Biogang:Projects/Protein folding
|
Home Projects Discussion Events FriendFeed - The Life Scientists |
This an idea I had some time ago but never got around to push it forward (mostly because I'm not a mathematician neither biophysicits). If the Biogang has any idea how to approach it, it's going to be very nice story :) Pawel Szczesny 19:22, 24 June 2008 (UTC).
The project is not really about protein folding as a process - it asks if there's anything similar on the mathematical level between tranformations of completely artificial representation of protein sequence to another artificial representation of protein structure. Or even if there's anything similar on any level when looking at transformation between abstracted sequence and abstracted structure.
The key here is to stop thinking about sequence and structure in biophysical terms. Pairwise alignment is scored by completely non-biological function. Gap penalties have no biological and biophysical meaning, dynamic programming algorithm is not approximation of biological events. However, we are still trying to predict protein structure ab initio using biophysical properties or statistically derived terms from the known structures.
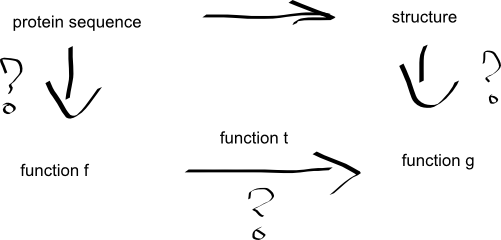
The graph above contains three question marks. First two mark abstractions of protein sequence and structure, the third asks about tranformation of one into the other. And while the obvious focus would be on the transformation, I think the most difficult would be to invent smart representations of proteins.
The basic idea is as follows:
- represent a protein sequence as a mathematical function f (requiring that particular f represent only one sequence)
- represent a backbone structure that the sequence folds into as a mathematical function g (requiring that particular g represent only one backbone topology - fold)
- compute a function t which transforms function f (representing sequence) to function g (representing structure)
- repeat above steps for two cases
- closely related sequences folding into the same structure
- completely different sequences folding into different structures