BME100 f2014:Group22 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis SystemIn order to test various patients for the presence of the disease-associated SNP, 34 teams of roughly 6 members each ran tests using a PCR reaction. Each group tested two individual patients, and prepared three replicated solutions of DNA and primer mix for each to be compared to positive and negative controls. The three replicant solutions from each pair of patients were then run through 30 cycles in the Open PCR machines along with the positive and negative controls, giving a total of 8 PCR results per group. In order to compare each of the results to one another and determine their concentrations of DNA, a calibration curve was created using the fluorimeter and solutions of DNA of known concentrations. For calibration, 6 drops were prepared using 80 μL of DNA solution (0, 0.25, 0.5, 1, 2, and 5 μg/mL concentrations) and 80 μL of SYBR GREEN I dye were placed on a hydrophobic slide and placed in the fluorimeter. Using the same camera settings and distance from the camera to the fluorimeter (within each group; setups and cameras varied from group to group), three photos were taken of each droplet in the fluorimeter to prevent error and increase the reliability of the data. Each of the 8 test solutions from the PCR machine were put in the fluorimeter in a similar fashion. Then in ImageJ, the photos of the both calibration solutions and the solutions from the PCR machine were analyzed, comparing the intensity of green wavelengths of light between the droplet and the background interference. This resulted in a quantitative measure of the intensity of fluorescence from the SYBR GREEN I dye that could be compared to the calibration solutions to find the approximate concentration of DNA in the solution. As a result of the entire class's data (34 groups, 68 total patients), 30 patients' tests returned a positive conclusion, while 24 returned a negative conclusion. Out of the remaining 14 patients' samples, 8 were inconclusive and 6 were failed tests with no conclusion at all. As far as diagnosing patients, the PCR tests across all 34 groups correctly diagnosed 10 patients out of 23 that had the disease, and 12 patients out of 45 that did not have the disease. What Bayes Statistics Imply about This Diagnostic ApproachCalculations 1 & 2: Conclusiveness and Errors of PCR Tests For calculations 1 and 2, the bayesian statistics had to do with the relationship between the likelihood of a PRC test (including replicates of the same patient) being positive and the likelihood of the overall conclusion (taking into account all three replicates of each patient) being positive. Overall, the likelihood of a positive PCR test resulting in a positive conclusion (sensitivity) and the likelihood of a negative PCR test resulting in a negative conclusion (specificity) were both fairly close to 1.0, although the system had a slightly lower specificity than its sensitivity. This means that the actual PCR test was fairly effective at producing positive conclusions, but it was slightly less conclusive when it came to negative ones. This discrepancy may be due to the fact that the conclusion was fairly arbitrary, requiring intuitive human judgement (across 34 different groups) to judge whether the concentration of DNA was more similar to the positive control than the negative one. Another source of error comes from the disparity between camera setups between groups, as neither the brand of phone nor the specific camera settings nor the distance between the camera and the fluorimeter were held constant between groups. Calculations 3 & 4: Accuracy and Errors of RNP Results in Detecting Disease Calculations 3 and 4 dealt with the reliability of the conclusion of the PCR test in correctly diagnosing a patient as either having the relevant mutated gene or being genetically healthy. The class as a whole correctly identified only a small portion of the patients that were positive for the disease as having the disease, as the bayes value was very close to 0. Therefore, the test was largely ineffective in providing true positive tests (low sensitivity). The lab class did only marginally better at correctly identifying patients that were