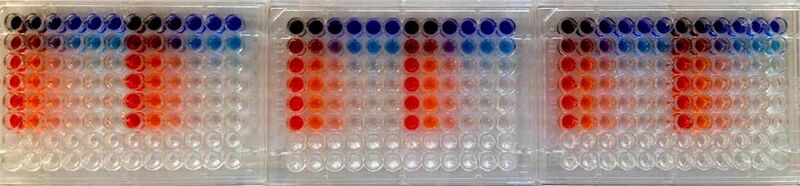
20.109(S15):Growth of phage nanowires (Day 1)
Growth of phage materials
Today's Goals:
Isolate bacteriophage from bacterial culture, measure phage concentration, and complex phage with gold nanoparticles at the assigned ratios.
Introduction


The accomplishments of the natural world can inspire us to great engineering feats. Biomineralization is one particularly impressive trick nature pulls off. Vertebrates, invertebrates and plants all have ways to precisely position inorganic substrates into crystalline order. For example, calcium carbonate will form unstructured dust in the absence of genetically-programmed organizers, but the same material can be made into the hard and luminous shells of sea creatures. Similarly, diatoms organize silicon dioxide into intricate patterns that manufacturers of electronic components can’t begin to approach. In one more instance, bacteria align iron inside their cytoplasm to form magnetic rods on the submicron scale. These feats are accomplished without harsh chemicals, without extreme temperatures, and without noxious wastes that poison the nests of the organisms themselves. Humans have a lot to learn from nature’s successes. In the upcoming weeks we’ll use a virus that infects bacteria, namely the bacteriophage M13, and we'll rely on the self-assembling coat of this virus to template carbon nanotubes or gold nanoparticles and TiO2. The interaction of these materials with a protein on the phage coat yields nanoscale-particles with useful energetic properties, as we’ll see.
About M13
M13 bacteriophage is a member of the filamentous phage family. This well studied phage is roughly 880 nm long, has a diameter of about 6.5 nm, and a mass of approximately 16.3 MDa. Of this mass, 87% is accounted for by the structural coat proteins which encase the single stranded DNA genome of about 6.4 kb. The M13 genome encodes 11 proteins, five of which compose the phage’s protein coat and six of which are involved in phage maturation inside its E. coli host. The phage coat is primarily assembled from a 50 amino acid protein called pVIII (or p8), which is sensibly enough encoded by gene VIII (or g8) in the phage genome. For a wild type M13, there are approximately 2,700 copies of p8 which pack together to build the majority of the ~880 nm long coat. The coat's dimensions are flexible though and the number of p8 copies adjusts to accommodate the size of the single stranded genome it packages. In contrast, there are typically only about five copies of each of the other four coat proteins, located at opposite ends of the mature virion. Of note, pIII (or p3) is a 406 amino acid protein found at one end of the phage, and is responsible for recognizing the host E. coli. From a biological engineering perspective, this bacteriophage provides an efficient and modular platform allowing for either random or specific genetic alterations of the phage DNA to be related to phenotypic changes of the phage surface. For example, genetically encoded fusions to the surface coat proteins can be used to display peptides with the ability to bind materials with which the wild type phage would normally not interact.
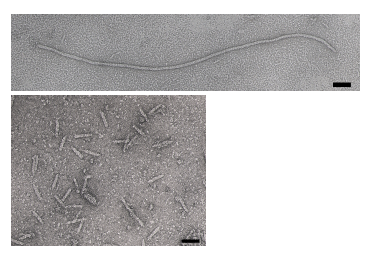
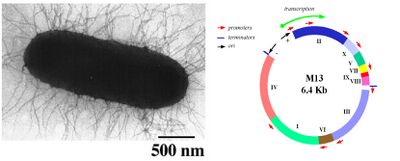
Phage life-cycle
The general stages to a viral life cycle are: infection, replication of the viral genome, assembly of new viral particles and then release of the progeny particles from the host. Filamentous phages use a protein at their tip, namely p3, to contact a bacterial structure known as the F pilus to infect E. coli. Through mechanisms which are not fully elucidated, the phage genome is then translocated into the cytoplasm of the bacteria where resident proteins convert the single stranded DNA genome to a double stranded replicative form (RF). This DNA then serves as a template for expression of the phage genes and produces new phage particles that shed off the surface of the infected cell. While other phage are known to lyse their host cells, M13 and E. coli co-exist in a balanced way, allowing the growth of both host and virus, although the infection does slow down the doubling time of the E. coli, causing plaques to form in a bacterial lawn.
