Biomod/2011/PSU/BlueGenes/results
|
Home ::: Overview ::: Methods ::: Results ::: Application ::: Literature ::: Team |
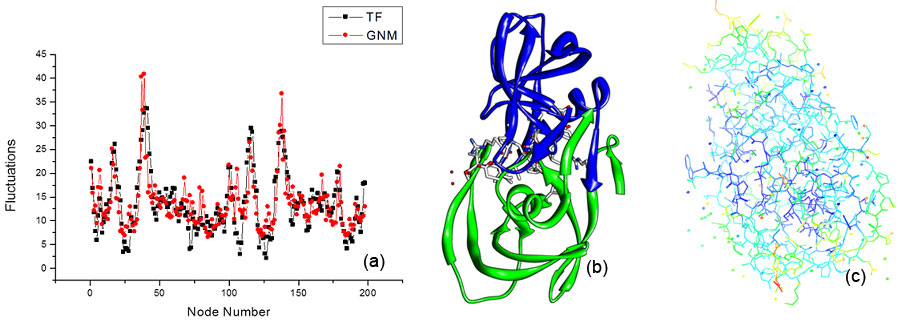
ResultsGNM with ProteinGNM has already been thoroughly and successfully applied to proteins. There is even a database known as iGNM that has calculated protein fluctuations for over 20,000 structures from the protein data bank or PDB. In most cases, carbon atoms are chosen as the nodes to represent the network. A common cutoff value is 7 Angstroms. To show you that GNM works, in Figure 1, we calculated the fluctuation for HIV-1 protease and compared it to the temperature factor, which are experimental values. There are 200 nodes in this structure.
Qualitatively, it is very easy to see that the GNM results (shown in red) match very well to the TF results (shown in black). Each dot represents a node and lines are simply to help guide your eyes. GNM has been successful in calculating protein fluctuations because of well matching results such as the one in Figure 1.
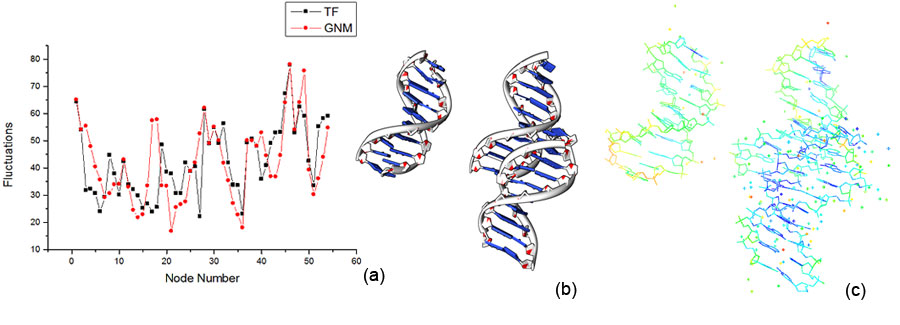
GNM with DNAThere haven’t been many applications of GNM on DNA structures. But since GNM filters out the chemistry of the atoms and simply looks at the bonds between the nodes, there is no reason why it shouldn’t work. In Figure 2, we tried GNM with DNA of PDB ID 307D. We used the phosphate atoms as the nodes and a cutoff distance of 13 Angstroms. There are 60 nodes. Results are pretty satisfactory but are slightly less accurate than the comparison with protein. This is because proteins have longer chains than DNA and more nodes, thus it rules out randomness. DNA is more flexible and thus harder to calculate the flexibility. We chose 307D strictly because it had a long chain.
Again, DNA fluctuation results from GNM (in red) are compared to TF results (in black). Points represent nodes and lines are to guide your eyes.
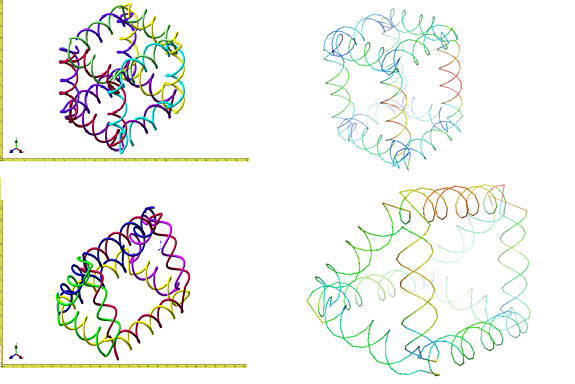
GNM with Synthetic DNANow that we have shown that GNM is applicable in both proteins and DNA, we want to use it to calculate fluctuations of synthetic DNA. Below you can see 2 structures that we've made using NanoEngineer-1, a cube and a prism. High fluctuations are shown in warm colors (red, orange, yellow/yellow-green) and low fluctuations are shown in cool colors (green, blue, dark blue). Both structures show similar satisfactory results. It makes sense to think that the vertices of the structure are the least flexible because they are the most confined. The legs of the prism are more flexible because they are not as rigidly confined.
