BME100 f2017:Group12 W1030 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
Dunder Mifflin Tempe
Dunder Mifflin Tempe LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System BME 100 students were given three replicate DNA samples from two "patients" in order to run a Polymerase Chain Reaction (PCR) in order to detect SNP in the patients. Using pipetting techniques, the DNA as mixed with a primer mix to put into an OpenPCR machine. Next, 8 PCR reaction samples were mixed with the buffers. The samples were mixed with the buffers and then SYBR GREEN solution and a liquid containing double-stranded Calf Thymus DNA (0.25, 0.5, 1, 2, and 5). These samples were then observed using a fluorescence technique. The division of labor throughout the 17 teams and 6 students allowed for the master chart of statistical analysis to be made while each student helped contribute the data and interpretation needed. There were many steps done to eliminate as much error as possible. This was done by using multiple replicates per patient to use as "safety nets" for samples. Along with that, there were multiple PCR controls to observe the samples which allowed the teams to take multiple pictures of the samples for the fluorescence readings, letting students play around with different images within ImageJ calculations for each specific PCR sample. Once all the calculations were done, each group sent them in to make one master statistical analysis sheet which allowed teams to calculate the statistics of the PCR tests and how accurate they were. Overall, it seemed the class had pretty accurate results compared to the doctor's actual diagnoses and there were only three inconclusive results. The experiment seemed to run well amongst the whole class and the only problems that were encountered seemed to be the confusion on calculating the results of the ImageJ calculations. However, after team collaboration, the correct method for calculations and graphing was determined and carried out. What Bayes Statistics Imply about This Diagnostic Approach
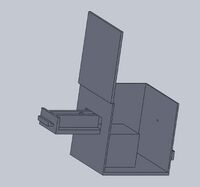
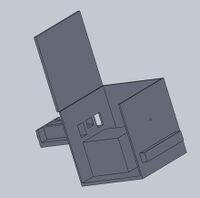
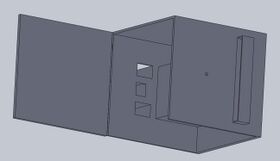
Intro to Computer-Aided Design3D Modeling Our Design
Feature 1: ConsumablesOur consumable kit product list is as follows:
The pipettor tips have been redesigned because the original design allows the tips to frequently fall off of the pipettor. This would cause PCR sample mix and primer solutions to be wasted due to the loss of the pipettor tip. Our company design prevents this problem by including a "bump" on the tip to act as a lock when connected to the pipettor. Feature 2: Hardware - PCR Machine & FluorimeterWe didn't make any changes to the OpenPCR machine because we thought it ran satisfactory. The only thing that we improved on was the fluorimeter system because we had trouble trying to get good pictures for the project, specifically, trying to keep it dark to take the picture. So, we improved on this by adding a separate component that allows for almost complete darkness while imaging the PCR results using the SYBR Green I solution. Increasing the darkness of the environment allows for a better image to be taken of the result, which will thus allow for more accurate data analysis and results. A separate box is to come with the PCR machine, to achieve these high quality images. This box includes a drawer, that the user can easily transfer their sample from the PCR machine to, and once closed, it is completely enclosed in darkness, aside from a tiny hole in the side of the box. This hole in the side of the box creates the desired user interface, allowing the user to put their phone up to the hole and take a picture of their result in almost complete darkness. Of course, the lasers activating the SYBR Green solution will be present in the sides of the box. Lastly, there is a built in and adjustable stand for the user's electronic device used to take pictures. That way, the user does not have to hold their phone and risk shaky hands affecting the outcome of their image, and thus, their results.
| ||||||