BME100 s2017:Group6 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
(http://www.promega.com/resources/protocols/product-information-sheets/g/gotaq-colorl
OpenPCR program
Research and DevelopmentPCR - The Underlying Technology Q1. The Component Functions of A PCR Reaction In the PCR reaction, template DNA is the strand of DNA with the target sequence. DNA polymerase uses template DNA to magnify the the target DNA. Primers are short strands of DNA or RNA that start the DNA replication process. They are very precise when doing their job and therefore, they are a good tool to use to identify a specific DNA sequence. Taq Polymerase is responsible for reading the DNA code and then attaching nucleotides (the genetic building blocks: A,C, G, T) to create new DNA strands. They allow the creation of billions of new DNA copies during the PCR reaction. Deoxyribonucleotides make up a string of DNA. They are made up of three different components: deoxyribose sugar, a phosphate group and a nitrogen base.
At the initial step of thermal cycling at 95 degrees Celsius for three minutes, the thermal cycler heats up the solution to almost boiling point. For the next thirty seconds, the DNA double helix splits creating two single-stranded DNA molecules. After the thermal cycler cools down to 50 degrees Celsius, because there is an ample amount of primers in the tube, the primers hurry and lock onto their target before the single strands of DNA can rejoin. Then, raising the temperature to 72 degrees Celsius for thirty seconds, the DNA polymerase are activated. When the polymerase finds a primer attached to a single strand of DNA, it begins to add nucleotides to it until it gets to the end of the strand and falls off. Once this process is complete, the temperature is decreased to four degrees Celsius to store and hold the DNA, preventing it from heating up again. This final process allows the newly replicated DNA stay within the double-strands and prevents it from further replication.
Nucleotides always pair with a corresponding nucleotide. Adenine (A) always pairs with Thymine (T) and vice versa. In additon, Cytosine (C) always pairs with Guanine (G) and vice versa. The nucleotides pair this way because it allows for hydrogen bonds to form.
Base-pairing occurs during both the anneal step at 57 degrees Celsius as well as the extension step at 72 degrees Celsius. During annealing, the system is cooled and the target DNA is binded with short DNA primers that serve as starting positions for replication. During the extension stage, the temperature is elevated and two taq polymerase come in and match the base primers with their pairs and extend the primers forming new nucleotide strands.
The temperature is increased to just below boiling point, at 95° C, in order to denature the double helix. Error creating thumbnail: File with dimensions greater than 12.5 MP The denatured DNA. Error creating thumbnail: File with dimensions greater than 12.5 MP Primers attach to the DNA when the temperature is lowered to 57° C. Error creating thumbnail: File with dimensions greater than 12.5 MP The temperature is increased once again but up to 72° C to allow Taq Polymerase to initiate replication. Error creating thumbnail: File with dimensions greater than 12.5 MP The first cycle is complete. SNP Information & Primer DesignBackground: About the Disease SNP A nucleotide is a group of molecules comprised of a phosphate group, bases, and a pentose sugar. The bases for DNA include adenine, guanine, cytosine, and thymine; in RNA, thymine is replaced by uracil. Polymorphism is a sequence difference, whcih includes Short Nucleotide Polymorphism, or SNP for short. It is a pathogenic disease caused by a mutation on chromosome 7, position 117587799. This mutation occurs when an adenine nucleotide is replaced by a cytosine nucleotide. It is found only in humans and is linked with the condition known as cystic fibrosis.
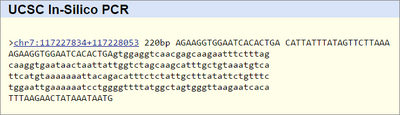
The non diseased primer was determined by finding the defective nucleotide causing Cystic Fibrosis Transmembrane Conductance Regulator (CFTR), and changing it.Using the UCSC database, the primers that were designed for non-diseased and diseased were searched. There was a match found for the non-diseased primer, with the position different from the expected. The diseased primer returned no results, since the disease is not part of the database.
Non-disease Primer Disease Primer | ||||||||||||||||||||||||||||||||||