AdvantagesThere are multiple advantages of our GNM method. They include:
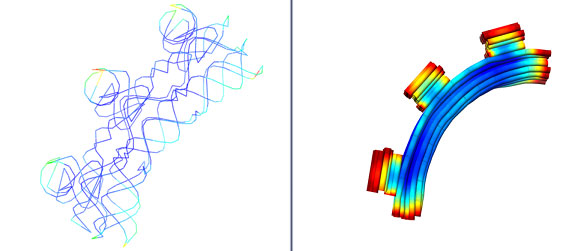
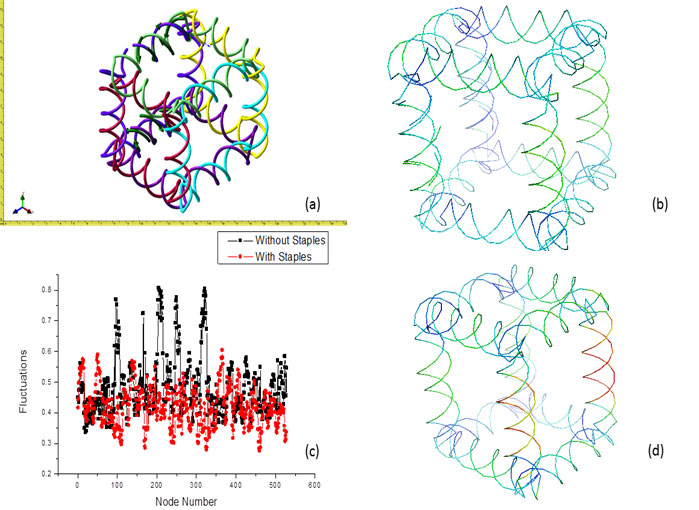
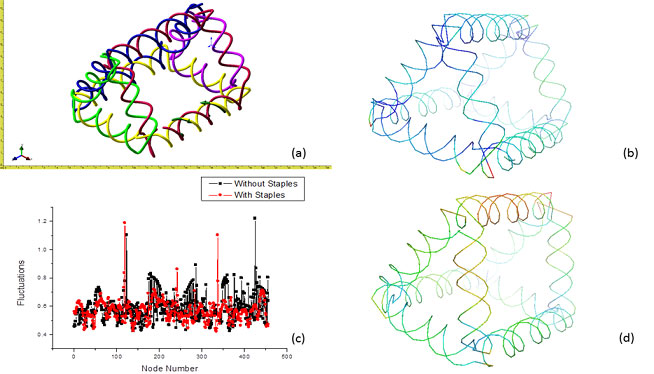
FeedbackStructural feedback is one of the most advantageous parts of our GNM method. It allows one to modify the structure fluctuations on a custom level. Figure 5 below displays this aspect of our method. Figure 5(d) shows the original structure. One may think that each leg of the structure should have even flexibility because it is symmetric. However, this is not the case as the 2 front legs seem to have a higher flexibility then the back legs and even their fluctuations are not the same. The reason for this is because these structures were built by hand. There will always some errors in the structures that cause these variations. Figure 5(b) on the other hand, shows the structure after feedback has been applied to it. Staples were added to the front legs thus stabilizing the structure (noting that warm colors are high fluctuation and cool colors are low fluctuation). Staples are simply a bond between one double stranded DNA strand to the next. Adding staples to the DNA structure introduces confinement to the strands, which in turns, decreases the flexibility. Figure 5(c) shows a comparison of the fluctuation data from before and after staple confinement. The red data shows structure flexibilities after staples were added while the black shows flexibilities from the original structure. Each point represents a node and the lines are simply to guide your eyes. It can be seen that the fluctuations are lessened after staples are added, emphasizing the point that staples within the structure help to stabilize and can be used to customize structure properties. Figure 5(a) is simply to show what the structure appeared like in NanoEngineer-1 for clarity.
FilesStructure files: Media: NE1 structures.rar |