BISC 219: Mod 2 Lab 8
PCR Reaction Cleanup and Sequencing
In order for our sequencing reactions to work properly we need to remove the components of our PCR reaction (like dNTPs, Taq and salts) that might prevent the sequencing from working properly.
We will be using the Qiagen QIAquick PCR Purification Kit (www.qiagen.com catalog #28104). We will follow the instructions as they are in the Qiagen protocol book.
- Combine all of your (40 μL) of remaining PCR reaction with 400 μL of Buffer PB in a new microfuge tube. Mix.
- Add all of the 440 μL of PB+PCR to a Qiaquick spin column
- To bind the DNA to the membrane: spin at top speed in the centrifuge for 60 seconds
- Discard flowthrough and put the column back in the collection tube
- Wash the sample by adding 750 μL of Buffer PE to the Qiaquick column and centrifuge for 60 seconds
- Discard flowthrough and put the column back in the collection tube
- Centrifuge the column for 1 minute at top speed to completely remove all of the Buffer PE
- Discard the collection tube
- Place the Qiaquick column in a clean 1.5 ml microfuge tube
- Add 30 μL of Buffer EB (10 mM Tris-Cl, pH 8.5) to the membrane
- Centrifuge for 1 minute at top speed
- Check to see a small amount of liquid at the bottom of the microfuge tube
- Discard column
- Label your collected DNA with your color, initials, day and name of DNA.
Sequencing:
BACKGROUND: Sequencing a portion of our genes of interest will provide us with direct evidence of the nature of the mutation (i.e: point mutation or deletion) in the mutant worms, as well as if RNAi has any affect on DNA sequence. It will help us make conclusions about where in the DNA to RNA to protein pathway RNAi actually works. This knowledge will be useful to establish a link with the type of functional defect present in the protein produced by the mutated gene.
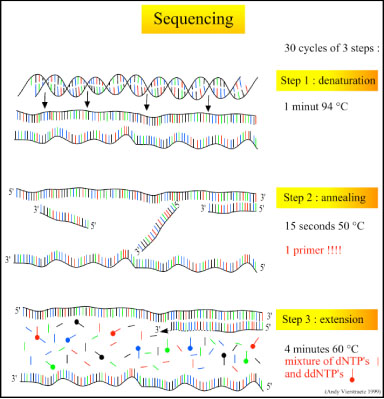
There are three major steps in a sequencing reaction, which are repeated for 25 cycles in a thermocycler:
- Denaturation at 94°C: During the denaturation, the double strand melts open to single stranded DNA and all enzymatic reactions stop, including the extension from the previous cycle.
- Annealing at 50°C: In each sequencing reaction, a specific primer is required. This primer is a short sequence of bases complementary to a region of the plasmid upstream of the target gene. The primer will anneal and stay, we hope, only at one position on the single stranded DNA template. In sequencing reactions there is only one strand copied whereas, in PCR, two primers are used so both strands of the DNA template are copied. The primer jiggles around, caused by the Brownian motion. Ionic bonds are constantly formed and broken between the single stranded primer and the single stranded template. The more stable bonds last a little bit longer (primers that fit exactly) and on that small section of double stranded DNA (template and primer), the polymerase can attach and starts copying the template. Once there are a few bases built in, the ionic bond is so strong between the template and the primer, that it does not break anymore.
- Extension at 60°C: Sixty degrees is the ideal working temperature for polymerase activity in sequencing even though, normally, it is 72 °C. Because the growing strand must incorporate ddNTP's which are chemically modified with a fluorescent label, the temperature is lowered so it has time to incorporate the 'strange' molecules. Well designed primers, where there are a few bases built in, already have a stronger ionic attraction to the template than the forces breaking these attractions. Primers that are on positions with no exact match, come loose again and don't give an extension of the fragment. The bases (complementary to the template) are coupled to the primer on the 3'side (adding dNTP's or ddNTP's from 5' to 3', reading from the template from 3' to 5' side, bases are added complementary to the template). When a ddNTP is incorporated, the extension reaction stops because a ddNTP contains a H-atom on the 3rd carbon atom (dNTP's contain a OH-atom on that position). Since the ddNTP's are fluorescently labeled, it is possible to detect the color of the last base of this fragment on an automated sequencer.