negative for the disease, correctly identifying not quite a third of the patients with true negative tests (fairly low specificity). Possible Sources of Error - The inaccuracies of the PCR tests and diagnoses may be due in part to the fact that the conclusion from each set of PCR tests was fairly arbitrary, requiring intuitive human judgement (across 34 different groups) to judge whether the concentration of DNA was more similar to the positive control than the negative one. Another source of error comes from the disparity between camera setups between groups, as neither the brand of phone nor the specific camera settings nor the distance between the camera and the fluorimeter were held constant between groups. Additionally, some error stems from the 8 inconclusive tests and 6 failed ones which, having no conclusion, subtracted from the overall accuracy and specificity of the conclusion. These may have been a result of difficulties that were presented in the setup of the fluorimeter itself, which did not lend itself to stability nor consistency in the way it moved when inserting or removing slides. Computer-Aided DesignTinkerCAD
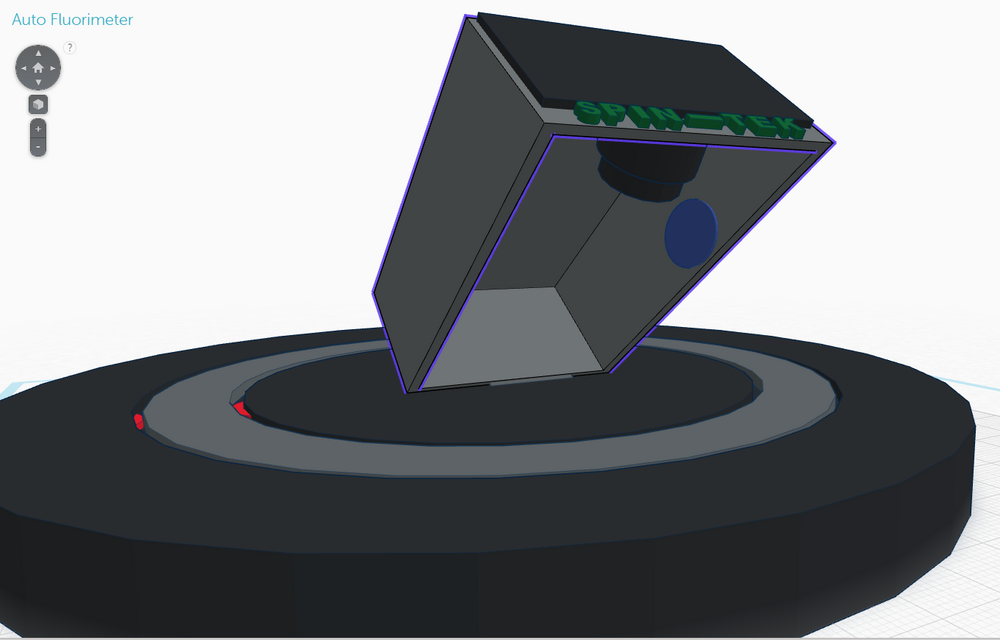
Feature 1: Consumables KitThe consumables kit would include: plastic sample tubes, micropipette tips, glass viewing slides, PCR plate. To improve the process of PCR, since most consumables cannot be reused immediately in an experiment until they are sanitized and autoclaved, there is little efficient improvements that can be made to the consumables themselves. However, an improvement to make PCR more efficient would be to modify the micropipette and redesign it to hold a stack of pipette tips on the inner barrel of the micropipette. To use this device, every time instead of attaching a pipette tip from a box of them, the operator would push a button located on the pipette tip that would push out one of the pre loaded tips from the inside of the device and it would then be used like a normal micropipette and once done the tip would be discarded into the sharps container. This design would still allow for the use of sterile tips, but it would allow the operator to save time in between pipetting different solutions by making the process of getting a new sterile tip to use much more efficient. Feature 2: Hardware - PCR Machine & FluorimeterIn our group's redesigned system, the design and function of the OpenPCR machine will be left unchanged, but the fluorimeter will be modified in order to address the issues of camera settings, distance, and movement (as shown in the 3D model above). In the A redesign to the fluorimeter set-up, as shown in the 3D model above, would include a single, circular slide with a hydrophobic surface (shown by the light grey ring in the diagram), which could be loaded with the fluorescent dye and DNA solution just once and rotated automatically using rotating discs (shown in red) to switch between droplets without having to remove and replace the slide, thus removing some of the risk of jarring the system. Additionally, pre-loading would allow fluorimeter readings to be done quickly and efficiently. Another added feature to the fluorimeter is the addition of a built-in dark box (shown in light grey) that is able to be lifted and replaced easily between samples, further reducing the chances of jarring the fluorimeter and camera set-up when removing or replacing the independent dark box that was included in the original design. Included in the dark box is a built in fixed camera and external LCD display (shown in black). The fixed camera serves to provide a major upgrade to the stability and consistency of the camera, as the image will be taken from the same distance each time, without having to adjust the view or settings. The external LCD display serves to provide near-instantaneous feedback, allowing for mistakes (blurred photos, uneven droplets, etc.) to be caught and corrected easily, and allows for a built-in program to analyze the images immediately and transferred to a USB. Additionally, the fluorimeter light in the dark box (shown in blue) assures that the the light is always the same distance from the camera and from the droplet, further improving consistency and precision.
| ||||||