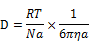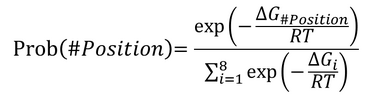Biomod/2011/TeamJapan/Tokyo/Project/Simulations
<html>
<style type="text/css">
/* ====================
主に全体に関わるCSS
==================== */
body {behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc);}
.clear {clear:both;}
body.mediawiki {
font-size: 14px;
background-color:#707070; background-position: center center; background-attachment: fixed; background-repeat: no-repeat; font-family: Calibri, Verdana, helvetica, sans-serif; }
h1 {
padding: 0px 20px 5px 20px;
font-size: 34px;
font-weight: bold;
}
h2 {
padding: 20px 20px 5px 20px;
font-size: 25px;
color: #0083eb;
text-decoration: none;
font-weight: bold;}
border: none;
h2 a {
color: #eb8300;
}
h3 {
padding: 20px 20px 5px 20px;
font-size: 20px;
color: #000;
font-decoration: none;
font-weight: bold;
}
h1.firstHeading {
display: none;
}
p { text-align: justify; } a:link { color: #00a5ea; text-decoration: none } a:visited { color:#00a5ea; text-decoration: none } a:hover {
color: #eb8300;
text-decoration: none } a:active { color:#f29400; text-decoration: none } #bodyContent {
width: 970px;
margin: 0px 0px 0px;
background-color:#ffffff;
border-width: 0px 1px 0px 1px;
border-color: #000000;
}
#content {
padding-left: 0px; width: 970px;
}
table#team_members {
text-align: justify;
border: 0;
}
table#team_members h2, table#team_members h3 {
clear: both;
}
#content * a:hover {
text-decoration:none;
}
#main_wrapper {
position:absolute;
left:0px;
top:20px;
margin-top: 0; width: 969px; height: 221px; align: center; border-style: solid;
border-color: white;
} /* ====================
メニューの画像を変更できる部分 ==================== */
#header {
position:relative;
left:0px;
top:0px;
margin-top: 0; width: 969px; height: 221px; align: left; background-color: #FFFFFF;
background-image: url(http://openwetware.org/images/e/e0/NEW_header.jpg);
} /* ====================
以下、特殊なclassに適用される ==================== */
#navigation { position:absolute;
left:18px; top:155px; width:1200px; height:69px;
z-index:100;
background-color: transparent;
float: left;
color: #0000FF; } #super_main_wrapper {
position:absolute;
left:0px;
top:227px;
width: 975px; align: center; background-color: #ede8e2;
heigth: auto;
}
#SubWrapper {
width: 645px;
padding: 0px;
border-left:4px solid #ede8e2;
float: left;
margin-top: 0px;
background-color: #ede8e2;
}
#SubWrapper * p, #SubWrapper p {
padding: 0 20px;
text-align: justify;
font-size: 12px;
}
#SubWrapper * h3, #SubWrapper h3 {
padding-top: 10px;
font-size: 18px;
}
#news {
width: 322px;
margin-top: 0px;
float: left;
background-color: #d8d5d0;
border-right:4px solid #ede8e2;
}
#news p {
padding: 0 20px 20px 20px;
text-align: justify;
font-size: 12px;
}
#news h3 {
padding: 10px 20px;
font-size: 18px;
}
#mission_box {
width:650px;
float: left;
}
#team_box, #heartbeat_box, #notebook_box, #parts_box, #gallery_box, #sponsors_box, .boxy {
width:215px;
float: left;
padding: 10px 0 0 0;
}
div.tleft {
border-width: 0px;
margin:0;
padding:0;
border-color:transparent;
}
/* ====================
ここからプルダウン周辺のデザイン ==================== */
- menu * {
margin: 0; padding: 0; }
- menu {
behavior: url(http://www.xs4all.nl/~peterned/htc/csshover3-source.htc); font-family: calibri, verdana, sans-serif;
font-color: #ffffff;
font-size: 19px; background-color: transparent; float:left; padding: 12px 0 0 0; }
- menu ul {
float: left; list-style: none; }
- menu ul li {
background-color:transparent;
position:relative;
float:left; list-style: none; padding: 10px 20px 0 0;
font-weight: bold;
width: auto;
}
- menu a {
color: #FFFFFF; display: inline; text-decoration: none; }
- menu a:visited {
color:#FFFFFF; text-decoration: none }
- menu a:hover {
color: #00a5ea; }
- menu ul li ul {
