BME100 f2014:Group30 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In the PCR lab experiment, the DNA of 68 patients total was amplified and analyzed for a specific disease indicator in order to predict the group's probability of disease detection. To divide the work for this experiment, 34 groups of 6 people each tested 2 patients to determine the likelihood that either patient would contract the disease. As with any experiment, steps were taken to prevent error. For example, 3 total replicates were tested for each patient as a way to minimize the likelihood that the patient will be misdiagnosed. In addition, the PCR mix had positive and negative control solutions, which helped to make sure that the PCR machines and fluorimeters produced the expected results for the known positive and negative solutions. Regarding the Image J processing, calibration controls of positive and negative solutions were utilized to prevent error in the sense that they provided a baseline for what was supposed to happen in patient samples with negative and positive test results. In other words, the positive control showed the SYBR green, which indicated that the PCR and fluorimeter machines worked to produce expected, controlled results, and it also indicates that other patient samples with the same green color test positive. To increase the accuracy of the Image J processing values, 3 drop images were taken per PCR sample and averaged together. Although the class's final data contained a majority of successful conclusions, there were discrepancies within the data collection that affected the overall Bayesian calculations for the specificity and sensitivity that a patient has a disease. For example, 8 out of the 68 patient results were inconclusive, which means that the calculated Bayesian probability statistics were higher or lower than the actual probability depending on whether the inconclusive results should have been positive or negative. Blank data was also discrepancy within the data collection, as the 6 blank conclusions decreased the total PCR conclusions that were used to calculate Bayesian probabilities. Since the total number of conclusions was decreased due to the number of blank conclusions, the statistics were higher than what they should have been if there had been 68 total conclusions. For example, if 20 out 68 people had a positive conclusion, there would be a higher probability of a positive test conclusion than if 20 out of 62 people had a positive test conclusion. Measures were taken to obtain the most accurate data collection, but there are things like inconclusive and blank test results that can still affect the data analysis.
Calculation one was working with the percentage of positive test results compared to the actual amount of positive PCR reaction for the disease SNP. Overall about half of the PCR tests came out positive. When looking at the ratio of positive test conclusions to the amount of positive PCR diagnostic signal, it is roughly close to 1. This means that the tests that we executed were roughly accurate for detecting a positive diagnostic signal. As for calculation 2 for the negative test results, a little less than half of the test results came out negative. The ratio of negative test results to the actual number of negative signals is very close to one. This shows that the PCR replicates were very accurate for the DNA that was not containing the disease SNP. Overall, the PCR replicates were reliable but more so for the negative reactions than the positive ones. Even though these results were reliable, there were a couple of