BME100 f2016:Group13 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
PCR Reaction List
OpenPCR program
Research and DevelopmentPCR - The Underlying Technology
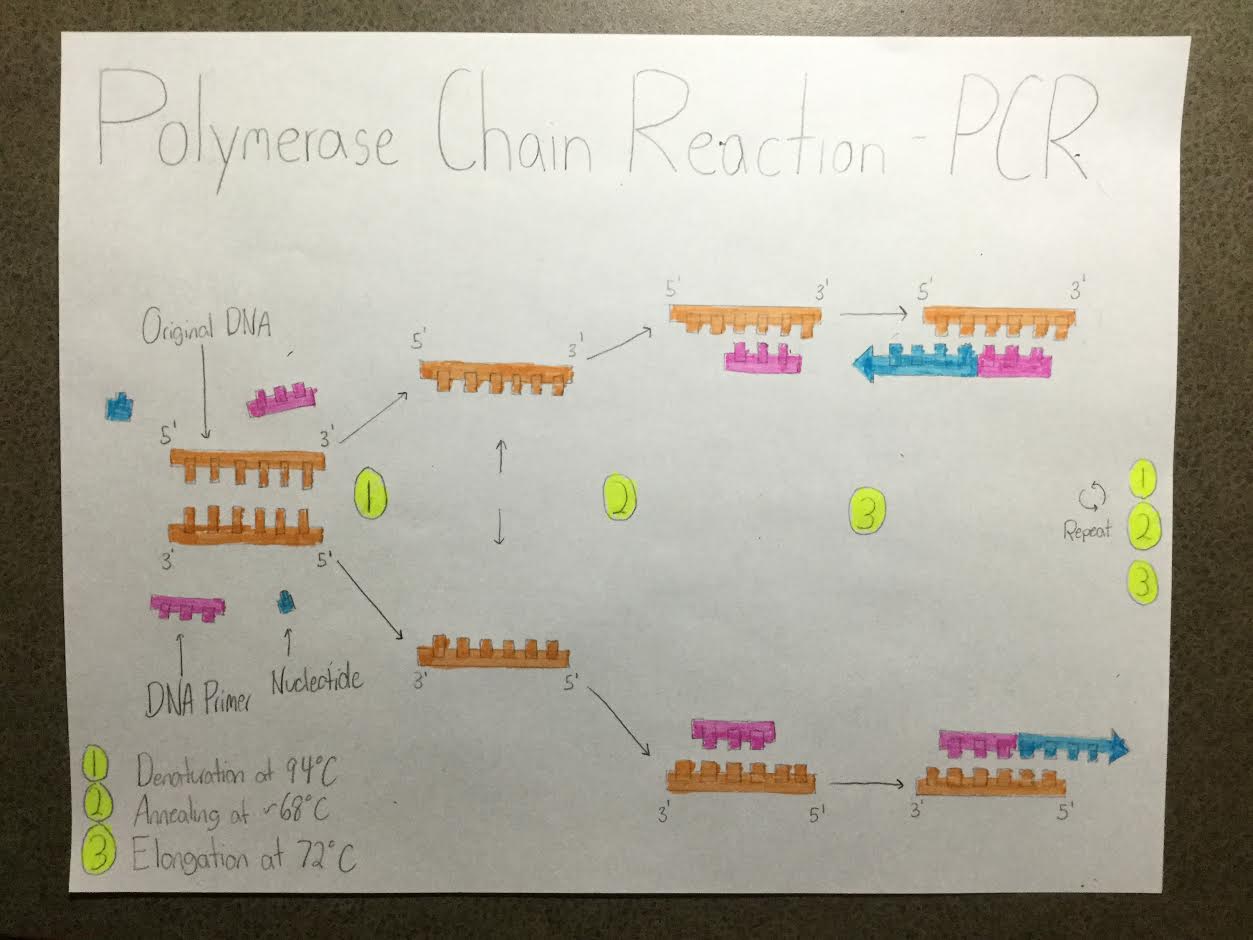
A polymerase chain reaction (PCR) is the rapid method by which specific sections of DNA containing genes of interest can be made and collected from a single complete piece of DNA. A PCR requires four components: Template DNA, Primers, taq Polymerase, and Deoxyribonucleotides (dNTP’s). Template DNA is just as isn’t name indicates. It is the original complete piece of DNA that is being copied in order to create the smaller sections of DNA for study. The primers are small sections of DNA that attach to the template DNA and work as an indicator to the taq Polymerase what section of the DNA needs to be replicated. The primer is designed specifically for the section of DNA containing the gene that is being studied. Taq Polymerase is what attaches complementary nucleotides to the template DNA to create a complete paired section of DNA. dNTP’s are the ‘loose’ nucleotides of building blocks of DNA that are bonded to the exposed nucleotides of the template DNA.
