BME100 f2016:Group13 W1030AM L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR COMPANY
Our Brand Name LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In order to test the patients for the disease-associated SNP, thirteen groups of five students diagnosed a total of twenty-six patients. Therefore, each group diagnosed a total of two patients. Precautions were taken to prevent the amount of error in each experiment for each patient. This included each group testing each sample three times. This made it so that if there was an error in a trial, it would less heavily weigh the results. By performing three trials, the groups were more confident in deciding if a patient was negative or positive. In order to decide whether the patients were positive or negative, their results were paired up with the negative and positive controls. If the results matched up to the negative control, then the results were negative. If the results matched up with the positive control, the results were positive. In addition to these precautions taken, while using ImageJ each group took controls by measuring the background of the image. This precautionary measure helped to eliminate differences in amount of exterior light in the picture that could possibly skew the data. Moreover, the groups examined multiple pictures of each drop in order to get an accurate representation of the Sybr-green dye in the solution. This technique ensures a more accurate reading of the amount of dye reacting in the solution because it can account fluctuations in the amount of exterior light and blurriness. In total, these precautionary measures were designed to aid in diminishing the level of error in our calculations in order to ensure the most accurate results.
Conclusions one and two imply that the individual PCR replicates are a fairly reliable way to conclude if a patient has the disease SNP or not. This is because the probabilities calculated were close to 100% accuracy. The accuracy for determining if a patient is negative appears to be higher because it is very close to 100%. The positive percent, in conclusion one, is closer to 75% accuracy than 100%, so it can be understood to be less accurate based on the testing done.
Conclusions three and four were both relatively inaccurate compared to calculations one and two. Conclusion three, the probability of patient being positive and correctly diagnosed, is relatively inaccurate, scoring below 50%. Meanwhile, conclusion four, the probability of the patient being negative and correctly diagnosed, is significantly more accurate, closing in on about 75% accuracy. This trend hints at the possibility that the test is more likely to fail and report the patient as negative, not having the SNP. This technique appears to be lacking in accuracy, something is pertinent to the diagnosis process.
A large part of the error can be attributed to the equipment used and human error. On one hand, the machines that conducted the heating and cooling lacked accuracy, with some of the machines working incorrectly or slowly. On the other hand, cross contamination and allowing the Sybr-green dye to be prematurely exposed to light could have hindered the accuracy of the data. Being that this was a first opportunity from many people to pipet, cross contamination can be expected until proper technique is achieved. In addition, while tin soil was placed over the Sybr-green dye, the situation was less than ideal. The tin foil did not fully cover the light, possibility resulting in a potential for the dye to prematurely react or react incorrectly. Intro to Computer-Aided Design3D Modeling
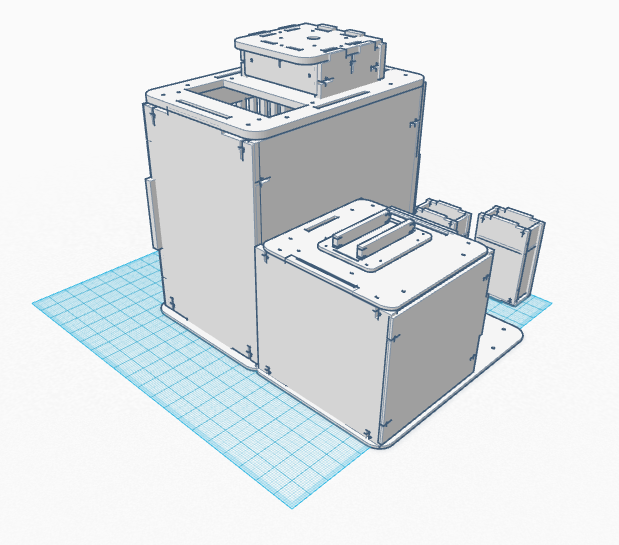
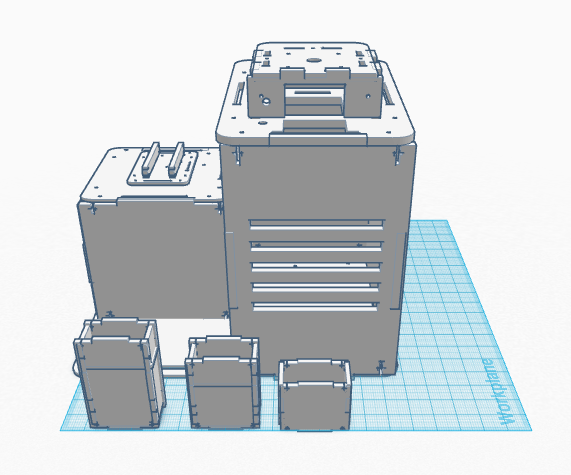
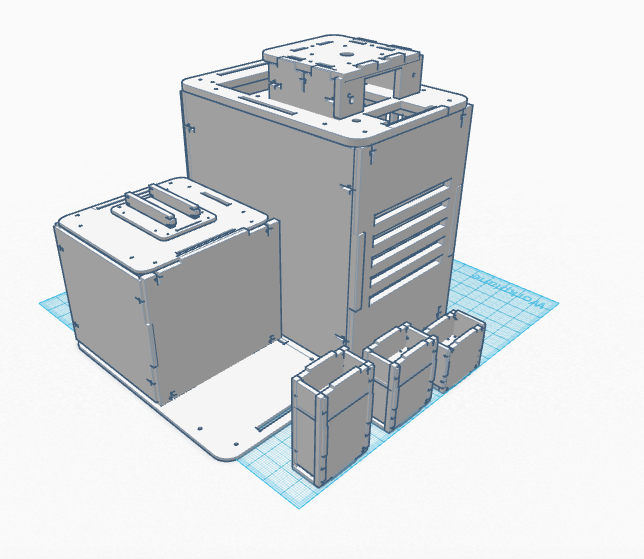
Our Design
In our design, we address the cyber green sensitivity to light by making PCR tubes with a black coating around them. Additionally, we are combining the PCR machine and the flourimeter into one functioning device. There is also a flexible extension off the machine that serves to prop up one's iphone. This allows the pictures to be taken more easily and in higher quality. Feature 1: ConsumablesThe most important consumables are the reagents such as the PCR mix and the primers, which will be included in the product. The plastic tubes for the sample will be darker in order to keep light out of the sample, which can be difficult using the standard test tubes. We will not include a pipette since this is a generic product that is easy to acquire. Feature 2: Hardware - PCR Machine & FluorimeterThe new PCR machine and Fluorometer all in one machine aims to eliminate the clutter of the current system. The system will be packed in a box which can double as the cover for the Fluorimeter component. The kit will also include a pipet, pipet tips, tube holders, and the PCR tubes. The all in one machine will also include multiple size phone stands that clip into the base of the Fluorimeter component. This allows for the stand to be sturdier than competing bundles while keeping the distance between the phone and the drop constant. In addition to this, the SYBR GREEN dye will have designated tubes eliminate light interference. This will improve the accuracy of the light sensitive marker diminishing the amount of time the dye in exposed to light.
| ||||||