BME100 f2017:Group10 W0800 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction Sample List
Heated Lid: 100 degrees Celsius Initial Step: 95 degrees Celcius for two min. Number of Cycles = 25
Final Step: 72 degrees Celcius for two min Final Hold: 4 celcius
Research and DevelopmentPCR - The Underlying Technology Q1. What is the function of each component of a PCR reaction? Template DNA: acts as a guide strand that is copied during replication Primers: attach to sites on the DNA strands that are at either end of the segment you want to copy. They are powerful tools for copying very specific DNA sequences since there is almost no chance they will target the wrong sites. Taq Polymerase: enzyme (protein) that assembles nucleotides into new strands of DNA Deoxyribonucleotides (dNTP’s): is the string of DNA that consists of a nitrogenous base, a deoxyribose sugar and one phosphate group. Q2. What happens to the components (listed above) during each step of thermal cycling?
Q3. DNA is made up of four types of molecules called nucleotides, designated as A, T, C and G Adenine (A): Thymine Thymine (T): Adenine Cytosine (C): Guanine Guanine (G): Cytosine Q4. During which two steps of thermal cycling does base-pairing occur? Explain your answers. During the two steps of thermal cycling, base pairing will occur between the annealing and extension steps. In the annealing step, the DNA is cooled down and the DNA primers bind to the target DNA. After this step, the extension step occurs and the temperature is increased, allowing Taq polymerase to match base primers with the corresponding pair. From this point, the primer pairs are extended and new nucleotide strands are made from the specific DNA sample and are made into numerous copies after each cycle. http://learn.genetics.utah.edu/content/labs/pcr/
SNP Information & Primer DesignBackground: About the Disease SNP Single nucleotide polymorphisms are a type of genetic variation that represent a difference in a single DNA building block called a nucleotide. These commonly occur about once every 300 nucleotides and can act as biological markers. When an SNP occurs in a gene or in the regulatory region near the gene, they may affect the gene's function and play a role in disease. Researchers have found that SNPs are useful for predicting a person's risk for developing certain diseases, responses to a certain drug or environmental toxin, and other phenotypic characteristics. An allele is one or two more alternative forms of a gene that arise bu mutation and are found at the same place on a chromosome.
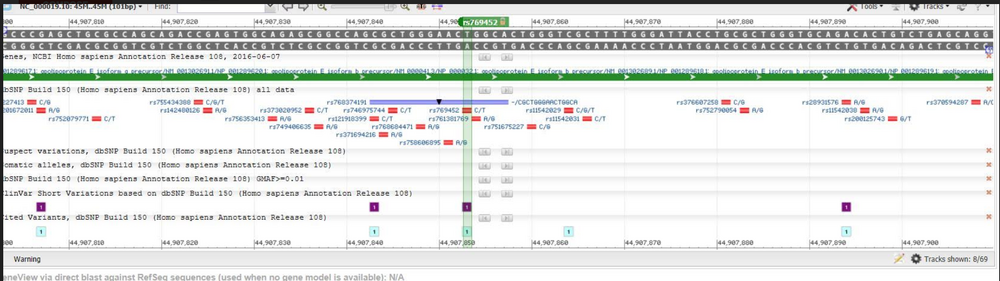
This species is a variation found in Homo Sapiens. The chromosome that the variation is located on is 19:44907853. The clinical significance is pathogenic. Alzheimers disease and strokes are the conditions associated with this SNP. The disease-associated allele contains the codon CCG. The numerical position for SNP is 44907853. Part 2: APOE stands for apolipoprotein E The APOE function is that it is cause for lipid binding, protein binding, as well as antioxidant activity. An allele is an alternative form of a gene that can arise through mutations and occurs at a specific location on a chromosome (encyclopaedia britanicca) The disease-associated allele of APOE contains the codon CCG. Part 3:
Sources Authors of Encyclopaedia Britannica"Allele" Encyclopaedia Britannica. Retrieved from https://www.britannica.com/science/allele. (Accessed on Ocotber 24, 2017). . | |||||||||||||||||||||||||||||||||