BME100 f2017:Group2 W1030 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAMLAB 4 WRITE-UPProtocolMaterials
HEATED LID: 100°C INITIAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 25
FINAL STEP: 72°C for 2 minutes FINAL HOLD: 4°C
Research and DevelopmentPCR - The Underlying Technology Source: PCR http://learn.genetics.utah.edu/content/labs/pcr/ (accessed Oct 18, 2017).
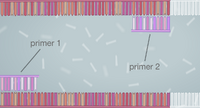
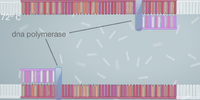
Template DNA is the original DNA used to begin the replication process. The function of the primer is to attach to the base of the targeted DNA and form nucleotides. TAQ polymerase copies the DNA of the targeted area prior to the denaturing of the DNA. Deoxyribonucleotides are the building blocks that DNA molecules are made of. Steps of Thermal Cycling The initial step in the PCR process is raising the temperature of the sample to 95 degrees Celsius, which is almost boiling point, for two minutes. This causes the double helix to separate into two separate DNA strands. The temperature is then lowered to 50 degrees Celsius to allow primers to attach to the base of the targeted strands. Then the temperature is raised to 72 degrees Celsius and the DNA polymerase is activated and locates a primer. After 30 seconds, nucleotides begin to form and extend to replicate another strand of DNA identical to the original. Finally, the temperature is lowered to 4 degrees Celsius to prevent further replication, and is a suitable temperature for storage. DNA Base Pairing There are four types of molecules called nucleotides, A, T, C, and G, that pair up to create DNA. The base Adenine (A) anneals with the base Thymine (T), and base Guanine (G) anneals with base Cytosine (C). Thermal Cycling Base-pairing occurs during annealation and extension. The reason being that annealing requires primers to pair up with the single DNA strands, and they can only be compatible if the bases properly pair up with the correct nucleotides. Extension is when the primers start building identical DNA onto another strand to bond with the original one, which also requires the proper set of nucleotides in order to be built correctly and bond together.
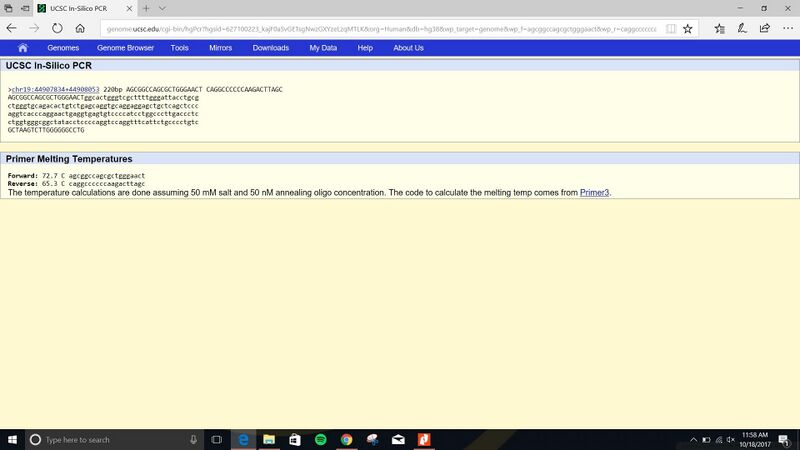
SNP Information & Primer DesignBackground: About the Disease SNP SNP is a single-nucleotide polymorphism that occurs throughout everyone's DNA. SNP's can replace nucleotides ( adenine, thymine, cytosine, and guanine) with a different type of a nucleotide resulting in the person having more than one allele. A nucleotide is a nitrogenous base, s-carb sugar phosphate group and is the building blocks of nucleic acids. The position of the disease is 44907853 and when it is diseased it appears as 44908053. A healthy allele in SNP goes from CTG to CCG when diseased. As you can see for an allele to be diseased a small change occurs which shows that just the smallest error can have serious effects like mutations and diseases.
| |||||||||||||||||||||||||||||||||