BME100 f2014:Group24 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
Our Team
ProtocolList Of Materials
PCR Reaction Sample List
OpenPCR Program
Denature at 95°C for 30 seconds, Anneal at 57°C for 30 seconds, and Extend at 72°C for 30 seconds
DNA Sample Set Up Procedure 1. Gather materials 2. Create two strips of four linked tubes by cutting the strip of empty PCR tubes in half. This needs to be done so that all the tubes will fit in the OpenPCR machine. 3. Label the sides of the empty tubes with the Tube Labels your group created. Do not label the lids of the tubes 4. Carefully place the tubes in the rack. 5. Start with the empty tube you labeled as the positive control. Using proper pipetting technique, transfer 50 μL of PCR Reaction Mix into this empty tube. Discard the disposable tip into the collection cup. Do not re-use tips and cross-contaminate your samples! 6. Using a fresh pipette tip, transfer the positive control DNA/primer mix into the same tube. The total volume in your positive control PCR reaction tube is now 100 μL. 7. Repeat steps 5 and 6 for the negative control. Use the appropriate DNA/primer mix for the corresponding tubes. When you are done, all of your labeled tubes should contain DNA/primer mix, resulting in a 100 μL complete PCR reaction in each tube. 8. Close the lids tightly on the PCR reaction tubes. 9. Take the tubes over to your assigned PCR machine. Place the tubes into the slots in the heating block. Do not start the machine until all 16 slots are filled (multiple groups need to run the reactions simultaneously). Research and DevelopmentWhat is the function of each component of a PCR reaction? Template DNA: The DNA that is being used as the base/template to build the new strand of DNA. Primers: Laboratory designed pieces of DNA that are made to have the desired sequence of nucleotides. The primers are made to match the segment of DNA being copied. Taq Polymerase: Responsible for copying a cell's DNA before it divides. It can attach itself near the end of a primer and add nucleotides from the surrounding liquid. Deoxyrybonucleotides:The four building blocks of the DNA molecule: Adenine, Thymine, Cytosine, and Guanine.
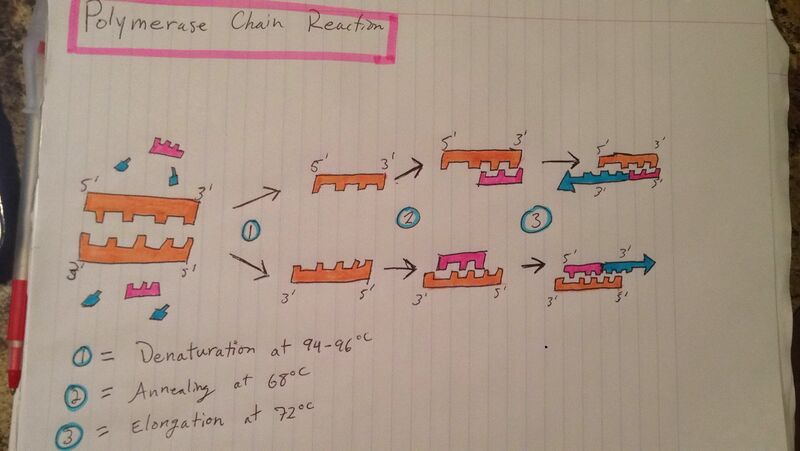
Initial step: 95 C for 3 minutes: The Template DNA begins to unwind its double helix shape. Denature at 95 C for 30 seconds: The Template DNA strand separates its two halves. Anneal at 57 C for 30 seconds: The primers anneal to the target sequence ends. Extend at 72 C for 30 seconds: The Taq Polymerase binds to the Primer sequence Final Step: 72 C for 3 minutes: The Taq Polymerase adds the deoxyrybonucleotides to the Template DNA Final Hold: 4 C: The target DNA sequence was made and copied multiple times with all the components and is now being held stable in the machine. Base Paring: Adenine pairs with Thymine Cytosine pairs with Guanine
Base-pairing occurs in the annealing and extension steps. Annealing is when in the template DNA, the primers attach to the two DNA strand. Extension is when the nucleotides are brought over by the taq polymerase, and thus extending the primers and pairing them in order of the DNA sequence.
What is a nucleotide? It is the basic building block of dna What is a polymorphism? a common variation of DNA among individuals. What species is this variation found in? Homo Sapiens What chromosome is this variation found on? 21:34370656 What is listed as the Clinical Significance of this SNP? pathogenic What gene is this SNP associated with? KCNW2 What diesease is linked to this SNP? Congentital Long QT syndrome What does KCNE2 stand for? potassium voltage-gated channel Describe the molecular function of this gene: This gene encodes a member of the potassium channel. This member is a small integral membrane subunit that assembles with the KCNH2 gene product, a pore-forming protein, to alter its function. This gene is expressed in heart and muscle. What is an allele? An allele is an alternative form of a gene that is located at a specific position on a specific chromosome. The disease-associated allele contains what sequence? CTC The numerical position of the SNP is: 34370656 Non-disease forward primer (20nt): CAT GGT GAT GAT TGG AAT GT The numerical position exactly 200 bases to the right of the disease SNP is 34370856 Non disease reverse primer: CCC TTA TCA GGG GGA CAT TTT Disease forward primer: CAT GGT GAT GAT TGG AAT GC Disease reverse primer (20nt): CCC TTA TCA GGG GGA CAT TT Bonus PCR Drawing |
||||||||||||||||||||||||||||||||||