BME100 f2014:Group5 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System DNA was amplified and analyzed in a open PCR machine from 68 different patients. There were 34 groups of six students each that analyzed two patients for a disease-associated SNP. After the DNA was amplified, three samples were made from each patient so that each group had six samples to test for the disease-associated SNP. The amplified DNA was mixed with SYBR green and analyzed through image analysis of drops in a fluorimeter. In order to limit error in the data analysis, groups only needed to test two patients. Each patient had three tests done with the fluorimeter so that our results were more accurate. In addition, error was reduced because of accurate imageJ analysis in which the oval perfectly matched up with the picture. All the pictures were taken with the same settings and were all focused. There were three pictures per drop as well making 9 total pictures to analyze per patient and 18 total per group. If everything went perfectly in the lab, the PCR conclusion and whether or not the patient had the disease should match up. If the PCR conclusion was “yes”, then the disease would have been present. If the PCR conclusion was “no”, then the disease would not be present. However, this is not the case for the class data as a whole. There were some inconclusive data sets from the PCR, and thus no data was able to support a claim about disease presence. There should be results for all 68 patients. 30 patients had a positive result, 24 patients had a negative result, 8 were inconclusive, and 6 tests failed to provide any data or were not tested at all. Overall, the PCR’s conducted only correctly identified 10 out of 23 patients that actually had the disease 12 out of 45 patients that did not have the disease. What Bayes Statistics Imply about This Diagnostic Approach For Calculations 1 and 2, there is a high reliability of the individual PCR replicates for concluding that a person has the disease SNP or not. Both values for the Bayes Statistic of 1 and 2 were close to 1 which indicated a high probability and connection. For Calculation 1, the probability that a patient will get a positive final test conclusion, given a positive PCR reaction was tested and was high (close to 1). For calculation 2, the probability that a patient will get a negative final test conclusion, given a negative diagnostic signal was tested and was also high (close to 1). A few possible sources of error that caused the numbers not to be 1 are as follows. One, the PCR machine could have not worked completely for some groups. In other words, the cycles and temperatures were not exact, thus the DNA segments were not copied. Two, due to human error, the PCR mixtures could have been messed up during micropipetting. If some of the samples lacked the SYBR Green for the fluorimeter or even the mixture of PCR components, the PCR would not work. Three, we could have accidentally mixed up the patients and the patients we thought we were testing would actually be mixed around. Calculations 3 and 4 were had a low value for the Bayesians Statistics. The value for both was close to 0 which provided a low probability and low reliability of the test. Calculation 3 tested the probability that a patient will develop the disease, given a positive final test conclusion. Calculation 4 tested the probability that a patient will not develop the disease, given a negative final test conclusion. Computer-Aided DesignTinkerCAD TinkerCAD is a web based design software similar to SolidWorks. It comes with preprogrammed shapes and teaches you how to use the program with ease. Our group imported shapes from the original PCR machine design and built an original device. The tools on TinkerCAD were used to make sure the machine was the correct size and shape. Our new device is essentially a slight modification of the old Open PCR Machine.