things that we could have changed to possibly make the test results come out correct more often. For one, on the imageJ detection process, the human error of outlining and capturing the droplet accurately could have had a negative effect on the PCR test results. The SYBR green solution was sensitive to light which was not optimal because we were working in a light environment so even though the vials of SYBR green was sealed by foil, it was subjected to light when we took it out to pipet it onto the slide. Lastly, when using the micropipette, more solution can be drawn in than desired because of the range of motion that the button gives. If you push the button down just a little bit too far, too much of the desired solution will enter the tip which can throw off the results.
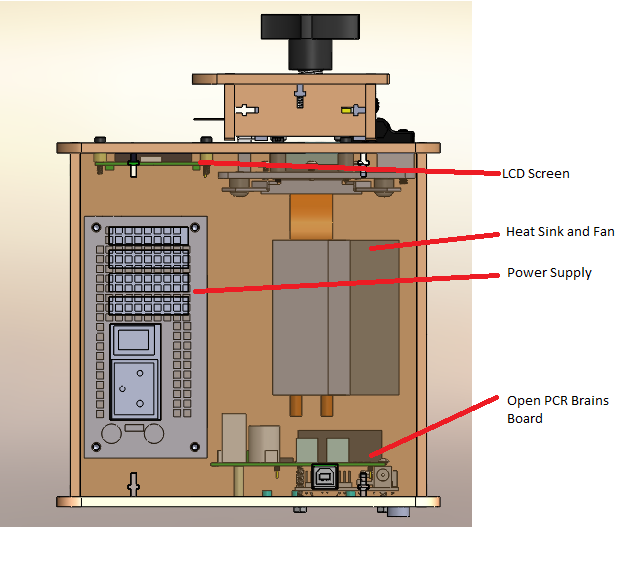
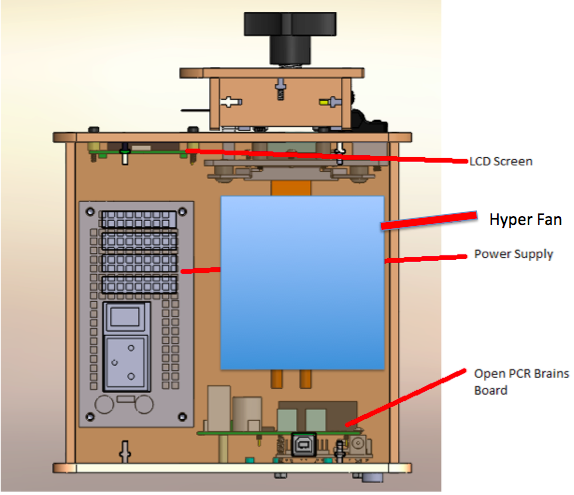
Calculations 3 and 4 have to do with the development of the disease in DNA that has the positive and negative diagnostic signal for the disease SNP. For the positive side of it, the probability that the patient with a positive test conclusion will develop the disease is very low. This implies that the PCR replicates do not show a very accurate prediction on whether or not the patient will develop the disease. If it was more reliable, the percentage of patients who develop the disease would be higher for the amount of positive test results. For the ration of patients who will not develop the disease when given a negative test result is significantly higher. This percentage is more than half. So the PCR replicates are more reliable at detecting the negative signals and predicting that a patient will not develop a disease than detecting positive test results and predicting that a patient will develop the disease. Computer-Aided DesignTinkerCAD Our Design Before Hyper Fan Addition* After Hyper Fan Addition*
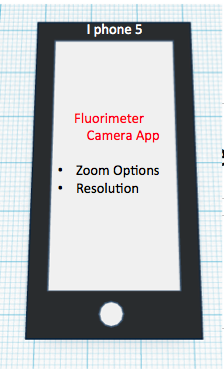
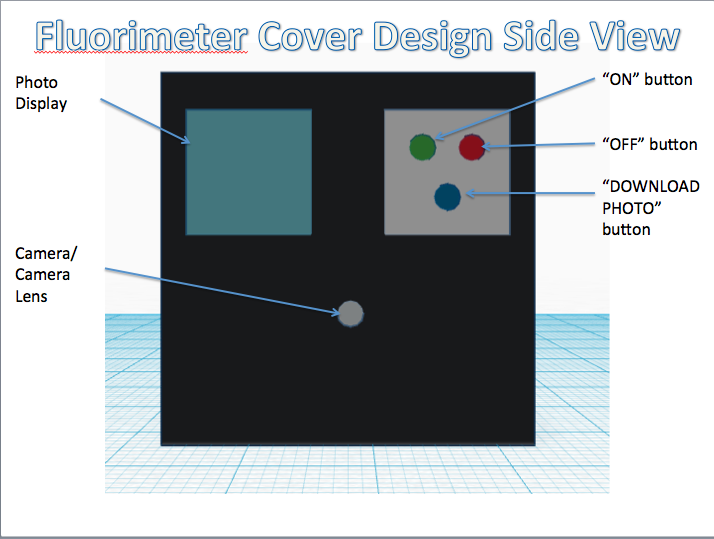
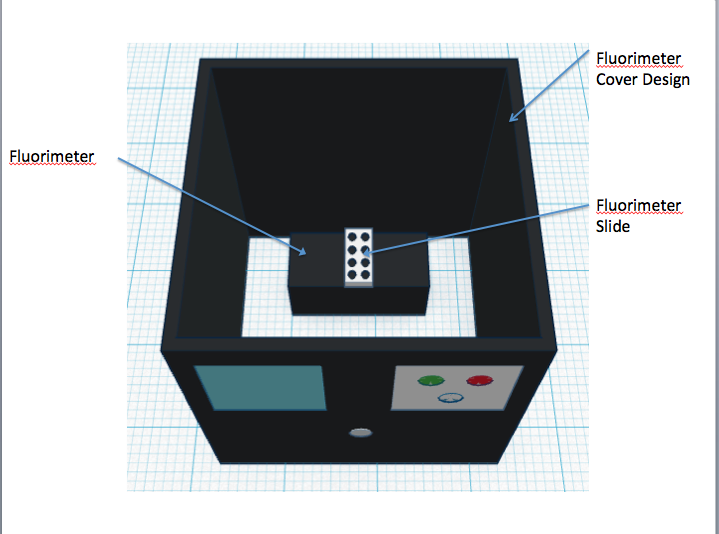
The newly designed OpenPCR machine implements a hyper-speed fan that will cool down the reaction quicker than the regular fan that contributes to the lengthy 2 hour PCR reaction duration time. In addition to the increased efficiency, there is a built-in camera into the side of the fluorimeter, which can be controlled by a smart phone to adjust zoom options and camera resolution.
Feature 1: Consumables KitThe consumables kit will include 16 dark plastic test vials, enough liquid reagents for 16 tests, a micropipette, and 2 hydrophilic glass slides
Feature 2: Hardware - PCR Machine & FluorimeterOne aspect of the PCR machine that Group 30 has redesigned was the fan system that is used to cool down the PCR reaction. In order to speed up the reaction in general, the group has redesigned the PCR machine to implement a hyper-speed fan that will cool down the reaction quicker than the regular fan that contributes to the lengthy 2 hour PCR reaction duration time. A weakness of the Fluorimeter system that was improved was the method for taking pictures. With the regular fluorimeter system, a phone camera had to be placed into a cradle, and it had to be adjusted or moved several times throughout the experiment to take the best quality of pictures possible. Thus, with the redesigned system, Group 30 has implemented a built-in camera into the side of the fluorimeter, which can be controlled by a smart phone app that allows the group to see inside the fluorimeter without moving the fluorimeter door. The features of the app also allow the experimenter to control zoom and camera resolution.
| |||||||