20.109(S15):Begin Western protein analysis and choose system conditions (Day2)
Introduction
Ku80 expression analysis via Western Blot
Last time you got familiar with the two cell lines that we will be using during Module 2, and plated a known quantity of each cell type in preparation for protein analysis. Today you will lyse the cells, isolate the protein fraction, and separate the proteins on an acrylamide gel. The teaching faculty will probe the gels for Ku80 expression with a primary antibody, next week you will complete the final steps of this assay and visualize your Western blot.
After completing the Day 1 exercise and hearing the Day 2 lecture, you should be well poised to understand that performing this Western blot serves a true research purpose. The Ku-80 deficient xrs6 cell line can revert to wild-type Ku80 expression under certain circumstances, and thus we want to validate that our current crop of xrs6 are not revertants. Recall that xrs6 have one XRCC5 allele that codes for truncated, non-functional Ku80, but also another, wild-type allele that is normally silenced by methylation. Should this allele become demethylated, the cells will no longer be repair deficient, and we’ll have some really boring flow cytometry results!
A great way to identify a specific protein from a complex mixture is to exploit antibodies – also called immunoglobulins. We will use antibodies to perform a Western blot, a traditional biochemical analysis tool.
Many species can be used to raise antibodies. Most commonly mice, rabbits, and goats are immunized, but other animals like sheep, chickens, rats and even humans can be used. The protein used to raise an antibody is called the antigen and the portion of the antigen that is recognized by an antibody is called the epitope. Some antibodies are monoclonal, or more appropriately “monospecific,” and recognize one epitope, while other antibodies, called polyclonal antibodies, are in fact antibody pools that recognize multiple epitopes. Antibodies can be raised not only to detect specific amino acid sequences, but also post-translational modifications and/or secondary structure. Therefore, antibodies can be used to distinguish between modified (for example, phosphorylated or glycoslyated proteins) and unmodified (or total) protein.
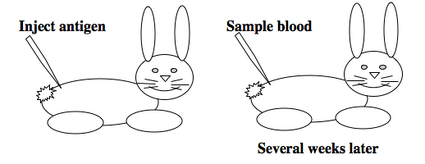
To raise polyclonal antibodies, the antigen of interest is first purified and then injected into an animal. To elicit and enhance the animal’s immunogenic response, the antigen is often injected multiple times over several weeks in the presence of an immune-boosting compound called adjuvant. After some time, usually 4 to 8 weeks, samples of the animal’s blood are collected and the cellular fraction is removed by centrifugation. What is left, called the serum, can then be tested in the lab for the presence of specific antibodies. Even the very best antisera have no more than 10% of their antibodies directed against a particular antigen. The quality of any antiserum is judged by its purity (that it has few other antibodies), its specificity (that it recognizes the antigen and not other spurious proteins) and its concentration (sometimes called its titer). Animals with strong responses to an antigen can be boosted with the antigen and then bled many times, so large volumes of antisera can be produced. However animals have limited life-spans and even the largest volumes of antiserum will eventually run out, requiring a new animal for immunization. The purity, specificity and titer of the new antiserum will likely differ from that of the first batch. High titer antisera against bacterial and viral proteins can be particularly precious since these antibodies are difficult to raise; most animals have seen these immunogens before and therefore don’t mount a major immune response when immunized. Antibodies against toxic proteins are also challenging to produce if they make the animals sick.
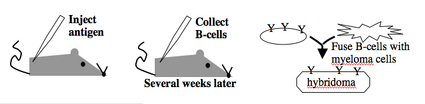
Monoclonal antibodies overcome many limitations of polyclonal pools in that they are specific to a particular epitope and can be produced in unlimited quantities. However, more time is required to establish these antibody-producing cells, called hybridomas, and it is a more expensive endeavor. In this process, normal antibody-producing B cells are fused with immortalized B cells derived from myelomas, and the two cell types are fused by chemical treatment with a limited efficiency. To select only heterogeneously fused cells, the cultures are maintained in medium in which myeloma cells alone cannot survive (often HAT medium). Normal B cells will naturally die out over time with no intervention, so ultimately only the fused cells, called hybridomas, remain. A fused cell with two nuclei can be resolved into a stable cell line after mitosis.