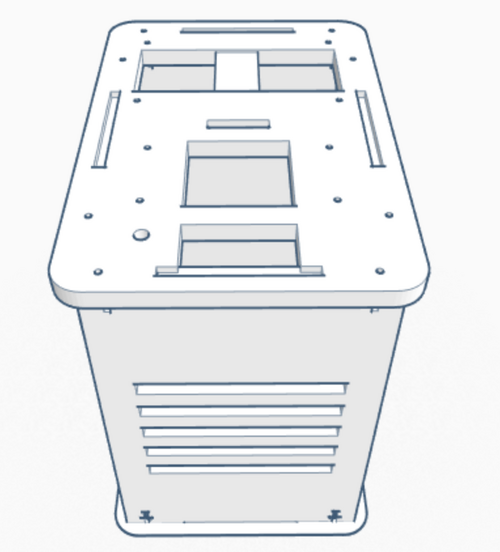
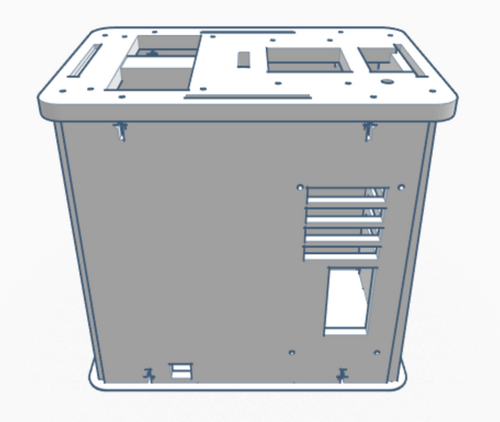
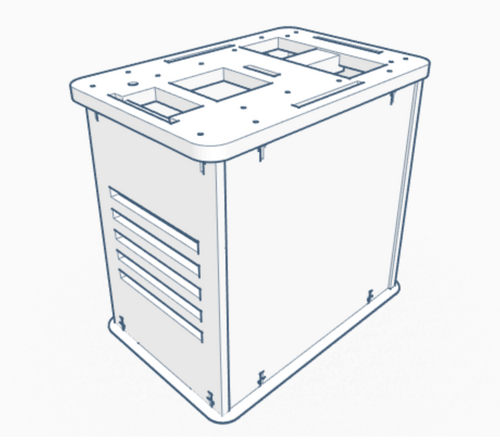
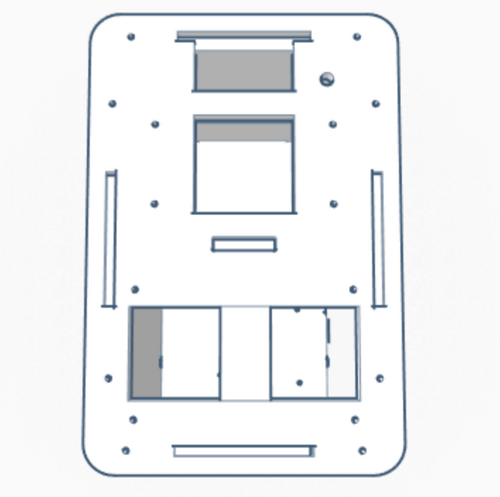
Our Design The group decided to add a second chamber to the machine. This change to the design allows for twice the data collection of the original device. The design was choose in order to allow for the machine to be more efficient.
Feature 1: Consumables KitConsumable kit includes:
We also have an option that includes a premixed solution made up of those parts. Because not all PCR is testing for the same disease or required to amplify the same section of DNA, primers will have to be bought separately or created yourself. The main difference between the original PCR machine, which has a hatch on top that 16 test tubes can fit inside, and the upgraded one is that the new PCR machine contains two of these hatches; which doubles the amount of PCR space and also allows different types of PCR's to take place. Feature 2: Hardware - PCR Machine & FluorimeterThe process of amplifying DNA and analyzing it is generally a lengthy one even though there are only two devices to work with, a PCR and a fluorimeter. One way to improve the process is to combine the two devices into one. This device would have the same general look and materials as an original PCR, but it would house two hatches on the top instead of one. This means the machine will still be made of wood and will be rectangularly shaped. Having two hatches allows for more samples to be tested, or it could accommodate different types of samples testing at once. Not only that, but both hatches will be fashioned with a flourimeter, light detection system, as well as the PCR. The SYBR Green will already be mixed with the DNA samples placed in the PCR, that way the flourimeter and light detection system can analyze the DNA, as it is being amplified. The analysis of the DNA can be monitored form screens placed on the side of the machine. The major concern with the current process of DNA amplification and analysis is the time it takes, but with these modifications the time of the process will be dramatically lowered. |
|||||||