display: none; position: absolute; left: 0px;
width: 155px;
heigth: 1%;
font-size: 19px; opacity: 0.8; list-style: none;
top: 30px;
padding-top: 20px;
z-index:500;
}
- menu ul li:hover ul {
display: inline;
background-position: bottom;
}
- menu ul li ul li {
width: 100%; list-style: none;
background-color: #000;
margin: -1px;
padding: 0px 0 0 5px;
display: inline;
}
</style>
<script type="text/javascript">
/***********************************************
- CSS Vertical List Menu- by JavaScript Kit (www.javascriptkit.com)
- Menu interface credits: http://www.dynamicdrive.com/style/csslibrary/item/glossy-vertical-menu/
- This notice must stay intact for usage
- Visit JavaScript Kit at http://www.javascriptkit.com/ for this script and 100s more
- /
var menuids=new Array("verticalmenu") //Enter id(s) of UL menus, separated by commas var submenuoffset=-2 //Offset of submenus from main menu. Default is -2 pixels.
function createcssmenu(){ for (var i=0; i<menuids.length; i++){
var ultags=document.getElementById(menuids[i]).getElementsByTagName("ul")
for (var t=0; t<ultags.length; t++){
var spanref=document.createElement("span")
spanref.className="arrowdiv" spanref.innerHTML=" " ultags[t].parentNode.getElementsByTagName("a")[0].appendChild(spanref)
ultags[t].parentNode.onmouseover=function(){
this.getElementsByTagName("ul")[0].style.left=this.parentNode.offsetWidth+submenuoffset+"px"
this.getElementsByTagName("ul")[0].style.display="block"
}
ultags[t].parentNode.onmouseout=function(){
this.getElementsByTagName("ul")[0].style.display="none"
}
}
}
}
if (window.addEventListener)
window.addEventListener("load", createcssmenu, false)
else if (window.attachEvent)
window.attachEvent("onload", createcssmenu)
</script>
</html>
Simulations for track walking mode
Principles and methods of simulations
Method
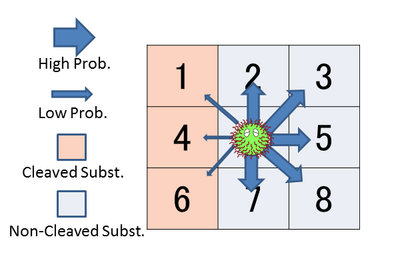
- In our cellular-automaton simulations, we defined cells as followings.
- Length of side of a cell is the same to the diameter of DNA ciliate body.
- A yellow cell represents DNA ciliate.
- Blue cells represent oligodeoxynucleotide substrates with a single ribose moiety.
- Red cells represent oligodeoxynucleotide substrates without a ribose moiety.
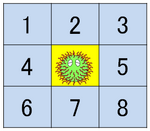
- Green cells represent UV-switching DNA which are spotted UV.
- A Yellow cell makes a move depend on its located cell. The rate of moving on each cell is derived as following steps.
- Yellow cells don’t move to the yellow cell. This means that DNA ciliate cannot move to the exact cell where the DNA ciliate is already allocated.
- We calculated the average time that DNA ciliate moves one micrometer, the rate that yellow cells don’t move at the present cell, and the rate that yellow cells moves to each cell. The detail of the calculations is following.
- Firstly, we calculated the average time that DNA ciliate moves one micrometer by Brownian movement as following formula.
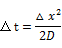
|
|
- Secondly, we calculated the rate that yellow cells don’t move based on two assumptions. One assumption is that DNA ciliate moves to another cell with cleaving substrate in 120 seconds at a rate of 0.5. In the other assumption, DNA ciliate moves to another cell without cleaving substrate in 1.2 seconds at a rate of 0.5. This assumption is based on this thesis[1]. In this thesis, the time which DNA releases substrate by using enzyme activity is slower about 100 times than the time which DNA releases substrate by not using enzyme activity. We defined the “step” as an indicator of time. The “step” is the unit time. The step of DNA ciliate which is 1μm in diameter is about 1.02 seconds. The step of DNA ciliate which is 200nm in diameter is about 0.00815 seconds.
