BME100 s2014:W Group1 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |
|
OUR COMPANY
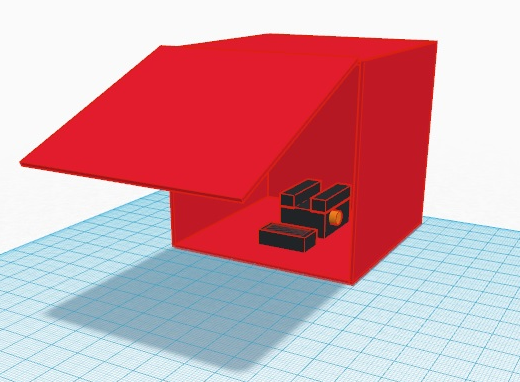
LAB 6 WRITE-UPComputer-Aided DesignTinkerCAD Tinkercad is a simple program to allow us to design items in three dimensions. We simply uploaded a preexisting section of the Open PCR using the tools on the left hand tool bar. At that point we uploaded two of the preexisting fan base. Positioning the fans we prepared to group them before adding fan blades to the design.
Our design is simple. To increase the cooling efficiency of the Open PCR we doubled the number of fans to double the cooling potential. Our PCR machine had issues cooling and this design change would negate the flaw we observed. This allows for more rapid cycle completion and efficient usage of time within the system. Less time will be wasted on inefficient cooling allowing more DNA replication.
Feature 1: Disease SNP-Specific PrimersBackground on the disease-associated mutation The SNP (Single Nucleotide Polymorphism) variation, rs237025, is found in Homo Sapiens on chromosome 6:149721690. This section of DNA has multiple variations, called polymorphisms. The genes associated with SNP are SUM04 and TAR2 which linked to Type I and II diabetes. SUM04 is the Small Ubiquitin-like Modifier 4. Regulation of Nf-Kappa-β-dependent transcription of ILI2B gene control the target proteins sub-cellular localization, stability or activity. An allele alternative form of a gene that ultimatly determines the phenotype. Primer design
When the DNA double helix is separated, the primers will only bind to the specific site on the DNA that contains perfectly complementary nucleotide bases. If the sequences are not exactly complementary, they primer(s) will not bind. This will cause the DNA amplification to fail.
Feature 2: Consumables KitThe consumables will be in a separate box that has the option to be included with the box containing the entire PCR kit. The consumables box will include all of the materials used in the lab. All of the similar parts will be in separate, sealed & labeled bags. The micropipettor will be in its own, smaller box to avoid breakage. The following list shows all the consumables included in the PCR Fandom kit:
Feature 3: Hardware - PCR Machine & FluorimeterIn our new design we add two fans to increase the rate of cooling of the lead, the older design was slower in decreasing the temperature of the polymerise chain reaction which gave us inaccurate data measurements. The new design provides an accurate real time polymerise chain reaction and that helps in copying one strand of DNA. For example, we could copy the HIV DNA because it has one strand. In the design we copied the original fan and added two fans. Although it will consume a higher amount of electricity and have a much higher price, you will get more accurate results and less chance of bias. As mentioned before, PCR Fandom, an adaption of Open PCR, has a more accurate temperature control. The main difference in added fan that will facilitate temperature changes to get more accurate results by reaching the desirable temperatures to duplicate DNA. Both fans will be placed on the left side in the inside of the PCR at the same level. PCR Fandom hardware is made of wood, and will also have an additional layer of stainless steel which will make it more resistant and durable. The fluorimeter will be incorporated to the system by having a more stable structure. This will be accomplished by having the base of the fluorimeter attached to the box so that we can get more precise pictures without the need to adjust it every time a new sample is placed in the ridge. To increase or decrease the height of the system or to move the cell phone base forward or backward, a button mechanism will be implemented. The way in which the box is designed will enable a better background and an easier way to handle the samples. The top part of the box can be opened completely to facilitate samples's positioning.
Bonus Opportunity: What Bayesian Stats Imply About The BME100 Diagnostic ApproachIn order to check the reliability of PCR, we used Bayesian statistics. This calculations describe the sensitivity of the system regarding the ability to predict the disease and the specificity of the system regarding the ability to predict the disease. For calculation 3, the sensitivity was calculated (the probability that a patient with the disease will get a positive result). The value was less than one in a significant way. This means that the probability for a patient with the disease to get a positive, and hence accurate result, is low. On the other hand, for calculation 4, where the specificity was calculated (the probability of a patient without the disease to have a negative result), the result was almost one. The latter means that the chances to get a negative result when the patient does not have the disease are high. Since just one calculations seems to have a high probability, it can be concluded that PCR tests are not always accurate. For other calculations the values were bigger than one. This suggests that in order to get more accurate and reliable results we need a greater range of samples to be analyzed. The deficiency in this test might be due to the lack of samples, errors in the experimental design, the misuse of Open PCR, and not having the best work place to take place the experiment. The limitations of our diagnostic test are noticeable, but still can be improved by making use of the material appropriately and having a more significant number of samples. | |