BME100 f2014:Group32 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System
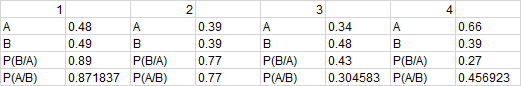
These results were then analyzed using Bayes statistics to calculate the probablity of the results. The final data resulted in 23 positives and 45 negatives however, the data that was given only required 30 positive samples and 24 negative samples. Some of the data was either discarded or blank because groups made errors within the experimental method that biased the results.
The Bayes Statistics approach to this implied that for calculations 1 and 2 the results seemed to be reliable in determine whether or not the patients had the disease SNP or whether they did not have the diseased SNP. The calculations for both were close enough to 1 to be considered alright, but the discrepancies in the data accounted for the large error away from 1. The calculations from 1/2 compared the diagnosis from the fluorimeter and the PCR machine as positive or negative, gave a numerical value for how accurate the PCR readings were. Because these numbers were fairly close to 1 within the error, one can say that the diagnosis method used was fairly accurate. However the discrepancy in the calculations cannot be ignored. The discrepancies were because of idealizations inherent in the theory as well as errors and uncertainties in the experimental method. The camera placement each time in the experiment might have been slightly off more and more each time, thus propagating the error of the experiment exponentially. The box itself was not closed properly during all pictures as well which could explain the discrepancies in the data; light let into the box distorted the images and made the data processing error ridden. The images taken by the phone itself were neither of the highest nor clearest qualities, thus the images themselves had slight distortion by the camera as well. Error within the experiment could have occurred in the human portion as well; during the experiment the samples could have gotten contaminated by oils or foreign material, different materials not supposed to be combined could have been combined or incorrect amounts of material between the tubes could have been placed in the PCR machine causing unsatisfactory results. The experimental method could be wrong as well; not enough sample could have been used or maybe there was not enough SYBR green solution per droplet in order to test for DNA sampling. Incorrect analysis of the results leading to an incorrect calibration curve could have led to incorrect results as well and thus a less reliable method. Calculations 3 and 4 give a direct numerical accuracy of the device used in this diagnostic method for the detection of the disease. The values were not statistically significant because of how small they were and thus one cannot say that this device is extremely accurate at measuring the disease based on the data taken. Because these values were not as good as the first two calculations, this suggests that this experimental method design is wrong or needs significant improvement in order to raise the standards of this device and experimental data. Both of the values calculated for the test were very small and below 0.5 which means that there was less than a 50% chance of detecting the correct diagnosis given the positive or negative test. 1)The calculation of the probability that a patient will get a positive test (SNP), if he/she were given a positive PCR reaction is close to one. Therefore, the PCR can be considered reliable for these calculations within the limits of accuracy. Possible reasons for error:
2)The probability of a patient getting a negative test (SNP) given a negative diagnostic signal is less than the first value yet close to one within limits of accuracy. Possible reasons for error in values:
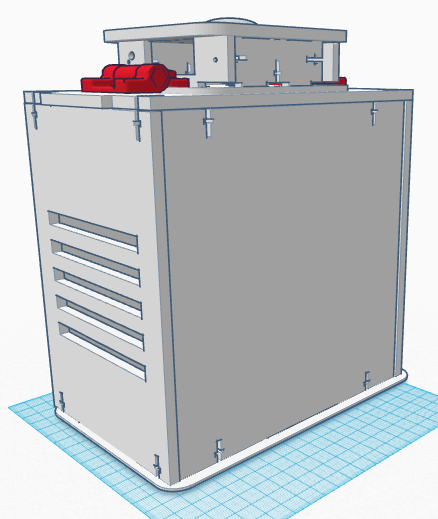
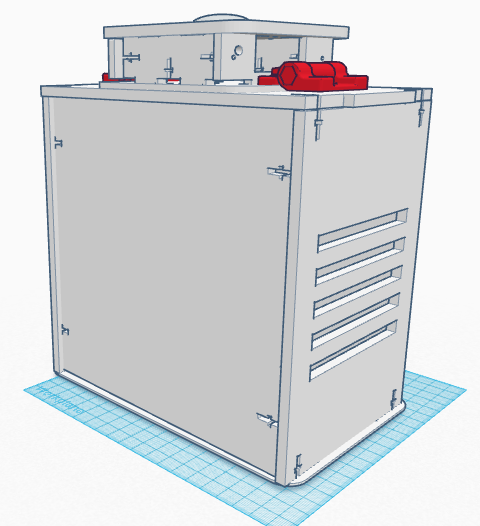
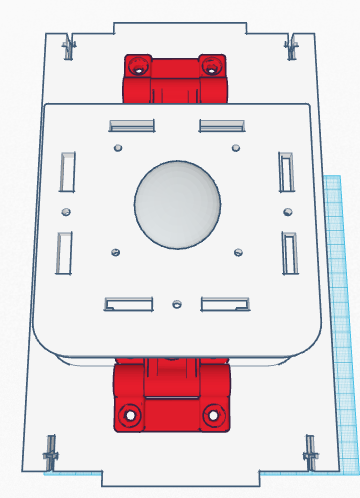
Computer-Aided DesignTinkerCAD TinkerCAD was an online software tool used in order to construct different objects using the preprogrammed objects built into the software itself and combining them into different shapes. TinkerCAD first had a detailed summary and lessons on instruction on the process that one can use in order to design different objects. TinkerCAD was used to build an Open PCR machine using objects imported into the software and then assembling them in the CAD software itself. TinkerCAD software assembled the premade parts quickly and with precision; the shape tool showed small details and sizes of the different objects for ease of assembly and several other tools made TinkerCAD a good software for novice assembly of the Open PCR machine.
