BME100 f2017:Group11 W1030 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
OpenPCR program HEATED LID: 100°C INITAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 25 Denature at 95°C for 30 seconds Anneal at 57°C for 30 seconds Extend at 72°C for 39 seconds FINAL STEP: 72°C FINAL HOLD: 4°C
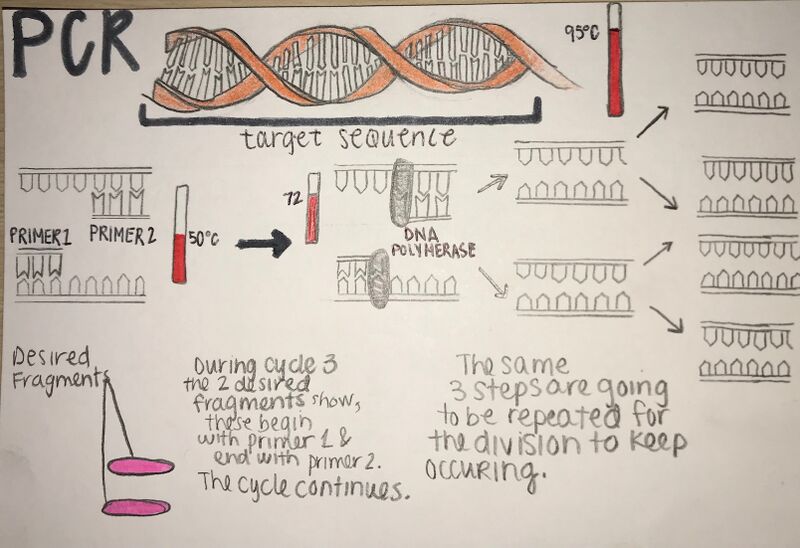
Research and DevelopmentPCR - The Underlying Technology Q1. What is the function of each component of a PCR reaction?
Base-pairing occurs duirng the second and third steps of thermal cycling. The second step, denaturing (which happens at 95°C for 30 seconds) is where the double helix separates and forms two single DNA strands. Base-pairing also occurs during the third step, annealing (which happens at 57°C for 30 seconds) where the primers jump in and crowds the DNA, blocking it from going back to a double helix. Then the primers attach to the single strands.
SNP Information & Primer DesignBackground: About the Disease SNP A nucleotide is a compound consisting of a nucleoside linked to a phosphate group. Nucleotides form the basic structural unit of nucleic acids such as DNA. Thymine, cytosine, adenine, guanine are the four nitrogenous bases in the DNA strand. Each type of the nitrogen bases will be bonded with their matching bond. (T to A and G to C) through hydrogen bonding which will in the end form the double helix of the DNA strand. Polymorphism on the other hand, in biology is the occurrence of different forms among members of a population, or in the cycle of life of an individual or an organism. However, in genetics polymorphism is the presence of the genetic variation within the population, in which natural selection can operate. The gene variation SNP (single nucleotide polymorphism), rs769452, is typically found in homo sapiens or humans, specifically in chromosome 19:44907853. The clinical significance that was listed for this SNP was that it is pathogenic, which means that it is linked to Alzheimer’s disease and subarachnoid hemorrhage in homo sapiens.
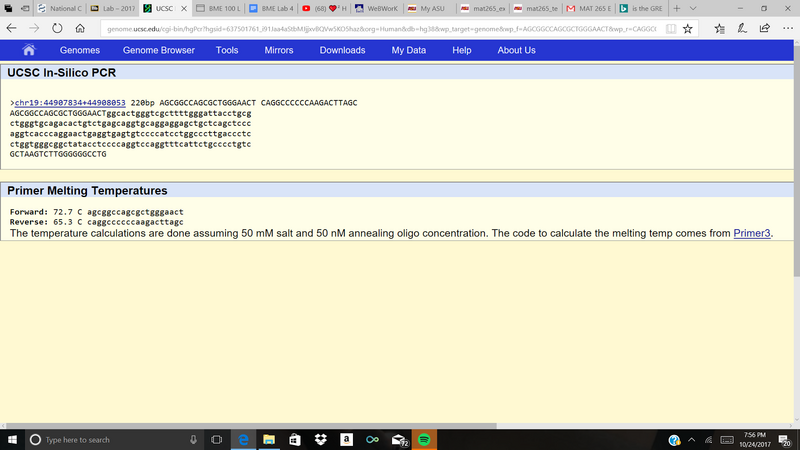
The APOE gene, provides instructions for making a protein called apolipoprotein E, which is what APOE stands for. This protein combines with lipids in the body to form something called lipoproteins which are responsible for bundling cholesterol and other fats and carrying them through the bloodstream. Maintaining normal levels of cholesterol is essential for the prevention of (cardiovascular diseases), including heart attack and stroke. The major alleles are called e2, e3, and e4. An allele is a variant form of a given gene. After looking at the Gene View on the illustrated and moving the cursor over the green flag that shows the SNP, rs769452, the numerical position of this SNP is 44907853. Non-disease forward primer: 5’ AGCGGCCAGCGCTGGGAACT 3’ Non-disease reverse primer: 5’ CAGGCCCCCCAAGACTTAGC 3’ Disease forward primer: 5’ AGCGGCCAGCGCTGGGAACC 3’ Disease reverse primer: 5’ CAGGCCCCCCAAGACTTAGC 3’
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||