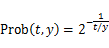
|
|
- Then, each rate is following.
- The diameter of DNA ciliate body is 1μm.
- On the blue cells, yellow cells do not move at a rate of 0.9941
- On the red cells, yellow cells do not move at a rate of 0.555.
- On the white cells, yellow cells do not move at a rate of 0 (means to move absolutely).
- The diameter of DNA ciliate body is 200nm.
- On the blue cells, yellow cells do not move at a rate of 0.999529.
- On the red cells, yellow cells do not move at a rate of 0.0053006.
- On the white cells, yellow cells do not move at a rate of 0 (means to move absolutely).
- Thirdly, we calculated the rate that yellow cells move to each cell.
- Yellow cells can move to adjacent cells at one step and cannot move to distant cells.
- The direction of yellow cells' movement is stochastically dependent on free-energy of the adjacent cells. The free-energy of each cell is followings. Blue, red and green cells have the free energies of non-cleaved substrate(-19.02[KJ/mol]), cleaved substrate (-16.03[KJ/mol]) and UV-switching DNA (-49.90[KJ/mol]). For example, Yellow cells are about 211 times more likely to move to blue cells than to red cells.
- ΔGi: An free energy of cell i which is adjacent;
- R: Gas constant. 8.3145[P/mol*K]
- T: Temperature. 298[K].
- Based on these three calculations, we simulated the movement of DNA ciliates.
Results
- We simulated movement of DNA ciliate
Simulation movie of track walking mode (a single DNA ciliate)
- We simulated the movement of DNA ciliate on DNA track walks. In the simulation, the width of the DNA track was 4μm, DNA ciliate was 1μm in diameter, and the length of the DNA track was 50μm.

<HTML><body>
</body></HTML>
- In this simulation, we confirmed that DNA ciliate moved from left to the right.
- To confirm “track walking mode” works correctly, we made line graphs which means the migration length of DNA ciliate. The vertical axis shows the distance from origin. The horizontal axis shows the number of steps. Three lines are different in the DNA types of DNA track. The red line's DNA track is substrate, so DNA ciliate moves with cleaving substrate. The blue line's DNA track is cleaved substrate, so DNA ciliate moves without cleaving in this situation. The green line's DNA track is complementary strands for deoxyribozyme, so DNA ciliate can hybridize with, but cannot cleave the DNA. Figure 1 and 2 are the graphs of the movement of 1μm and 200nm DNA ciliate, respectively.
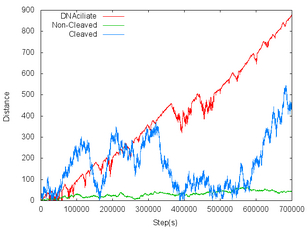 | 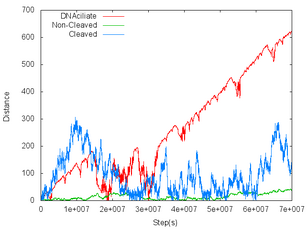 |
- From these graphs, we found that DNA ciliate moving on substrate moves faster and more directly than the others.
- To confirm DNA ciliate is forced motion, we make mean square displacement graph. These graphs are the average of the movement by 30 simulations. In mean square displacement, Brownian motion is presented by linear function and forced motion is presented by quadratic function.
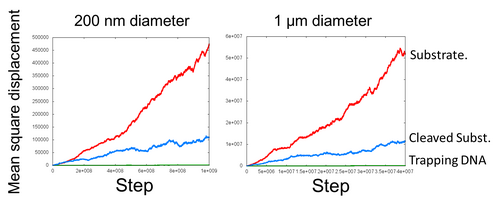
- In this simulation, the moving distance on substrate field is approximately presented by quadratic function. We confirmed DNA ciliate is forced directly and moves intended direction on substrate in this simulation. Furthermore, DNA ciliate on DNA track walks is faster than on cleaved substrate.
