BME100 f2016:Group16 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
, and dNTP’s
have the same forward primer and reverse primer
PCR Reaction Sample List
-Initial step: 95°C for 2 minutes -Number of Cycles: 25
-Final step: 72°C for 2 minutes -Final hold: 4°C
Research and DevelopmentPCR - The Underlying Technology Primers in a PCR reaction are designed to match the DNA segment that is wanted to be copied, the two primers attach to the top and bottom of the segment of DNA being copied. Template DNA is the DNA gene that is being amplified in the PCR reaction. In PCR reaction the Taq polymerase is the enzyme that copies the DNA segment. DNA, deoxyribonucleotides, in a PCR reaction provides the building blocks for replicating, and creating new DNA. Q2: What happens during each step of thermal cycling? In the initial step, the lid is heated to 95°C because that is the temperature at which DNA denatures. The samples are held at 95°C for 30 seconds allowing the DNA to denature. The double helix unravels into two separate stands. At 57 degrees Celcius, the annealing process occurs. During this process, the primers bind to the DNA; one primer binds to each strand. The extension, or elongation step, occurs at 72°C. During this step the tap polymerase binds to the primer and begins to replicate the DNA. Only a portion of the DNA will be replicated. After each chain reaction the portion of the DNA replicated will become smaller until only the target gene sequence is being replicated. In the final step the sample is held at 72°C to ensure that replication has been completed. Finally, the sample is held at 4°C. This is when the DNA strands rebind to form the double helix. Q3: Base pairs Adenine (A) will anneal to thymine (T), while cytosine (C) will anneal to guanine (G). Both of these processes form what are known as base pairs. Q4: Base-pairing during thermal cycling Base-pairing occurs during the steps at which the sample is at 57°C and 72°C of the thermal cycling process. At 57°C, primers bind to the template DNA stand by forming hydrogen bonds to an appropriate section of the template strand according to the corresponding bases of the primer. At 72°C, the Taq polymerases binds to the primers and attaches the appropriate nucleotides to it by creating base-pairs with the template strand.
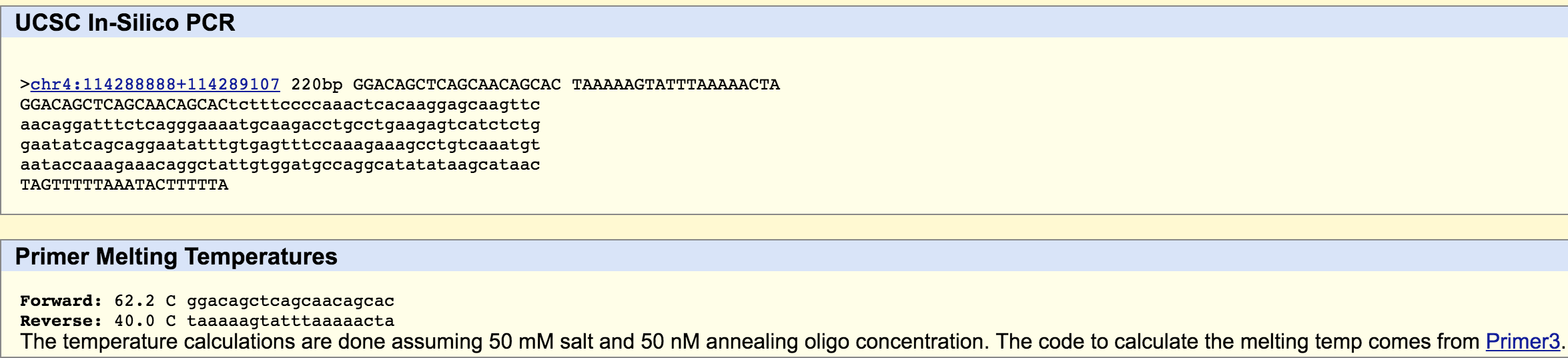
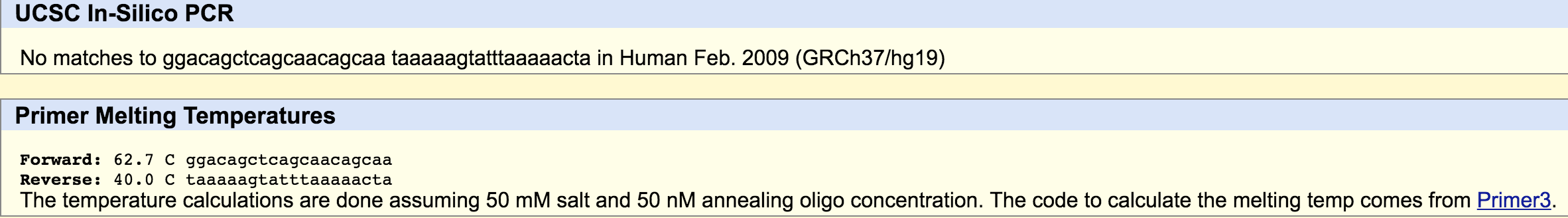
SNP Information & Primer DesignBackground: About the Disease SNP A nucleotide is a compound of a molecule bounded to a phosphate group. These compounds form the basic structural units of nucleic acids. A polymorphism is a variation in certain genes. The SNP rs35530544 is found in Homo Sapiens and is located on the chromosome 4:113367751. It has a pathogenic clinical significance and is linked to a condition called type 4 long QT syndrome, which is an inherited cardiac arrhythmia. This SNP is located on the ankyrin 2, neuronal (ANK2) whose function include; ATPase binding, cytoskeletal adaptor activity, and enzyme binding. An allele is a variation of a gene found at a certain poinr in the chromosome. The non-disease allele of this gene contain the codon "CTC", but in the diseased allele the first "C" is replaced with an "A" resulting in the codon "ATC". This "C" to "A" replacement is the single nucleotide polymorphism (SNP) that occurs at the numerical position 113367751. Primer Design and Testing The numerical position of the SNP we are evaluating is 113367751. The non-disease forward primer is: 5' GGACAGCTCAGCAACAGCA 3'. This primer was determined by writing down the sequence of base pair 20 before the numerical position of of the nucleotide where the SNP occurs. It should end with the base pair at that position. The non-disease reverse primer is: 5'TAAAAAGTATTTAAAAACTA 3'. The reverse primer was determined by reading the bottom (3' to 5') strand of DNA at a point that is 200 base pairs away form the position of the SNP nucleotide (113367951). The strand was read from right to left to, starting with the nucleotide at 1133677951. The disease forward primer (5' GGACAGCTCAGCAACAGCAA 3') is the same as the non-disease forward primer except the final base pair is an "A" not a "C" because of the single nucleotide polymorphism. The disease reverse primer (5'TAAAAAGTATTTAAAAACTA 3') is the same as the non-disease reverse primer.
| |||||||||||||||||||||||||||||||||