BME100 f2014:Group17 L4
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||
|
OUR TEAM
LAB 4 WRITE-UPProtocolMaterials
Values to Input in the Program
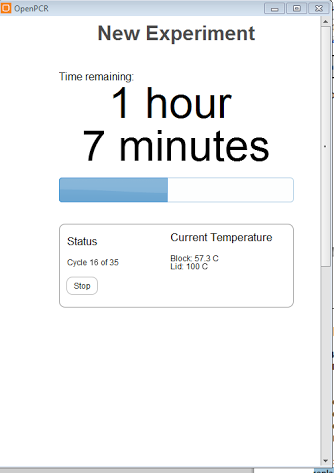
Screenshot of what is displayed on computer monitor during the PCR cycles, the OpenPCR program displayes cycles completed, temperature as well as time remaining for the total number of cycles.
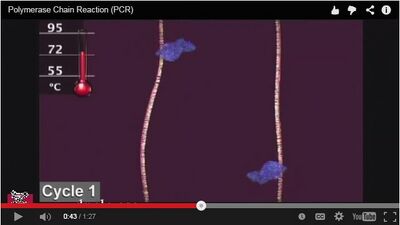
Research and DevelopmentPCR - The Underlying Technology PCR technology amplifies a single copy of DNA into thousands or billions of copies. Open PCR does this through repeated cycles of heating and cooling which separates the double helix into single strands. By initially heating the DNA the hydrogen bonds are broken, allowing for two separate single strands to be formed. Generally, PCR is used to replicate a portion of DNA (DNA template), rather than an entire genome. This technology is important in the medical field as it can help to identify disease or learn how to treat diseases; more specifically it is used in human diagnostics, environmental monitoring, and scientific research. Template DNA, primers, taq polymerase, and deoxyribonucleotides (base pairs) are the basic components necessary to complete an OpenPCR cycle. The primer is an important part of the polymerase chain reaction. Primers are short portions of the DNA sequence, they bind to certain areas of the single strands of DNA during the PCR process. Taq polymerase binds to the primers at the parts to be copied during higher cycle temperatures; it then adds nucleotides to the single strands. The image above shows the Taq polymerase in blue attached to the primers on the two single strands (image screenshot from https://www.youtube.com/watch?v=2KoLnIwoZKU). The neucleotides complete the second strand of DNA, as well as the first cycle in the OpenPCR. The process is repeated for the programmed number of cycles. It is important to note that the four bases are paired together, each base needs to have its corresponding base on the second strand of DNA (GC and AT). PCR can replicate DNA faster than cancer cells are capable of completing the same process (https://askabiologist.asu.edu/pcr-polymerase-chain-reaction). If the sequence of DNA selected to be replicated has a mutation (cancer) the replicated DNA would also be mutated.
| ||||||||||||||||||||||||||||||||||