For Western analysis, a high quality antibody can have a relatively low affinity for its target protein. This is because the target is localized and concentrated on a blot, allowing the antibody to bind using both antibody “arms” thereby strengthening the association. Even an antibody that is loosely bound to the blot under these circumstances may dissociate then re-associate quickly since the local concentration of the target protein is high. The lower limit for protein detection is approximately 1 ng/lane, a value that varies with the size of the protein to be detected and the Western blotting apparatus that is used. For most acrylamide gels, the protein capacity for each lane is usually 100 to 200 ug (that would be 20 ul of a 5-10 ug/ul protein preparation). Thus 1 ng represents a protein that is approximately 0.001-0.002% of the total cellular protein (1 ng out of 100,000-200,000 ng). Obviously proteins that make up a more significant fraction of the total protein population will be easier to detect.
Design of the NHEJ reporter plasmid
In addition to the Western protein analysis, you will become intimately familiar with the plasmid reporter assay for measuring NHEJ that we will use in Module 2. You will examine the different elements present in the plasmid, and more importantly you will try to reconstruct some of the design choices that were made when different variants of the reporter were conceived.
A key topic that you will need to understand to complete today’s (re-)design exercise is the function and (sequence-level) structure of restriction enzymes. Restriction endonucleases, also called restriction enzymes, cut (“digest”) DNA at specific sequences of bases. The restriction enzymes are named for the prokaryotic organism from which they were isolated. For example, the restriction endonuclease EcoRI (pronounced “echo-are-one”) was originally isolated from E. coli giving it the “Eco” part of the name. “RI” indicates the particular version on the E. coli strain (RY13) and the fact that it was the first restriction enzyme isolated from this strain.
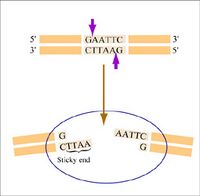
The sequence of DNA that is bound and cleaved by an endonuclease is called the recognition sequence or restriction site. These sequences are usually four or six base pairs long and palindromic, that is, they read the same 5’ to 3’ on the top and bottom strand of DNA. For example, the recognition sequence for EcoRI (see also figure at right) is
5’ GAATTC 3’
3’ CTTAAG 5’

Notice that EcoRI leaves a 5' overhang on either DNA strand; some restriction enzymes leave 3' overhangs instead. Still other restriction enzymes cut precisely in the middle of the palindromic DNA sequence, thus leaving no overhangs after digestion. As you may recall from Module 1, the single-stranded overhangs resulting from DNA digestion by enzymes such as EcoRI are called "sticky" or cohesive ends, while double-stranded ends resulting from digestion by enzymes such as HaeIII are called blunt ends. HaeIII recognizes
5’ GGCC 3’
3’ CCGG 5’
In its simplest description, our reporter assay works as follows: a blue-fluorescent-protein-expressing plasmid is cut by a restriction enzyme(s), then transfected into cells, and repaired at some frequency that we evaluate by measuring the blue fluorescence of said cell population. In fact, the assay design is more subtle than this description encompasses in several respects. The high-level design considerations that we take into account include accuracy of the assay (both technical and biological) and its ease of use. To be confident of the technical accuracy, the assay design includes a second reporter – this time green fluorescent protein – as a transfection control. There is also a control for variable expression of GFP versus BFP, which we will discuss in more detail on Day 4. As another bit of internal validation, one that helps us avoid wasting time with failed products, we have a way to test for successful digestion of the BFP plasmid. Finally, as an example of ease of use, in cases where we cut with multiple restriction enzymes, convenience demands that they are compatible with the same buffer system. We'll dig into some of these considerations further below.
Protocols
Part 1: Prepare cell lysates
- You will each have an ice bucket at your bench with the following pre-chilled items inside: two empty eppendorfs, RIPA buffer, protease inhibitors, and PBS. Begin by labeling the eppendorf tubes as K1 and xrs6 (plus your section).
- Pick up your cell dish from the incubator in TC, and place it at a 30-45 degree angle tilted downward in your bucket.
- Add 10 μL of protease inhibitors to your 1000 μL of aliquotted RIPA buffer.
- Aspirate the media from each well and add about 2 mL (very approximate!) of ice-cold PBS per well by pouring.
- Obtain two pre-chilled scrapers from the fridge.
- Aspirate the ice-cold PBS and repeat the wash once more – make sure to remove ALL of the PBS after this wash.
- Add 100 μL of lysis buffer across the top of each well, allowing it to run down the well.
- Collect the cells to the bottom of the well by scraping each well with a fresh cell scraper.
- First tilt the plate back and forth to coat the cells with lysis buffer. Then go from top to bottom windshield wiper style to pool them down towards the bottom of the tilted dish.
- Add the contents of each well to its respective eppendorf tube.
- At this step it is important to make sure there are no aggregates of cell lysate remaining in the plate. Do this by tilting the plate so that light reflects off the bottom. If you see a 'chunk', pipette your lysate on top of it to dissolve it and collect everything into the eppendorf tube.
