BME100 f2017:Group4 W0800 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
, and dNTP’s
have the same forward primer and reverse primer
the samples will become cross-contaminated
HEATED LID: 100°C INITIAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 25 Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds FINAL STEP: 72°C for 2 minutes FINAL HOLD: 4°C
Research and DevelopmentPCR - The Underlying Technology Components of the PCR Reacttion The different components of the PCR reaction include template DNA, primers, taq polymerase,and deoxyribonucleotides. The template DNA is the initial strand of DNA used to create a complementary strand in PCR. The primers are short sequences of DNA that are custom built to target the segment of DNA that is desired. These primers attach to the template DNA. The taq polymerase is the enzyme that then builds the sequences and adds the deoxyribonucleotides to form the second strand. PCR Process The initial step is to heat the DNA at 95 degrees for two minutes to asljkdfh;asdhfl;k. After this, a cycle of three steps is repeated until the amount of DNA desired is attained. First the DNA is denatured, at 95 degrees Celsius for 30 seconds. This is when the template DNA separates into single polynucleotide strands. The next step in the cycle is to Anneal the strands at 57 degrees(C) for 30 seconds. This is when the short, primer strands attach to the template DNA. The third step in the cycle is to extend the new strands at 72 degrees(C) for 30 seconds. This is when the taq polymerase enzymes builds the sequences and adds the nucleotides. Then, when the cycle goes back to denature the new DNA formed. After the cycle is repeated as desired the DNA is held at 4 degrees(C). Base Pairing DNA is made up of four types of nucleotides. The four types are Adenine, Thymine, Cytosine, and Guanine. These bases are paired by hydrogen bonding and Adenine always pairs with Thymine, and Cytosine always pairs with Guanine. Base pairing occurs during both the anneal and extension steps of the PCR cycle.
SNP Information & Primer DesignBackground: About the Disease SNP A nucleotide is a compound consisting of a nucleoside linked to a phosphate group. Nucleotides form the basic structural unit of nucleic acids such as DNA. Polymorphism is the presence of genetic variation within a population upon which natural selection can operate. The variation rs769452 occurs in Homo sapiens. The chromosome variation occurs in 19:44907853 and its clinical significance is pathogenic. It is also linked to Alzheimer's disease.
APOE stands for apolipoprotein E. Its function is to combine with lipids in the body to form molecules called lipoproteins. An allele is one of two or more alternative forms of a gene that arise by mutation and are found at the same place on a chromosome. The disease-associated allele contains the codon CCG. The numerical position of the SNP is 44907853. Non-disease forward primer: 5'AGCGGCCAGCGCTGGGAACT3' Non-disease reverse primer: 5'TCGCCGGTCGCGACCCTTGA3' Disease forward primer: 5'AGCGGCCAGCGCTGGGAACC3' Disease reverse pimer: 5'GCTAGAACCAGCAGACC3' 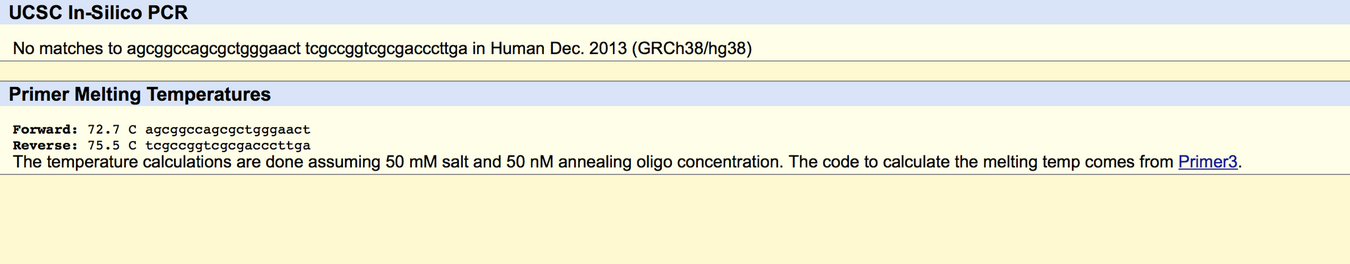 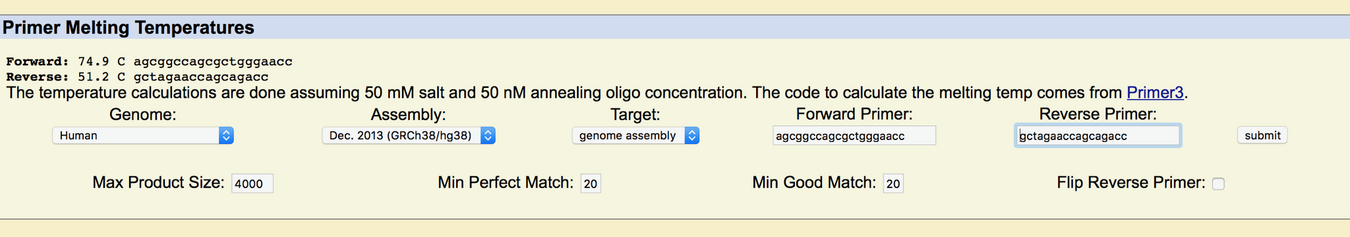 | |||||||||||||||||||||||||||||||||