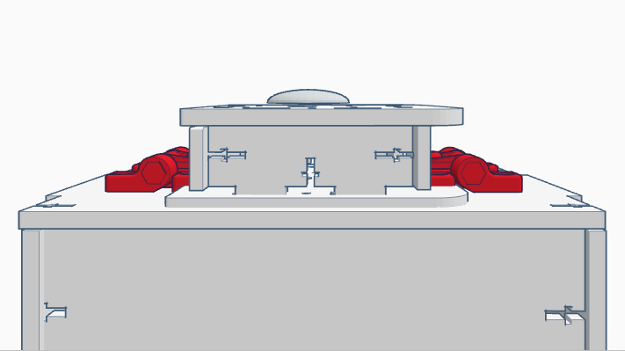
Feature 1: Consumables KitThe consumable kit that came with the original PCR reaction was fairly thoroughly prepared and well kept. The consumable kit that comes with this PCR machine will include the same products such as gloves, the micropipetters, SYBR Green solution, the master mix containing: taq polymerase, dNTPs, Buffer solution and primers; tray tables, micropipet tips and a disposable waste container. The consumables kit had several different problems with it during the experiment that added error and uncertainty to the experiment. The consumables kit that comes with our PCR machine includes an efficiently designed tray table that holds different test tubes of any size within the tray table itself. Furthermore, instead of wrapping the SYBR Green solution in an aluminium foil wrapping to keep it from deteriorating, the consumables kit comes with reaction tubes of SYBR Green that are all black on the outside in order to hold light sensitive solutions. Furthermore several reaction tubes will be included of different colors in order to differentiate between different samples, and on those tubes there will be rough strips to label the samples that are resistant against heat erosion. The tray table itself also has features of attaching to the actual PCR machine for ease of storage and space. The micropipettes have presets on the volume levels within this consumables kit. The micropipettor is now fully digitized so that the volume intake can be inputted electronically and distributed electronically with less human interaction at the press of a single putton. Preloaded pcr material is included in the forms of a master mix with all the materials combined and individual materials for further usage and combination of PCR products. Feature 2: Hardware - PCR Machine & FluorimeterThe PCR machine, while rather effective when the seal atop the machine is fully fastened, can show very skewed results if the hatch fails to be closed due to human error. With the current design, the latch may not fasten all the way because one must be sure to make sure the cap is all the way closed and the screw is completely tightened. Based on group observations of the machine, it is far too easy for this hatch to be improperly shut, thus causing the data to suffer accuracy. The proposed solution to such a problem is to diminish the likely hood of this human error. The group has decided to apply latches to the hatch atop the machine. These hatches will make sure that the top is both completely closed, as well as securely fastened throughout the duration of the PCR machine cycle. As for the fluorimeter, the group found the greatest problem to be that the pictures taken of the droplets were not as accurate as they could be. With the way the front flap works, it is difficult for group members to place the camera inside the fluorimeter. Also, the flap fails to completely close, meaning that excess light is being let in which will affect imageJ results. The group proposed a possible curtain mechanism which will both make the fluorimeter easy to access and close better so as to keep light from entering.
| |||||||