BME100 f2014:Group15 L6
| Home People Lab Write-Up 1 | Lab Write-Up 2 | Lab Write-Up 3 Lab Write-Up 4 | Lab Write-Up 5 | Lab Write-Up 6 Course Logistics For Instructors Photos Wiki Editing Help | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
OUR COMPANY
LAB 6 WRITE-UPBayesian StatisticsOverview of the Original Diagnosis System In this investigation, the BME 100 class utilized PCR analysis to determine the existence of the disease-associated SNP in multiple people. In order to run the tests and acquire accurate data, 34 groups of six students ran 186 PCR tests and diagnosed 68 patients with the disease-associated SNP. Each individual group ran their own PCR tests on the DNA of two different patients to determine if they were negative or positive for the SNP. The results were compiled into one large document, and Bayesian calculations to analyze the data and determine our probability with our results. In order to prevent error within our testing, Dr. Haynes used her significantly more precise PCR machine and analyzed the three replicates of DNA per patient for both patients for the BME 100 class. When analyzing the fluorescence of SYBR Green fluorescent, we ensured that the fluorimeter and camera system (for later analysis in ImageJ) were isolated from surrounding light via the dark box. We also analyzed two control samples of DNA, one that was negative for the SNP, and one that was positive for the SNP, in order to ensure our SYBR green would detect the SNP in our experimental samples. Positive results would glow green when exposed to UV light, negative results would remain clear. For the ImageJ image analysis program, which was used to analyze the concentration of SYBR green fluorescence in the experimental samples, the split channel tool was applied, and only green colored images were analyzed. The other displayed colors of blue and red of the RGB color scheme were discarded in the analysis. Each calibration trial used three images for analysis, and the mean average taken and used for our data.
What Bayes Statistics Imply about This Diagnostic Approach
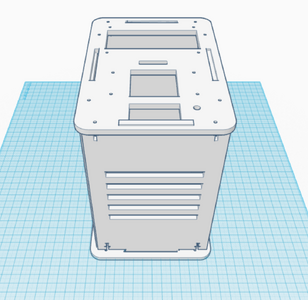
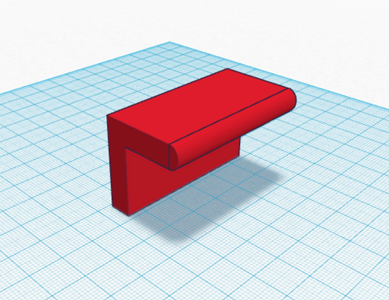
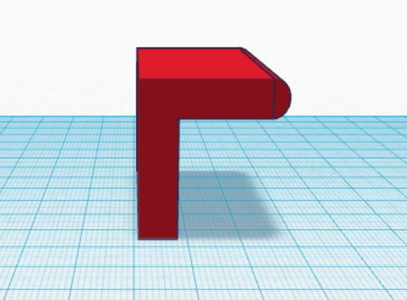
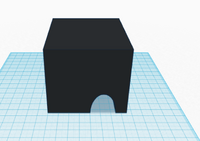
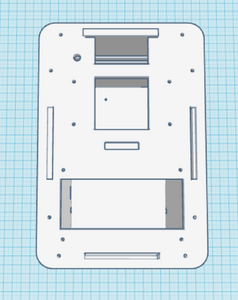
The third and fourth Bayesian calculations were used to determine a “yes” or “no” diagnosis of disease-associated SNP from the positive and negative conclusions calculated in the first two calculations. A “yes” reading shows that the patient does have the cancerous disease, and a “no” reading shows that the patient does not have the disease. In the table of calculations, these two values are displayed very small compared to the 100% standard. This means that if a sample of DNA is concluded positive for the SNP by the PCR, there is a small chance that the patient actual has the disease, and vice versa for a negative conclusion. This error once again adds to the lack of reliability in PCR testing. Computer-Aided DesignTinkerCAD Our Design
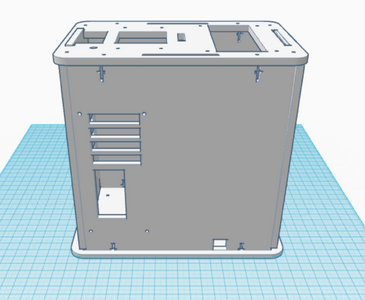
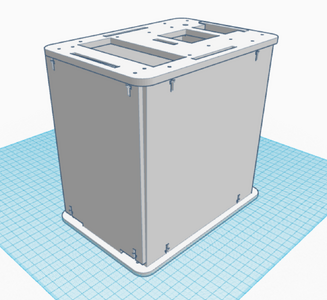
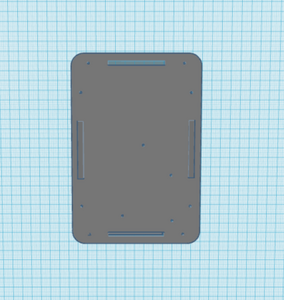
The new design for the Open PCR machine includes the addition of clamps on the outside of the machine and an improved tray to hold the test tubes. The clamps would be added to the top of the PCR in order to more effectively hold down the lid and prevent heat loss. The tray that holds the test tubes during the PCR process would be made of a metal that is a better conductor of heat so that it can warm up and cool off much more efficiently. The area that the tray is sitting in would have to be a good insulator to keep the temperature constant which is why the physical tray needs to be an effective conductor. These changes are necessary because the DNA needs optimal conditions to satisfy the temperature requirement of the DNA Polymerase for it to make as many copies as possible.
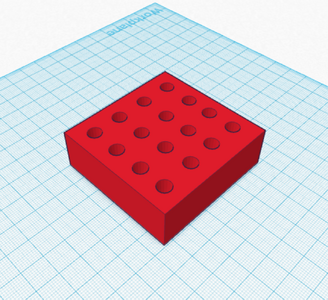
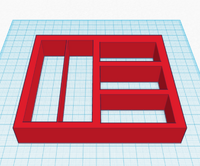
Feature 2: Consumables Kit
Feature 3: Hardware - PCR Machine & FluorimeterThe new system will be very similar to the devices now. The only difference for the PCR machine will be the addition and revitalization of some parts, including the clamps on the exterior portion of the lid and the newly designed tray. For the fluorimeter, the outer flap that is suppose to be closed before the picture is taken will have a small area cut out so that a person's arm can fit underneath, but the lid will still close completely. This should reduce any error that occurs because of an additional light source. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||