BME100 s2017:Group9 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
OpenPCR program 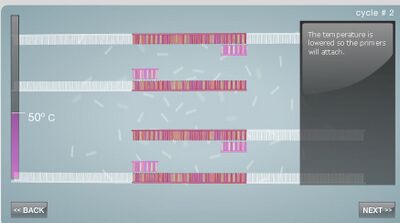  HEATED LID: 100°C
Research and DevelopmentPCR - The Underlying Technology Component functions of the PCR reactionA template DNA strand is used as a model for the newly synthesized strand of DNA. It contains the genes that are to be copied into the new strand. DNA primers are used to indicate the starting point of DNA replication. The Taq polymerase enzyme will recognize and bind to primers on the template DNA strand and then begin replication. The enzyme Taq polymerase binds to the primed sequences of DNA and adds nucleotides to extend the second strand. The deoxyribonucleotides are the monomers of a DNA strand, containing a nitrogenous base, deoxyribose sugar and a phosphate group. Taq polymerase synthesizes these nucleotides to grow the replicated DNA strand. The deoxyribonucleotides bind to each other through complementary base pairing. What happens to the components (listed above) during each step of thermal cycling?The first step of PCR is applying heat and raising the temperature to 95°C for 3 minutes. This simply heats up the solution. During the second step, or denaturing at 95°C for 30 seconds, the strands of the template DNA split up. The third step is lowering the temperature to 57°C for 30 seconds, which allows the primers to bind to the complimentary target sequence. The next step is to raise the temperature slightly to 72°C for 30 seconds, which is when the taq polymerase attaches to the primers. The last step is to hold the 72°C for 3 minutes, which is when the taq polymerase arranges nucleotides to make a new DNA sequence. The final hold at 4°C stops the reaction until the next run. Which DNA base pair anneals to each baseThere are four nucleotides, and they are thymine, adenine, guanine, and cytosine, or T, A, G, and C, respectively. T and A correspond, and C and G correspond. So when the taq polymerase is making its way along a DNA strand replicating, it comes accross each of these nucleotides. Whent he polymerase reads a T, it places an A on the new strand. When it reads a C, it places a G on the new strand. When it reads an A, it places a T. And lastly, when it reads a G, it places a C. During which two steps of thermal cycling does base-pairing occur?Base-pairing occurs during the third and fourth steps of thermal cycling. It occurs initially when the Taq polymerase enzyme adds DNA nucleotides to the replicated strand after it has bonded to the primer sequence of DNA. Base-pairing then continues to occur when the PCR cycle is repeated through 35 cycles, creating billions of replicated DNA strands. 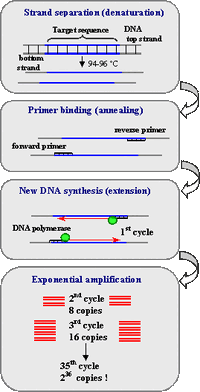 Image source: https://www.ncbi.nlm.nih.gov/probe/docs/techpcr/ SNP Information & Primer DesignBackground: About the Disease SNP SNPs are single nucleotide pairs that are unique for each individual. These differences in the DNA strands are only one nucleotide different than everyone else but they are undetectable by the naked eye. These changes alter how humans react to drugs and the strength of their reaction. Primer Design and Testing
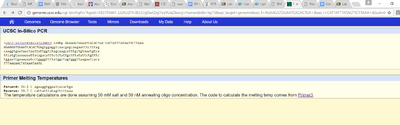 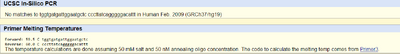 | ||||||||||||||||||||||||||||||||||