- Incubate the eppendorf tubes on ice for 10 min.
- Meanwhile, take two fresh eppendorf tubes and begin to chill them for a later step.
- Spin the tubes at max speed in the cold room centrifuge for 10 min to pellet insoluble material. Bring your eppendorf tubes to the TA who will spin them for you.
- This step is typically referred to as "clearing" the lysate.
- Transfer the supernatant to the new set of eppendorf tubes – be careful not to disturb the pellet at the bottom! Keep these samples on ice when not directly in use.
- The pellet at the bottom contains the DNA from the cell and genomic DNA can get very soupy making it difficult to load your lysate on the SDS-PAGE gel.
Part 2: Measure protein content
You will now measure the total protein concentration in each cell lysate to determine the volume required to evaluate equal protein amounts by Western blot. We are using the Precision Red Advanced Protein Assay from Cytoskeleton.
- Right before it is your turn to use the spectrophotometer, add 10 μL of cell lysate to a plastic cuvette: prepare K1, xrs6, and a blank "lysate" using your leftover RIPA buffer.
- Be careful not to allow 10 μL of lysate to sit in the cuvettes for more than a couple minutes before starting the next step -- it will dry up!
- Bring the cuvettes to the spectrophotometer and add 990 μL of Precision Red reagent. Mix by pipetting up and down a couple times without introducing bubbles.
- After 1 minute, measure each sample at 600 nm, using the RIPA sample as a blank.
- Calculate the two stock protein concentrations using the following information
- 1 absorbance unit = 100 ug protein/mL reagent / cm
- the path length of the spec is precisely 1 cm
- don't forget to account for the dilution factor
- Next, calculate the volumes of lysate and water required to add 10 - 20 μg of total protein to the SDS-PAGE gel in a total volume of 20 μL, per each lysate.
- if your concentration is greater than 1μg/μL, use water to make up the remaining volume
- if your concentration is less than 1μg/μL for at least one sample, scale both samples down to a lower amount, such as 10 μg
- Do not throw away the remainder of your cell lysate! We will store these samples for you at -80 °C in case another Western blot needs to be completed.
Part 3: Separate proteins by SDS-PAGE
Two teams will share one gel.
- Working inside the hood, add 4 μL of 6X Laemlli sample buffer to each of the two lysates (but not to the ladder). Briefly vortex each tube, then quick-spin the solution back down.
- Toss your β-mercaptoethanol-soaked tips into the empty pipette tip box inside the hood so they can air out for a while before being dumped in a burn box. Humans can sense this stinky compound at the order of one part per million!
- Put lid locks on the eppendorf tubes – including the ladder – and boil for 5 minutes in the water bath that is in the fume hood. Repeat the vortex and quick-spin step.
- Meanwhile, get trained in groups of 2-4 people about how to load a protein gel.
- When your samples are ready, you should load them according to the scheme below.
- It is okay to let your samples boil 2-3 extra minutes if there is no gel box free, but don't boil them more than 10 min total.
- It is also okay to let them cool a couple of minutes.
- In your notebooks, document the starting and stopping time of electrophoresis, which will be initiated by the teaching faculty at 200 V and run for 30-40 minutes
| Lane | Sample | Volume to load |
|---|---|---|
| 1 | Group 1, K1 | 20 ul |
| 2 | Group 1, xrs6 | 20 μL |
| 3 | "Dual Color" protein molecular weight standards | 2 μL |
| 4 through 6 | BLANK | N/A |
| 7 | "Dual Color" protein molecular weight standards | 2 μL |
| 8 | Group 2, xrs6 | 20 μL |
| 9 | Group 2, K1 | 20 μL |
| 10 | BLANK | N/A |
Part 4: Transfer proteins to membrane
- Wearing gloves, disassemble the electrophoresis chamber.
- Blot the gel to nitrocellulose as follows:
- Place the gray side of the transfer cassette in a tupperware container which is half full of transfer buffer. The transfer cassette is color-coded so the gray side should end up facing the cathode (black electrode) and the clear side facing the anode (red).
- Place a pre-soaked ScotchBrite pad on the gray side of the cassette.
- Place 1 piece of filter paper on top of the ScotchBrite pad.
- Place your gel on top of the filter paper.
- Place a piece of nitrocellulose filter on top of the gel. The nitrocellulose filter is white and can be found between the blue protective paper sheets. Wear gloves when handling the nitrocellulose to avoid transferring proteins from your fingers to the filter.
- Gently but thoroughly press out any air bubbles caught between the gel and the nitrocellulose.
- Place another piece of filter paper on top of the nitrocellulose.
