BME100 s2017:Group7 W1030AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Thermal Cycling
http://perlong.en.made-in-china.com/ Heated Lid: 100°C Initial Step: 95°C for 2 minutes Number of cycles: 25
Final Step: 72°C for 2 minutes Final Hold: 4°C
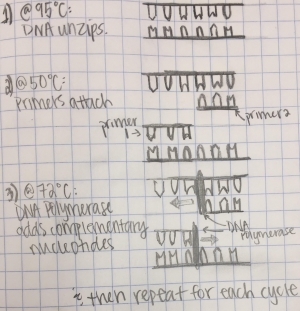
Research and DevelopmentPCR - The Underlying Technology Functions of Each Component: Steps of Thermal Cycling: During thermal cycling some DNA, dNTP's, DNA polymerase, and primers to the PCR tube, and then placed into the DNA Thermal Cycler. For 3 minutes the temperature is set to 95°C for 3 minutes. At this step it is warming up, almost boiling, and the DNA is getting ready to separate its strands. At 95°C for 30 seconds the DNA starts to denature, meaning that it is separating and as a results makes two single-stranded DNA molecules. At 57°C for 30 seconds, a single stranded DNA molecules naturally attempt to pair up. The primers crowd up and lock to a target before strands can rejoin. At 72°C for 30 seconds The DNA polymerase is activated. When the polymerase finds a primer attaches to a single DNA strand it begins to make a complementary nucleotides on the strand. At the final step for 3 minutes it is set to 72°C, where it goes on until it reaches the end of the strand and falls off. A final hold can be placed at 4°C and this is used to limit the activity of Taq DNA polymerase sot this can prevent potential non-specific binding of primers. cite: http://learn.genetics.utah.edu/content/labs/pcr/ Nucleotide Pairing:
Where Base-Pairing Occurs: Base-pairing happens between the steps of Annealing and Extending. At annealing the temperature is at 57°C. At this step the primer attaches to the separated DNA strands. At the extending stage the temperature is at 72°C. This lets the DNA polymerase attach to the primer. Then, the DNA polymerase begins to make complementary nucleotides on the strand.
SNP Information & Primer DesignBackground: About the Disease SNP A SNP is a single nucleotide polymorphism. A nucleotide is a basic structural unit of nucleic acids. Polymorphism describes the variety that nucleotides can have in a singular strand. In this lab, we are looking into the disease related SNP. Disease SNP are nucleotides that cause illness from their creation. Specified SNP SNP: rs121908757 This is variation of SNP is found in humans, Homo sapiens. Within humans, the seventh chromosome is where it is located. This SNP is pathogenic, disease causing, linking it to cystic fibrosis and sever cardiac impact. Being called CTFR or, Cystic fibrosis transmembrane conductance regulator, creates a protein that makes a channel across the membrane of cells that produce mucus, sweat, saliva, tears, and digestive enzymes. Terms associated with CTFR, such as, PDZ domain binding, intracellular ATPase-gated chloride channel activity and protein binding, describe where CFTR has its effect on. Going into more detail, an allele is variant gene created by codons. what causes a problem is when a mutation occurs to a specific allele,"AGT", where the A is switched with a C. This SNP is located numerical on position 117,587,799.
For the primer design, we will need the 20 codons that come before SNP location, having the last one being the change. In this case, the primer would be 5'- AGAAGGTGGAATCACACTGA .For the PCR process, we need reverse primer, which we create by going 200 codons past the location position of the mutation, in this case, 117,587,999. For the Reverse primer, it would be 5'- CATTATTTATAGTTCTTAAA. Now, we would need to pair this with the disease primers. Now to create these, we only need to change the last codon of the first primer into the disease causing mutation. we also keep the reverse primer the same. We validate these primers and with confirmation, the primers are done at last.
| ||||||||||||||||||||||||||||||||