PCR does not simply occur once the four components are placed together in solution, several cycles of differents heat at different times are required to perform a PCR. The initial step is begins at 95°C for 3 minutes. During this time and at this temperature the template DNA strands will straighten out looking like a latter opposed to DNA’s standard shape of a double-helix. At the same temperature of 95°C for 30 seconds after the DNA is straightened it is denatured, meaning that DNA is nucleotide bonds are broken and the DNA is separated. The temperature is then reduced to 57°C for 30 seconds. Over this time the the cooling allows the engineered primers to anneal or attach to the appropriate binding site on the template DNA. Once the primers have bonded to the template DNA, the temperature is increased to 72°C for 30 seconds, at which time the taq polymerase will move forward along the template DNA bonding the appropriate nucleotides the the exposed nucleotides of the template strand. These same step are repeated over several cycles for several hour to obtain an adequate amount of DNA segments for studying. Once the final PCR occurs the DNA is held at 72°C for 3 minutes to ensure that the DNA strands will remain fully extended. After all this is completed the DNA held at 4°C while it is being tested and studied.
The A base stands for Adenine which bonds to the T base, Thymine. Cytosine is the name of the C base and it will bond with Guanine, which is the G base. During PCR there are two step in which base-pairing will occur. The first step is when the temperature is reduced to 57°C for 30 seconds. This is the time when the primer will anneal to the template DNA. The second is when the taq polymerase binds to the primer and extends forward bonding the ‘loose’ nucleotides to the exposed nucleotides. This bonding occurs when the temperature is increased to 72°C for 30 seconds.
SNP Information & Primer DesignBackground: About the Disease SNP To better understand more about this disease SNP there are some general words that need to be understood:
This particular Disease SNP is found in humans, latin name Homo Sapiens. The variation is located on chromosome 4. The Clinical significance lists this SNP a pathogenic, which often times means that it causes a disease. Research on this particular SNP has linked it to Cardiac Arrhythmia Syndrome.
It is important to understand the DNA sequence of the SNP to design a primer to isolate it. An allele is the alternative forms of a genes formed from a mutation that is located in the same position in a chromosome. The disease-associated allele contains ATC opposed to the CTC of the non-disease-associated allele. This SNP is located at 113367751.
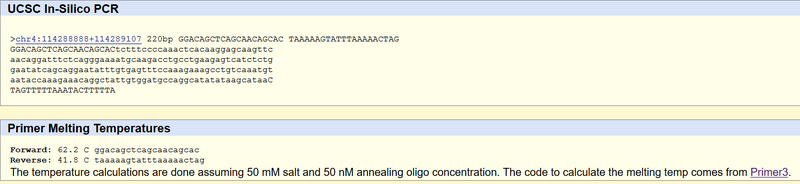
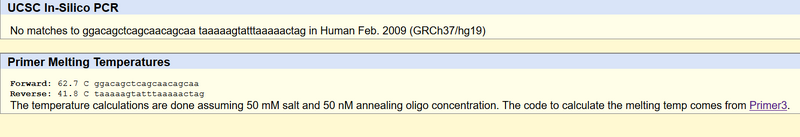
Non-disease forward primer (20 nt): 5’- GGACAGCTCAGCAACAGCAC The numerical position exactly 200 bases to the right of the disease SNP is 113367951. Non-disease reverse primer (20 nt): 5’- TAAAAAGTATTTAAAAACTAG Disease forward primer (20 nt): 5’- GGACAGCTCAGCAACAGCAA Disease reverse primer (20 nt): 5’- TAAAAAGTATTTAAAAACTAG
The Non-Disease Primer Test Results
The Disease Primer Test Results
| |||||||||||||||||||||||||||||||||