Phage display
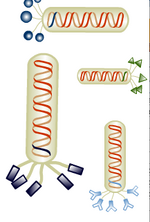
Despite phage display techniques being available for more than a generation, this tool has been applied only recently to the search for novel materials. Largely it’s been Angela Belcher and her lab who highlighted and then demonstrated the usefulness of this search tool for finding peptides that interact with materials to meet human needs. That M13 could interact with inorganic materials could not have been predicted from the original genetic studies on the phage, but there was also no one who had tried it. Phage that can bind to cobalt oxide, gold, iridium and indium tin oxide are all in-hand thanks to their work (e.g see reference [3]). Today you will harvest phage that can bind to gold nanoparticles and TiO2 since these can be used to build nanocomposites that will be assembled into a photovoltaic device before this module is over.
Protocols
Pre-lab work:
In preparation for this lab, a bacterial host E. coli K12 ER2738 (New England Biolabs) was infected with the M13 phage clone named p8#9 that can bind thin gold films (PMID: 16178252). This infection allowed for amplification of phage as the bacterial culture was incubated over night. These particular phage clones are modified with an 8 amino acid addition to the surface-exposed terminus of the p8 protein -- VSGSSPDS. Phage were obtained by panning, starting with a library of p8 mutants and isolating phage that could bind materials.
Behind the scenes Cherry and members of the Belcher Laboratory have also prepared another phage clone, DSPH, that can bind single-walled carbon nanotubes (SWNTs) (doi:10.1038/nnano.2011.50) by incorporating the DSPHTELP peptide into p8. They will prepare solar cells utilizing the DSPH phage and SWNTs that you can use to as a comparison for your gold nanoparticle-based solar cells.
Today, you will isolate phage from the infected bacterial culture and measure its concentration using the spectrophotometer. Finally, you will bind the phage to 5, 20, or 50 nm gold nanoparticles. These phage will be used as a template for TiO2 nanowire synthesis incorporated in the dye-sensitized solar cell (DSSC) in future lab sessions.
Part 1: Phage purification
- Divide the overnight culture (~80 ml volume) into 2 x 50 ml conical tubes.
- Label the tubes with your group color and phage type.
- Spin 3,250 rpm, 10 minutes using a fixed angle rotor. The TA will help you load your samples into the centrifuge located on the TA bench.
- Transfer the supernatant to new 50 ml conical tubes, splitting the supernatant between them. The transfer should be done with a plastic pipet and a bulb so you can measure the volume of supernatant.
- Add a 1/6th that volume of 20% PEG-8000/2.5M NaCl solution.
- Invert to mix, then incubate on ice 60 minutes. During this time, you can finish up your reflections, or start thinking about the culminating 20.109 assignment described here.
- Spin at 3,250 rpm for 15 minutes. A white pellet may be visible: these are your precipitated phage. If you can't see a pellet keep going, but be aware of where the pellet you can't see is in the tube and don't scrape a tip against it or you will accidentally remove it. Hint: Mark the tube where you expect the pellet to form when as you load it into the centrifuge.
- Remove the supernatant by pouring most down the sink and the rest with aspiration (carefully so as not to disturb the pellet).
- Resuspend the pellet in 3 ml sterile H2O. This is best done by adding 3 ml of H2O to one of the conical tubes, washing the water up and down the side of the tube with the phage pellet, and then moving the 3 ml of phage solution to the second tube and dissolving that pellet as well by washing the water up and down the side of the tube.
- Split the phage solution between three eppendorf tubes. For this part of the protocol, you will be given special eppendorf tubes that can hold 2 ml.
- Spin tubes in a room temperature microfuge for 1 minute to remove residual bacterial residues. Transfer supernatant to fresh eppendorf tubes.
- Add a 1/6th volume of 20% PEG-8000/2.5M NaCl solution.
- Invert to mix, then incubate on ice for 15 minutes.
- Spin the tubes full speed in a microfuge for 10 minutes.
- Aspirate the supernatant and resuspend the pellets (if you can see them) in 0.2 ml TBS--using 0.2 ml to resuspend one pellet and moving that volume to resuspend the next pellet, and then moving that volume again to resuspend the third pellet. This is your phage stock (yay!).
