group of aminated DNA, so polystyrene beads combine DNA through the linker. In second method, we use NHS and EDC to alter carboxylic acid to NHS. NHS has very high reactivity, and DNA's amino group reacts with NHS of porystyrene beads. The DNA ciliate body is developed in this process. :
As a motor of DNA ciliate, we used deoxyribozyme which is the enzyme comprised of DNA. Deoxyribozyme cleaves its substrate at an RNA base, if there are 2+ metal ions. Using this reaction, DNA ciliate can move.
Principle and methods
- Two experiments were needed to complete developing DNA ciliate body.
First experiment was creating DNA ciliate by attaching DNAs to polystyrene beads. This process is used the reaction of connecting aminated DNAs’ amino group and polystyrene beads’ carboxylic acid. We took two methods to react. Both methods are used the common reaction, but chemical materials are different. First method is used EDC and NHS. This induces transforming carboxylic acid to succinimide which is been able to react with aminated DNAs and connect.

- Second method is used EDAC. EDAC reacts with both aminated DNAs and polystyrene beads’ carboxylic acid.

Second experiment is confirming whether deoxyribozyme is attached to polystyrene beads and able to cleave substrate. We confirm deoxyribozyme activity by urea-PAGE. Making mixture of DNA ciliate and substrate and Zn2+ ions. If DNA ciliate has deoxyribozyme activity, substrate is cleaved and the band of cleaved substrate appears as a band.
Protocol
- Creating DNA ciliate, we use (1) and (2) protocols.
- (1) The method of using EDC and NHS is here.
- (2) The method of using EDAC…
- Confirming DNA ciliate, we use (3) and (4) protocols
- (3) The method of electrophoresis is here.
- (4) The method of making sample is
Results
- Following is the result of electrophoresis. Result 1 is DNA ciliate using NHS and EDC. Result 2 is DNA ciliate using EDAC. Meanings of the numbers above results are here.
- deoxyribozyme
- substrate
- negative control (Zn(2+) 0 mM)
- positive control (Zn(2+) 10 mM)
- no deoxyribozyme beads (Zn(2+) 0 mM)
- no deoxyribozyme beads (Zn(2+) 10 mM)
- DNA ciliate (Zn(2+) 0 mM)
- DNA ciliate (Zn(2+) 10 mM)
- Three bands were there. From a top, A is the band of deoxyribozyme, B is the band of substrate (didn’t be cleaved), and C is the band of cleaved substrate. If there is cleaved substrate band, it means deoxyribozyme activity is appeared and deoxyribozyme is attached to polystyrene beads successfully.
[Result 1]
- PAGE of φ200 nm polystyrene beads using NHS and EDC.
- PAGE of φ1 um polystyrene beads using NHS and EDC.
[Result 2]
- PAGE of φ200 nm polystyrene beads using EDAC.
Discussions
- First, we explain this experimentation’s appropriateness for checking DNA ciliate body. To create DNA ciliate body, it was necessary to attach deoxyrobozyme to micrometer-sized polystyrene beads firmly. Furthermore, we had to check deoxyribozyme activity of DNA ciliate body. In this method, these two things could be checked. First, the degree of fixation can be checked. DNA ciliate is removed before loading to polyacrylamide gel, so if deoxyribozyme can’t be attached to polystyrene beads firmly, the leg band is appeared. Second, the deoxyribozyme activity of DNA ciliate body can be confirmed. If there is deoxyribozyme activity of DNA ciliate body, the cleaved substrate band is appeared. Based on the above, this method is appropriate for checking DNA ciliate body.
- Second, we explain necessity of lanes. Lanes of 1 to 4 are needed for checking deoxyribozyme activity and the positions of each band of DNAs. Lane 1 and lane 2 are control lanes. The band in lane 1 is deoxyribozyme band, and the band of lane 2 is substrate band. Lane 3 and lane 4 are lanes to check deoxyribozyme activity. The solutions in lane 3 doesn’t contain of Zn(2+), the solution in lane 4 contains of Zn(2+). The cleaved band is appeared in lane 4, but doesn’t be appeared in lane 3. This means normal deoxyribozyme isn’t active in no metal ions solutions. Lane 4’s bottom band means position of cleaved substrate.
- Lanes of 5 to 8 are needed for checking deoxyribozyme activity of DNA ciliate body. Lane 5 and 6 are lanes for checking to polystyrene beads. If polystyrene beads had deoxyribozyme activity, the cleaved band would be appeared. Lanes of 7 and 8 are needed for checking DNA ciliate body’s deoxyribozyme activity. If DNA ciliate has normal deoxyribozyme activity, the cleaved band is appeared in lane 8 because metal ions are needed for deoxyribribozyme activity.