BME100 f2016:Group12 W8AM L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||
|
Group 12, Best Group
PCR LABProtocolMaterials
PCR Reaction Sample List
OpenPCR program
-Denature at 95 °C for 30 seconds, Anneal at 57 °C for 30 seconds, and Extend at 72 °C for 30 seconds
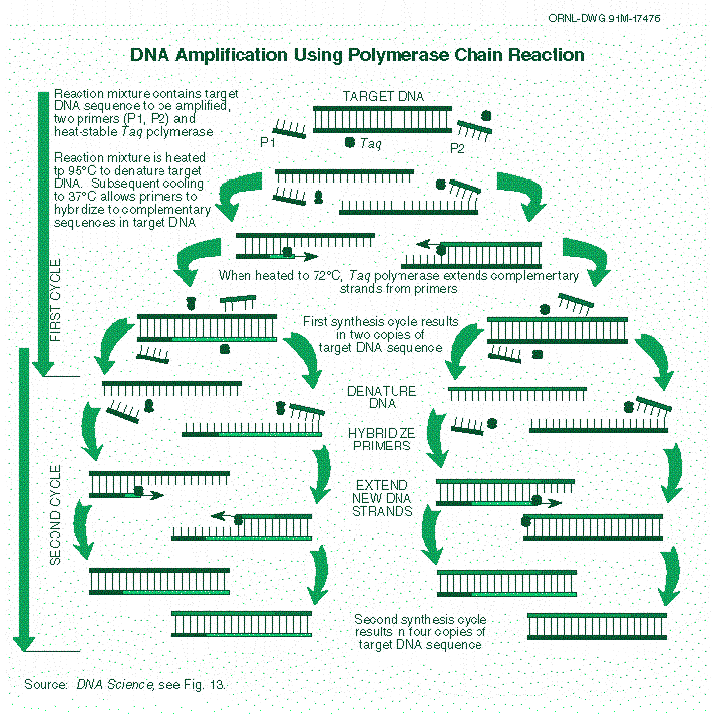
Research and DevelopmentPCR - The Underlying Technology Function of each component of a PCR reaction:
What happens to the components during each step of thermal cycling
DNA is made up of four types of molecules called nucleotides. Which base anneals to each base?
During which two steps of thermal cycling does base pairing occur?
(http://www.contexo.info/DNA_Basics/polymerase_chain_reaction.htm)
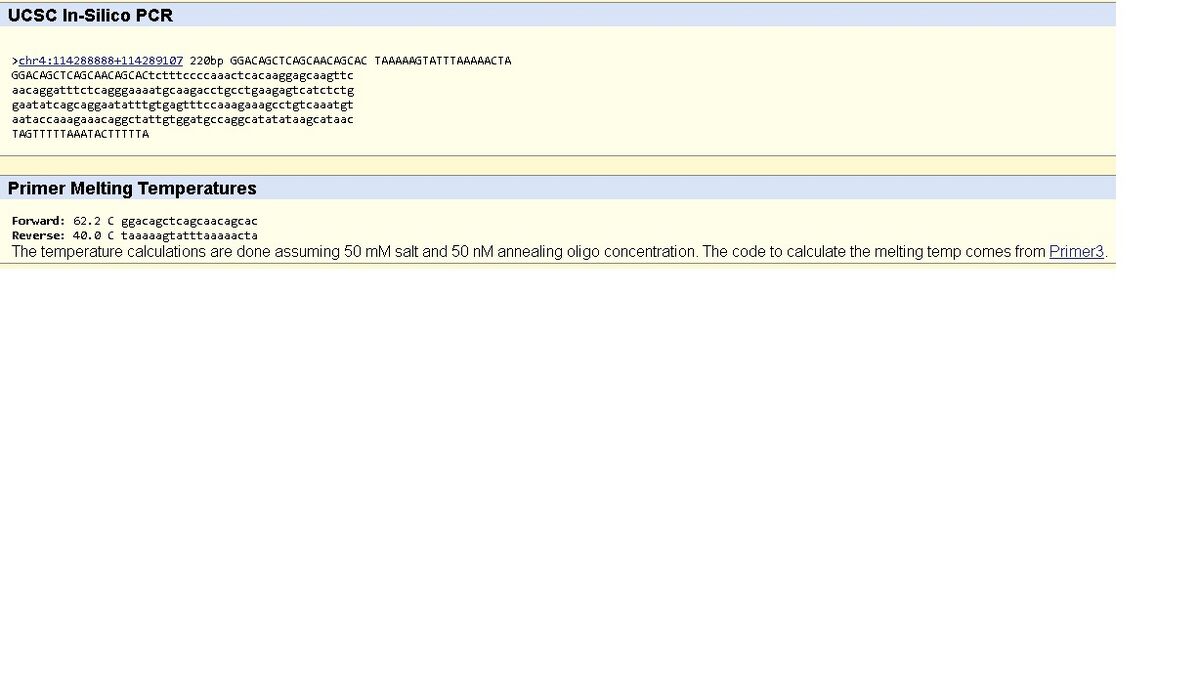
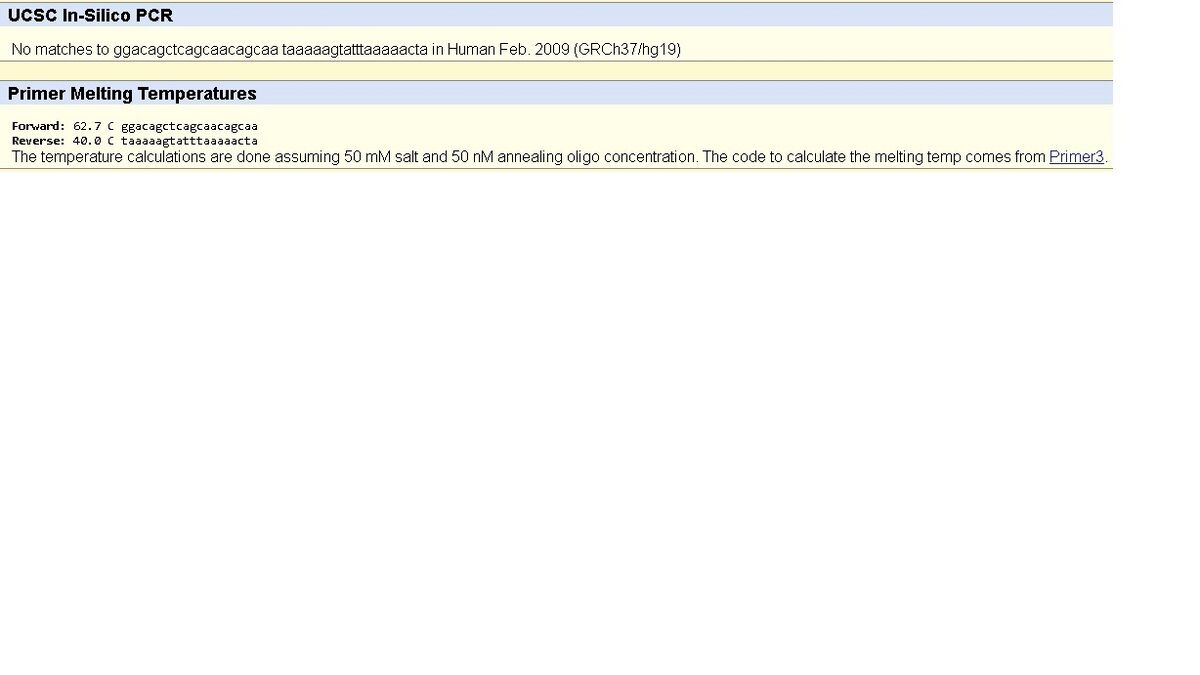
SNP Information & Primer DesignBackground: About the Disease SNP SNPs or single nucleotide polymorphisms are among the most common type of genetic variation amongst the population of the world. An SNP will generally appear about once every 300 nucleotides in someone's DNA. While most SNPs are harmless, some can be directly tied to a specific disease due to the fact they can alter a gene's function. Scientists are able to use SNPs to act as an indicator that a person has a specific disease, making diagnostics a lot easier. For example, researchers have been able to use SNPs to predict a person's likely response to individual drugs, susceptibility to environmental factors, and even a person's risk of certain genetic diseases. The specific SNP our group is looking at is found in homo sapiens (humans). It is located on the chromosome 4:113367751. This specific SNP is linked to pathogenic factors, more specifically, it is closely linked to cardiac arrhythmia. Sources: https://ghr.nlm.nih.gov/primer/genomicresearch/snp A short while ago, a test was done to find which SNP's within genes could be related to what caused the Arrhythmia. The study had 273 people, and some of which had died from from cardiac death, and 20 that were revived. The study showed that NOS1AP and KCNQ1 are two of some the SNP's that are related to cardiac arrhythmia, and are the highest risk factored ones too. http://www.sciencedirect.com/science/article/pii/S1547527113011429 Primer Design and Testing The non-disease primer that was created consists of two primers which amplify the DNA. It consists of both a forward and a reverse primer. The non-disease primer is correct because it resulted in 220bp. The diseased primer that was created consists of two primers which alter the amplification of the DNA. It consists of both a forward and a reverse primer. The diseased primer does not work correctly because it shows a result of no match. the "no match" is due to the change in the non-diseased primer from "C" to "A".
| |||||||||||||||||||||||||||||||||