BME100 s2017:Group3 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||
OUR TEAMLAB 4 WRITE-UPProtocolMaterials
Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds
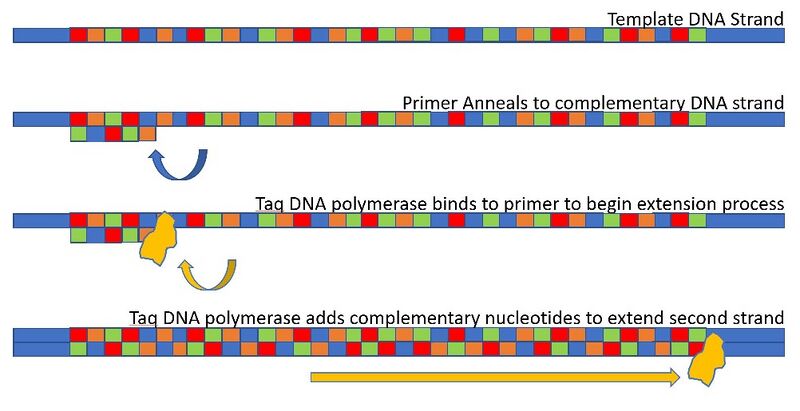
Research and DevelopmentPCR - The Underlying Technology Components of a PCR reaction A PCR reaction begins with a template DNA used as a base for creating the complementary base pairing strand. Primers are what binds to the template strand to bracket the target sequence to start replication. Afterwards, a taq polymerase attaches to the primer and assembles nucleotides into a new DNA strand. Deoxyribonucleotides (dNTP's) are the complementary nucleotides added to extend the second strand of DNA. Steps of Thermal Cycling The changes in temperature allows for the PCR reaction to occur. The initial step of increasing the temperature to 95 degrees Celsius for 3 minutes heats up the DNA and the strand begins to unwind. Afterwards, denaturing occurs, where the DNA separates into a single strand when the temperature is at 95 degrees for 30 seconds. At the anneal stage where the temperature is at 57 degrees for 30 seconds, primers bind to complementary DNA sequences. The temperature is then raised again to 72 degrees for 30 seconds for Taq DNA polymerase to bind to the primers for extension. Finally, at 72 degrees for 3 minutes, dNTP's are added to extend the second strand of DNA. Base-pairing There are four types of nucleotides that make up DNA. These nucleotides are Adenine (A), Thymine (T), Cytosine (C), and Guanine (G). Hydrogen bonding binds these nucleotides to create base-pairing. Nucleotides A and T binds together and C and G nucleotides bind together. Base-pairing occurs during the annealing and extension steps of thermal cycling. During Annealing DNA is bonded with the DNA primers as a starting position for replication. Temperature is then slightly elevated as the thermal cycling stage of extension occurs. Here, two taq polymerase come and match with the base pairs.
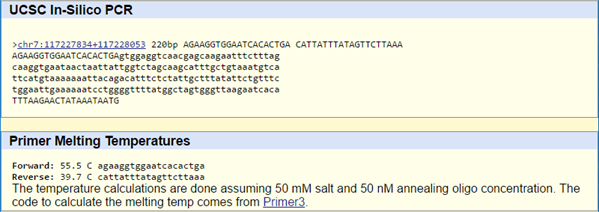
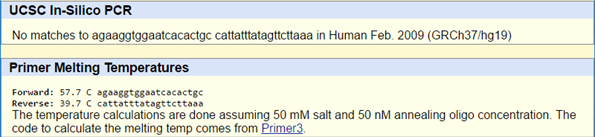
SNP Information & Primer DesignBackground: About the Disease SNP The disease SNP (single nucleotide polymorphism) is a pathogenic variation of SNP found in homo sapiens, or humans. The chromosome of the SNP is found in 7:117587799. It is linked to cystic fibrosis and severe cardiac failure. The cause of the disease is linked to the disease-associated codon, in which the AGT strand is swapped for a CGT. To regulate the condition, we use CTFR (cystic fibrosis transmembrane conductance regulator) to encode a member of the ATP-binding cassette. Primer Design and Testing We ran two primer tests at the website, the first one using the non-disease primer and the second using our primer with the AGT codon swapped for the diseased CGT codon. The first one yielded a positive result and gave us a match, whereas the second test did not. This is likely because we are changing the strand at one part, which will not give us a proper match. That, and it is a diseased codon, which is going to mutate randomly at the AGT sites.
| ||||||||||||||||||||||||||||