Simulation movie of DNA ciliates on DNA track (several DNA ciliates)
- Actually, it is difficult to put a single DNA ciliate on DNA track walks, so it is necessary to confirm DNA ciliates can moves directly on DNA track if several DNA ciliates are put on at a time. We simulated the movement of several DNA ciliates on DNA track. The number of DNA ciliates is 12, the width of DNA track is 30 μm, and the length of DNA track is 100 μm.

<HTML><body>
</body></HTML>
- From the result, most of DNA ciliates move to the right. It is confirmed several DNA ciliates moves directly on DNA track if several DNA ciliates are put on at a time.
Simulation of light-irradiate mode on DNA track
- We simulated the movement of several DNA ciliates on DNA track which contains UV-switching DNA. The UV-switching DNA is spotted UV, so DNA ciliates are trapped by the DNA. Though we did not assume the substrate as the UV-switching DNA in the previous video, we assumed the substrate as the UV-switching DNA in the following video. The number of DNA ciliates is 12, the width of DNA track is 30 μm, and the length of DNA track is 100 μm.
<HTML><body>
</body></HTML>
- From the result, most of DNA ciliates gathered at the area of UV-switching DNA, so it is confirmed DNA ciliate on DNA track walks gather at UV-switching DNA.
Discussion
- We simulated the movement of each DNA ciliate on a long DNA track. The following graph is the mean square displacement graph for movement of DNA ciliate. This graph is same to figure3, except in the length of the x axis.
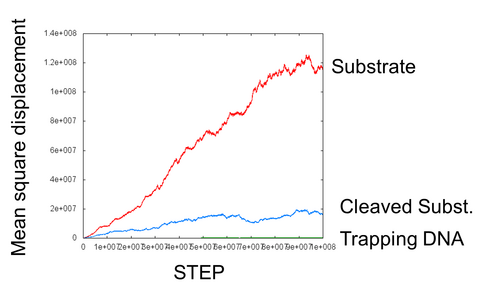
- In this graph, the red line (on DNA track) changes quadratic function to linear function little by little. It is thought that the movement of DNA ciliate which walk on DNA track change track walking mode to free moving mode.
- To consider about this incomprehensible behavior, we made line graphs which mean the movement of each DNA ciliate. Each line of these graphs means a single simulation, NOT the average of several simulations. The left graph is square displacement of each DNA ciliate on DNA track walks. The right graph is the movement of DNA ciliate on cleaved substrate.
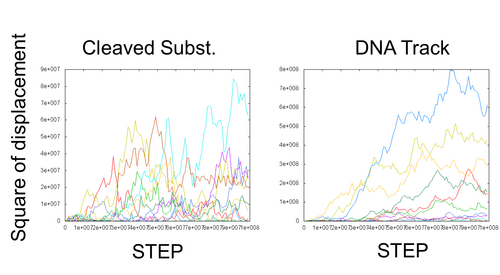
- In left graph, all lines are very rough. It is thought DNA ciliate moves at random on cleaved substrate.
- In right graph, most of lines are approximated quadratic functions at first, but some of lines became rough. It is thought DNA ciliate on DNA track walk along with track at first, but a short time later, some DNA ciliates became moving on cleaved substrate. It is thought that the number of the DNA ciliates which move on cleaved substrate become large little by little because the probability of moving to cleaved substrate and starting free moving become large as time passes.
- To reduce the probability of moving to cleaved substrate and starting free moving, it is thought to reduce the free-energy of substrate. However, DNA ciliate takes longer time to separate from substrate or cleaved substrate, which will result in slow movement of the DNA ciliate. The speed and the distance of keep walking on DNA track are trade-off.
- In conclusion, we confirmed by a simulation that DNA ciliate moves by directional force, not by Brownian motion in “track walking mode”. Furthermore, we found the speed of movement on DNA track is faster than on cleaved substrate.
Program sources
Simulation for no light-irradiated DNA and single DNA ciliate
Simulation for no light-irradiated DNA and several DNA ciliates
Simulation for light-irradiated DNA and several DNA ciliates
References
[1] Mark J. Olah, Darko Stefanovic. "Multivalent random walkers: a model for deoxyribozyme walkers." The 17th International Meeting on DNA Computing and Molecular Programming, LNCS 6937, pp.160-174, 2011.