- If the solution looks at all cloudy, spin in a room temperature microfuge for 1 minute more and move supernatant with the phage to a new tube.
Part 2: Measuring concentration of phage
where
- the molar extinction coefficient of the phage and the average size of a DNA base are used collected into the constant
- the absorbance at 269 nm reflects the protein and DNA content in the solution
- the absorbance at 320 nm corrects for the naturally high baseline value of the solution
- the number of DNA bases in DSPH is ~7220.
This method for titering the phage stock is less informative than the traditional plaque method (known as titering) since materials other than phage might be contributing to the absorbance readings. Thus, the number of infectious particles isn't truly known. Since infectivity is not critical for the synthesis of SWNT-TiO2 nanowires,however, we will be using spectrophotometry only.
- Dilute the phage stock you have 1:10 by adding 70 ul of the phage to 630 ul of TBS, vortex to mix and then move this solution to a quartz (not plastic!) cuvette.
- A few things to be aware of when using quartz cuvettes:
- They are very expensive.
- The lab has very few.
- When you are done using your cuvette, you should carefully clean it by shaking out the contents into the sink and rinsing it once with 70% EtOH, then two times with water. Quartz cuvettes get most of their chips and cracks when someone is shaking out the contents since it is so easy for the cuvette to slip from wet fingers or be hit against the sink. Don’t let this happen to you.
- A few things to be aware of when using quartz cuvettes:
- Read the absorbances of your phage dilution at 269 and 320, using TBS in a second quartz cuvette to blank the spectrophotometer at each wavelength.
- Calculate the number of phage particles/ml using the formula shown above.
Part 3: Binding phage to solar cell enhancers: Complexing phage with gold nanoparticles
The class will be optimizing the solar cell's performance by comparing the influence of gold nanoparticles versus SWNT’s on efficiency metrics. In order to make the best use of our time, each team will bind their phage to one of the material parameters in the table below. Performance results between all groups and conditions will be compared after solar cell assembly at the end of the module.
| Group (T/R) | AuNP Size (nm) | Group (W/F) | AuNP Size (nm) |
|---|---|---|---|
| Orange | 12 | Red | 5 |
| Yellow | 50 | Orange | 12 |
| Green | 12 | Yellow | 50 |
| Blue | 5 | Green | 5 |
| Purple | 50 | Blue | 12 |
| Pink | 50 | ||
| Purple | 5 | ||
| Platinum | 12 |
Part 3: Complexing phage with Gold nanoparticles
- Each group will complex the gold and phage at the same mass ratio:
1.45x10^-17 grams of gold / phage
- You'll need the following information about your teams specific AuNP size:
5nm particles 6.32x10-5 g/ml
12nm particles
50nm particles 5.68x10^-5 g/ml
- Calculate the volume of AuNP solution that you need using the following equation:
total gold needed in ml = (Total number of phage) * (1.45X10^-17 g gold / phage) * (1 / (concentration of gold in nanoparticles))
- You should use all of the phage that you have left in your reaction.
- Obtain a glass scintillation vial and mix phage and nanoparticles together. Store in the refrigerator and leave it until you return to lab.
Clean up
- The quartz cuvettes can be returned, clean, to the instructors bench at the front of the room.
DONE!
Homework
Due on M3D2
- The primary assignment for this experimental module will be for you to develop a research proposal and present your idea to the class. For next time, please describe five recent findings that could potentially define an interesting research question. You should hand in a 3-5 sentence description of each topic, in your own words, and also formally cite an associated reference from the scientific literature. The topics you pick can be related to any aspect of the class, i.e. DNA, system, or biomaterials engineering. During lab next time, you and your partner will review the topics and narrow your choices, identifying one or perhaps two topics for further research.
- Note: for this assignment, you do not need to have a novel research idea sketched out; you simply have to describe five recent examples of existing work. However, you can start to brainstorm how to build off of those topics into something new if you want to get ahead of the game.
Reagents list
20% PEG 8000, 2.5 M NaCl
TBS
- 50 mM Tris
- 150 mM NaCl
- pH 7.6
10mM NaCl
- 0.5844 g/liter, pH to 5.3 or 10
gold nanoparticles
- Ted Pella 5, 20, and 50 nm gold particles
Navigation Links
Next Day: Biotemplating on phage nanowires Previous Day: Data analysis