Biomod/2011/TeamJapan/Tokyo/Project
<html>
<style type="text/css">
/* ====================
主に全体に関わるCSS
==================== */
body {behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc);}
.clear {clear:both;}
body.mediawiki {
font-size: 14px;
background-color:#707070; background-position: center center; background-attachment: fixed; background-repeat: no-repeat; font-family: Calibri, Verdana, helvetica, sans-serif; }
h1 {
padding: 0px 20px 5px 20px;
font-size: 34px;
font-weight: bold;
}
h2 {
padding: 20px 20px 5px 20px;
font-size: 25px;
color: #0083eb;
text-decoration: none;
font-weight: bold;}
border: none;
h2 a {
color: #eb8300;
}
h3 {
padding: 20px 20px 5px 20px;
font-size: 20px;
color: #000;
font-decoration: none;
font-weight: bold;
}
h1.firstHeading {
display: none;
}
p { text-align: justify; } a:link { color: #00a5ea; text-decoration: none } a:visited { color:#00a5ea; text-decoration: none } a:hover {
color: #eb8300;
text-decoration: none } a:active { color:#f29400; text-decoration: none } #bodyContent {
width: 970px;
margin: 0px 0px 0px;
background-color:#ffffff;
border-width: 0px 1px 0px 1px;
border-color: #000000;
}
#content {
padding-left: 0px; width: 970px;
}
table#team_members {
text-align: justify;
border: 0;
}
table#team_members h2, table#team_members h3 {
clear: both;
}
#content * a:hover {
text-decoration:none;
}
#main_wrapper {
position:absolute;
left:0px;
top:20px;
margin-top: 0; width: 969px; height: 221px; align: center; border-style: solid;
border-color: white;
} /* ====================
メニューの画像を変更できる部分 ==================== */
#header {
position:relative;
left:0px;
top:0px;
margin-top: 0; width: 969px; height: 221px; align: left; background-color: #FFFFFF;
background-image: url(http://openwetware.org/images/e/e0/NEW_header.jpg);
} /* ====================
以下、特殊なclassに適用される ==================== */
#navigation { position:absolute;
left:18px; top:155px; width:1200px; height:69px;
z-index:100;
background-color: transparent;
float: left;
color: #0000FF; } #super_main_wrapper {
position:absolute;
left:0px;
top:227px;
width: 975px; align: center; background-color: #ede8e2;
heigth: auto;
}
#SubWrapper {
width: 645px;
padding: 0px;
border-left:4px solid #ede8e2;
float: left;
margin-top: 0px;
background-color: #ede8e2;
}
#SubWrapper * p, #SubWrapper p {
padding: 0 20px;
text-align: justify;
font-size: 12px;
}
#SubWrapper * h3, #SubWrapper h3 {
padding-top: 10px;
font-size: 18px;
}
#news {
width: 322px;
margin-top: 0px;
float: left;
background-color: #d8d5d0;
border-right:4px solid #ede8e2;
}
#news p {
padding: 0 20px 20px 20px;
text-align: justify;
font-size: 12px;
}
#news h3 {
padding: 10px 20px;
font-size: 18px;
}
#mission_box {
width:650px;
float: left;
}
#team_box, #heartbeat_box, #notebook_box, #parts_box, #gallery_box, #sponsors_box, .boxy {
width:215px;
float: left;
padding: 10px 0 0 0;
}
div.tleft {
border-width: 0px;
margin:0;
padding:0;
border-color:transparent;
}
/* ====================
ここからプルダウン周辺のデザイン ==================== */
- menu * {
margin: 0; padding: 0; }
- menu {
behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc); font-family: calibri, verdana, sans-serif;
font-color: #ffffff;
font-size: 19px; background-color: transparent; float:left; padding: 12px 0 0 0; }
- menu ul {
float: left; list-style: none; }
- menu ul li {
background-color:transparent;
position:relative;
float:left; list-style: none; padding: 10px 20px 0 0;
font-weight: bold;
width: auto;
}
- menu a {
color: #FFFFFF; display: inline; text-decoration: none; }
- menu a:visited {
color:#FFFFFF; text-decoration: none }
- menu a:hover {
color: #00a5ea; }
- menu ul li ul {
display: none; position: absolute; left: 0px;
width: 155px;
heigth: 1%;
font-size: 19px; opacity: 0.8; list-style: none;
top: 30px;
padding-top: 20px;
z-index:500;
}
- menu ul li:hover ul {
display: inline;
background-position: bottom;
}
- menu ul li ul li {
width: 100%; list-style: none;
background-color: #000;
margin: -1px;
padding: 0px 0 0 5px;
display: inline;
}
</style>
<script type="text/javascript">
/***********************************************
- CSS Vertical List Menu- by JavaScript Kit (www.javascriptkit.com)
- Menu interface credits: http://www.dynamicdrive.com/style/csslibrary/item/glossy-vertical-menu/
- This notice must stay intact for usage
- Visit JavaScript Kit at http://www.javascriptkit.com/ for this script and 100s more
- /
var menuids=new Array("verticalmenu") //Enter id(s) of UL menus, separated by commas var submenuoffset=-2 //Offset of submenus from main menu. Default is -2 pixels.
function createcssmenu(){ for (var i=0; i<menuids.length; i++){
var ultags=document.getElementById(menuids[i]).getElementsByTagName("ul")
for (var t=0; t<ultags.length; t++){
var spanref=document.createElement("span")
spanref.className="arrowdiv" spanref.innerHTML=" " ultags[t].parentNode.getElementsByTagName("a")[0].appendChild(spanref)
ultags[t].parentNode.onmouseover=function(){
this.getElementsByTagName("ul")[0].style.left=this.parentNode.offsetWidth+submenuoffset+"px"
this.getElementsByTagName("ul")[0].style.display="block"
}
ultags[t].parentNode.onmouseout=function(){
this.getElementsByTagName("ul")[0].style.display="none"
}
}
}
}
if (window.addEventListener)
window.addEventListener("load", createcssmenu, false)
else if (window.attachEvent)
window.attachEvent("onload", createcssmenu)
</script>
</html>
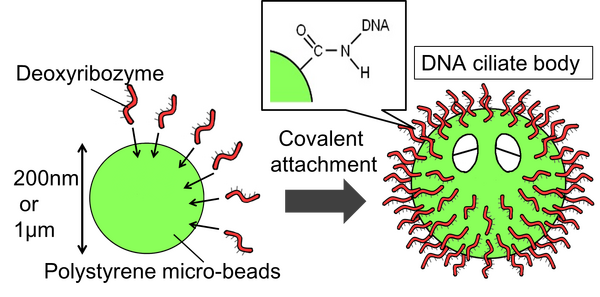
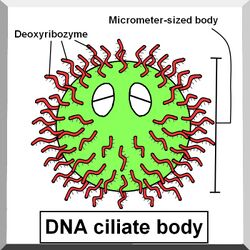
Project
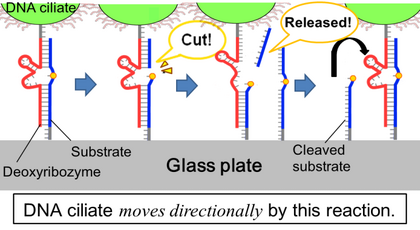
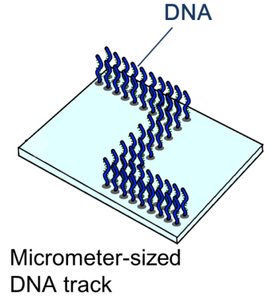
The body of the DNA ciliate