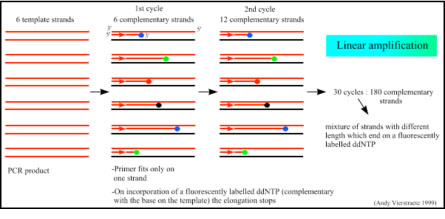
Because only one primer is used, only one strand is copied during sequencing, there is a linear (not log as in PCR) increase of the number of copies of one strand of the gene. Therefore, there has to be a large number of copies of the gene in the starting mixture for sequencing. If there are 1000 copies of the wanted gene before the cycling starts, after one cycle there will be 2000 copies: the 1000 original templates and 1000 complementary strands with each one fluorescent label on the last base. After two cycles, there will be 2000 complementary strands, three cycles will result in 3000 complementary strands and so on.
Procedure/Protocol
You will set up the sequencing reactions for your PCR reaction containing the amplified DNA from your RNAi worms. The region of interest in the amplified DNA will be sequenced with primers constructed from the flanking regions. It is best to use a sense (forward) and an antisence (reverse) primer for sequencing. We will be doing both reactions today.
| Gene name | Primer | Code | DNA μL | HPLC H2O | 10 mM primer μL | Big Dye μL |
|---|---|---|---|---|---|---|
| Your gene name - forward | Forward | Lab day/color - tube 1 | ||||
| Your gene name - reverse | Reverse | Lab day/color - tube 2 |
- Label two 0.2 ml tubes in your team color with your sequencing code (see table above). Treat these thin walled tubes with care (no squeezing or rough handling) as they may crack and then leak when you spin them later.
- Once the aliquot of BigDye™ has thawed place 1 μl on the bottom of each tube, you don’t need to change tips since the tubes are empty (PLEASE MAKE SURE YOU USE THIS REAGENT FIRST- Your instructor will pipet this precious reagent into your tubes for you).
- Add the gene specific primers, in the volume according to the table above. Remember, only one primer per reaction. It is advisable to add the primer to the wall of the tube. Change tips!
- Add 4 μl of cleaned worm pcr product to each tube and 3 μl of sterile HPLC water. Your total volume should be 9.6 μl in each tube. Consult with your instructor if you have any question about the appropriate volumes for your reactions. Give your remaining purified DNA back to your instructor for freezing.
- Shake down or tap the tubes very gently to make sure all reagents are mixed and are in the bottom of the tubes. To reiterate, do not treat these tubes roughly as they are thin walled and will break easily (and leak)! Because BigDye is light sensitive, keep the tubes on ice and covered with aluminum foil until everybody has finished their reactions and we can load them all in the thermal cycler (PCR machine).
The reactions will run for 2.5 hours. We will clean up the sequencing reactions as soon as they are ready using the Qiagen DyeEx 2.0 Spin Kit (catalog #63204). We will follow the manufacturer's instructions for this procedure.
- Gently vortex the spin column to resuspend the resin
- Loosen the cap of the column 1/4 turn (to avoid a vaccuum inside the tube)
- Snap off the bottom closure of the spin column and place the column in a 2 ml collection tube
- Centrifuge for 3 minutes at 750 x g
- Transfer the column to a clean centrifuge tube
- Slowly apply the 10 ul sequencing reaction to the CENTER of the top of the gel bed
- Centrifuge for 3 minutes at 750 x g
- Remove the spin column from the microfuge tube
- The small amount of liquid in the microfuge tube is your "cleaned up" sequencing reaction!
- Add 10 μL of HPLC water to each sample for a total of about 20 μL in each reaction
Once your samples are cleaned up we will load them into the sequencer to run overnight. Your instructor will post the results tomorrow.
Wild type (N2) and mutant samples have been sequenced previously and will be provided for your analysis.
While your reactions are running - you will proceed with part 1 of your transgenic plant analysis.
RNAi Schedule of Experiments
RNAi General Information
Media Recipes
Lab 4: Picking your gene to RNAi
Lab 5: Plasmid DNA isolation and transformation
Lab 6: Induction of RNAi plasmid and C. elegans feeding
Lab 7: Single Worm PCR and Agarose Gel Electrophoresis