BME100 s2014:W Group4 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR TEAM
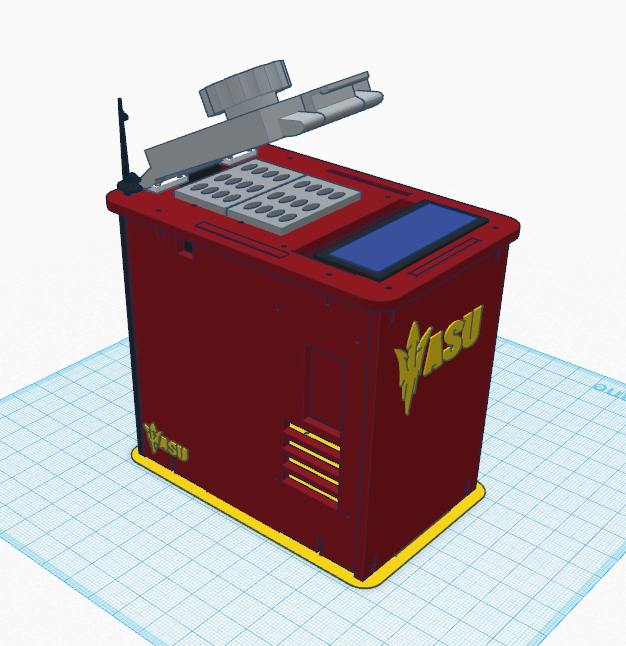
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD TinkerCAD is a website designed to make 3D modeling easy and available for everyone. Using the training/intro tutorial on the website, we were able to familiarize ourselves with the tools available. In addition, in collaboration with the website www.thingiverse.com, we were able to import the files provided by Dr.Haynes in order to build a skeleton of a PCR machine. Having a skeleton of the PCR machine allowed us to personalize it to our liking and incorporate our ideas.
The new and improved PCR machine has a couple of key features that improve the accessibility, safety, customization and efficiency of the device. The new lid is easier to open and has a torque controlled tightening mechanism that makes it simpler to access the wells. The Bluetooth and chargeable power source makes for a cable-less device,turning trip hazards into a thing of the past. By ordering the device online you are able to customize the appearance of the PRC machine, making it unique to your company or university. The addition of extra PCR well tubes and the Cellular PCR control App adds efficiency and freedom for any test possible.
Feature 1: Disease SNP-Specific Primers[Instructions: This information will come from the exercises you did in PCR Lab B.] Background on the cancer-associated mutation Based on searching rs237025 in the database of short genetic variations in the NCBI website, we found that this variation is found in Homo Sapiens. The variation in the gene is located on chromosome 6:149721690. No specific clinical significance was listed. This SNP is associated with the SUM04 and TAB2 genes and is linked to type I diabetes. In this SNP, the non-disease allele contains GTG but a change in the first G position is linked to the disease. The disease-associated allele contains the sequence ATG. The SNP is located at position 149721690. The disease-allele-specific forward primer ending with the nucleotide from the disease-associated allele is AACCACGGGGATTGTCAGTG. A PCR reaction needs two primers to amplify DNA so the position of the reverse primer 200 bases to the right of the disease SNP 149,721,890 and is AGTTTTCTAATTGAGAATGC.
Primer design
How the primers work: Based on the way the disease-specific primer is designed, the reverse primer will bind 100% to its complimentary template while the forward primer will bind 100% to a complimentary disease SNP-containing template. If either the forward or the reverse primer can't bind 100% then PCR is unable to occur. If the template contains the non-disease allele, the primers will not exponentially amplify DNA.
Feature 2: Consumables KitWe will package our consumables in a box that is full of vibrance and color. If the customer orders a design corresponding to their PCR machine, the box will correspond to this theme. It will grab the attention of all who pass by and will be lightweight and come with a convenient handle on top for easy transportation. Within the box, there will also be a tube rack with a covered section for the light-sensitive SYBR green material. Feature 3: Hardware - PCR Machine & FluorimeterOur newly designed PCR machine is much more efficient and easy to use, without any of the hassle. Instead of wires connecting the machine to the OpenPCR program on the computer, there is a bluetooth connection to both the computer and an app for a smartphone. This allows for multitasking while performing DNA replication. One can check the status of the replication by lookin gat their phone. There will also be a gas-cap-like mechanism that prevents the overtightening of the lid that could crush the samples on the PCR machine. The fluorimeter in our system is redesigned to be more stable. There will be a more concrete setup than flimsy plastic trays, and a knob to adjust the height of the table to allow for easy manipulation. No longer will the samples have such an easy time falling off the setup and messing up the data.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachBased on calculation 3, the probability that the patient will develop the disease is around one-half. This means that even if the final test conclusion is positive, it is not certain that the patient will develop the disease; only around half the time will this be the case. For calculation 4, the value is slightly less than one. This means that given a negative test conclusion, it is pretty likely that the patient will not develop the disease, making the negative test conclusion pretty accurate. However, it is not 100% accurate, so false negatives are still possible. The results of these calculations indicate that the PCR machine is not extremely accurate in positively or negatively diagnosing a patient with the disease.
| ||||||