BME100 f2016:Group9 W1030AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
2 and dNTP’s (http://www.promega.com/resources/protocols/product-information-sheets/g/gotaq-colorl ess-master-mix-m714-protocol/)
have the same forward primer and reverse primer
do, the samples will become cross-contaminated
HEATED LID: 100°C INITIAL STEP: 95°C for 2 minutes NUMBER OF CYCLES: 25 Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds FINAL STEP: 72°C for 2 minutes FINAL HOLD: 4°C
Research and DevelopmentPCR - The Underlying Technology
| |||||||||||||||||||||||||||||||||
| Question 1: What is the function of each component of a PCR reaction? | |||||||||||||||||||||||||||||||||
| Template DNA | provides DNA sample for primers to copy from | ||||||||||||||||||||||||||||||||
| Primer | matches the segment of DNA of interest, in PCR one primer attach to the top strand at one end and the other primer attach to the bottom strand at the other end | ||||||||||||||||||||||||||||||||
| Taq DNA polymerase | copies a cell’s DNA before it divides in two. In PCR it attaches itself near the end of the primer and starts adding nucleotides | ||||||||||||||||||||||||||||||||
| Deoxyribonucleotides (dNTP’s) | single unit of DNA, Taq DNA polymerase grabs them and attahcs them to the end of a primer | ||||||||||||||||||||||||||||||||
| Question 2: What happens to the components (listed above) during each step of thermal cycling? | |||||||||||||||||||||||||||||||||
| INITIAL STEP: 95°C for 3 minutes | prepare for the temple DNA split | ||||||||||||||||||||||||||||||||
| Denature at 95°C for 30 seconds | at 95°C separate single DNA molecule into two strands | ||||||||||||||||||||||||||||||||
| Anneal at 57°C for 30 seconds | at 57°C the system is cooled enough to introduce primers to attach to template DNA strands and copy them | ||||||||||||||||||||||||||||||||
| Extend at 72°C for 30 seconds | Add Taq DNA polymerase to make copies of the template DNA from primers, and Taq DNA polymerase works the best at 72°C | ||||||||||||||||||||||||||||||||
| FINAL STEP | 72°C for 3 minutes: let Taq DNA complete the process | ||||||||||||||||||||||||||||||||
| FINAL HOLD 4°C: | cool down the system so the newly formed DNA strands can bind to each other by hydrogen bonding | ||||||||||||||||||||||||||||||||
Question 3 - DNA is made up of four types of molecules called nucleotides, designated as A, T, C and G. Base-pairing, driven by hydrogen bonding, allows base pairs to stick together. Which base anneals to each base listed below?
Adenine (A): Thymine (T)
Thymine (T): Adenine (A)
Cytosine (C): Guanine (G)
Guanine (G): Cytosine (C)
Question 4-
Anneal: In the denature step the DNA double helix is separated by the high heat. In the anneal step the solution is cooled to a temperature of 57 degrees Celsius where primers can pair (using base pairing) with the desired section of DNA before the double helix is reformed.
Extend: Once the primers have paired with the DNA the solution is heated to 72 degrees Celsius where DNA polymerase can begin to add nucleotides (through base pairing) to the end of the primer.
SNP Information & Primer Design
Background: About the Disease SNP rs35530544 is an SNP found in Homo sapiens. It is located on chromosome 4:113367751 and affects the gene ANK2. It's clinical significance is pathogenic, which means that the mutation is induced by pathogens. Since this mutation is a non-synonymous substitution, it can result in the inability to produce certain proteins. Specifically, people with this SNP can experience cardiac arrhythmia.
Primer Design and Testing
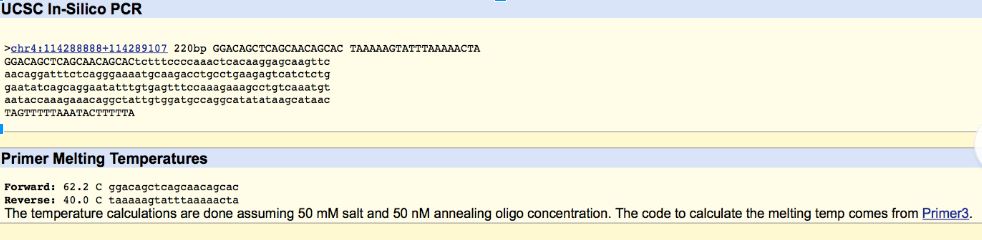
By looking at a model of the SNP and surrounding DNA the primers needed to cut this section of DNA could be read. The base that changes for disease and the 20 base pairs to the left of it make the forward primer. The primer in someone without the disease would look like this 5’- GGACAGCTCAGCAACAGCAC -3’. By moving 200 bases to the right down the DNA we find where our reverse primer will begin. This in on the bottom strand of DNA and is also 20 bases in length. It is 5’- TAAAAAGTATTTAAAAACTA -3'. By changing the single base at the end of the forward primer (C to A) we have our disease primers.
Disease forward primer (20 nt): 5’- GGACAGCTCAGCAACAGCAA -3' Disease reverse primer (20 nt): 5’- TAAAAAGTATTTAAAAACTA -3'