BME100 f2014:Group9 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||
|
OUR COMPANY
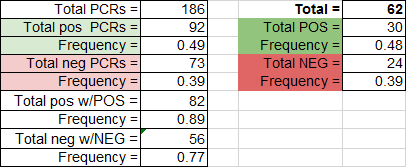
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In the BME100 lab, multiple teams of students diagnosed 68 total patients to analyze their DNA to determine if they had a specific disease. Each group of students received all equipment necessary to perform PCR on the two patients, such as the samples themselves, the PCR mix, machine, and pipettor. The procedure for PCR was to combine a specific amount of the PCR mix with the target DNA by using the micropipettor, and place the samples in the OpenPCR machine. Multiple samples for each patient were run in the machine to prevent error. Two groups shared one OpenPCR machine, for a total analysis of 4 samples. Overall, the labor was divided among 34 groups for the analysis of all samples. Once the PCR was completed, the samples were added to a slide with SYBR Green I dye, and placed on a fluorimeter. This allowed the groups to analyze the amount of green fluorescence produced through the use of smartphone cameras and ImageJ software. Other samples with a positive and negative control were also analyzed, so that the patient samples could be compared to the ImageJ results of the controls. Three photos of each sample were analyzed, and the data from ImageJ allowed us to determine if the patient tested positive for the SNP. The results for positive and negative results, along with the determination of the presence of the SNP can be seen in the following:
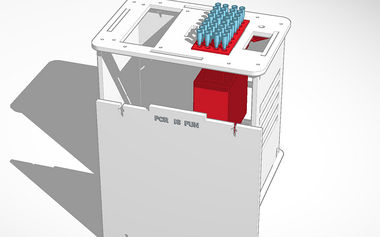
Computer-Aided DesignTinkerCAD TinkerCAD allowed the group to make a 3D visual of the PCR machine using online templates uploaded from www.thingiverse.com. Using simple shapes and ergonomic tools, manipulation of the original design was effortless. Since the dimensions are displayed, dragging the corner of any shape to change the length, width, and depth allowed quick and precise measurements to fit the scale needed. Of all the features, the movable grid plane was the most efficient, allowing the fast placement of objects on any surface of the design.
Our design changed the OpenPCR machine to enable PCR on 32 samples rather than 16. This allows for more tests to be run quickly, maximizing efficiency of PCR. Also, since more samples can be run, one machine can serve the purpose of many, so the costs can be cut down.
Feature 1: Consumables KitThe consumables kit in our improved PCR system will be the exact same as before, except with double the components. Included in the package will be the test tubes, SYBR Green I dye, and PCR mix. A micropipettor tool with tips will be in the packaging as well. A holster for all of the samples will also be included. Feature 2: Hardware - PCR Machine & FluorimeterThe improved PCR machine will be able to run up to 32 samples at once. This allows for testing to be run quickly and more efficiently than before. Also, the improved fluorimeter system will have a cradle and camera in a set position from the sample slide. This will eliminate or marginalize variance in the photographs of the samples, thus improving accuracy of the testing. Also, ImageJ will be easier to run on these photographs because the samples should be in the same position every time.
| |||||||