- Place a second pre-soaked ScotchBrite pad on top of the filter paper.
- Close the cassette, then push the clasp down and slide it along the top to hold it shut.
- Place the transfer cassette into the blotting tank so that the clear side faces the red pole and the gray side faces the black pole.
- Two blots can be run in each tank. When both are in place, insert the ice compartment into the tank. Fill the tank with buffer. Connect the power supply and transfer at 100 V for one hour.
- After an hour, turn off the current, disconnect the tank from the power supply, and remove the holders. Retrieve the nitrocellulose filter and confirm that the pre-stained markers have transferred from the gel to the blot. Cut the blot next to each ladder so that you end up with two blots, each only three or four lanes wide (not losing track of whose side is whose!), and then move each mini-blot to blocking buffer and store it in the refrigerator until next time.
The teaching staff will do step #4 for you if it runs past 5 pm.
Part 5: Reverse engineering pMax-BFP-MCS
Understanding existing pMax-BFP plasmid
- Begin by downloading the pMax-BFP file and opening it in ApE. This plasmid is a recent iteration of the BFP component in the NHEJ assay developed in Samson lab.
- Also open the product page for the original pMax cloning vector from Lonza, found here.
- You will use the above two resources to answer several questions below.
- What three components are related to propagating the plasmid in bacteria? Why is propagation in bacteria useful?
- What three components are related to plasmid expression in mammalian cells? What is the purpose of each?
- Look at the product page MCS, and compare it to the MCS that you view in ApE. How do they differ? Let's break this question down further.
- What restriction site(s) have been deleted? (See hint below!)
- What restriction site(s) have been added? (Ditto.)
- Hint: choose "Graphic Map + U" under the Enzymes file menu to see single-cutters. Use base-pair numbers and feature labels, along with the Lonza MCS sequence, to orient yourself.
- Recall that mammalian cells require a Kozak sequence for translation. Can you find one here?
- So, by now you should see that much of the original MCS has been deleted.
- This plasmid was prepared from pMax in two steps. First, the MCS deletion was done. Next, the insertion of BFP and an additional restriction site was done. Of the sites remaining in the MCS, which appear to have been used to clone in BFP?
- What's special about the cut topology of the newly introduced restriction site that is present in the BFP insert? How does it differ from those remaining in the truncated pMax MCS?
- To answer this question, you will need to look up the individual restriction enzymes on the New England Biolabs website. For example, the page for KpnI is found here
- Note that NEB indicates cut sites for the restriction recognition sequence with small triangles.
(Re)-designing pMax-BFP-MCS plasmid
By now you should see that pMax-BFP has a restriction site that produces blunt ends upon cutting. With respect to assay ease of use, blunt ends are convenient because the associated DNA requires minimal or even no purification after digestion. In contrast, after digestion that produces cohesive ends, it is wise to get rid of the tiny fragments, lest they fill back in on the plasmid and change re-ligation efficiency.
For our study, however, we are interested in comparing many different kinds of digested ends. Given that we are performing extra steps anyway, wouldn't it be nice to be confident that we successfully cut our DNA? Based on techniques you learned in Module 1, think about how you might accomplish this validation. We'll come to the answer a little later.
In order to create various digested ends, we will design an extended MCS. It is important that the restriction enzymes that we pick don't occur somewhere else in pMax or in the BFP code-determining sequence. Why? To find these restriction enzymes, we will use another tool from NEB, namely NEBcutter.
- Copy the pMax-BFP sequence into the query box. You can keep most of the default parameters, but one should stand out to you as "wrong." Adjust it, name your project, and then submit the sequence.
- View the "0 cutter" enzymes, i.e., the enzymes that don't cut this plasmid. How many restriction sites (of those recognized by NEB-available enzymes) are absent from pMax-BFP? How many of these are cut in a blunt fashion?
- Assuming that you will need about ten restriction sites in your MCS, how would you possibly go about narrowing this zero-cutter list down? Brainstorm a few ideas with your partner before reading ahead.
- In fact, the following considerations entered into our design decisions
- enzymes that are all or mostly all compatible in the same buffer
- enzymes that are relatively common (i.e., have uses in other lab modules)
- enzymes that we already had in the lab
- enzymes that avoid long stretches of AT or GC
- enzymes that don't require many base-pairs on either side of the recognition site for successful cutting see here
We eventually jettisoned the last consideration, when in a planning meeting we came up with an idea for validating DNA cutting. Did you think of any ideas yet? Take a moment to brainstorm… We decided that instead of making one MCS, we would make two, and that these would be separated by a nonsense DNA fragment large enough to see on a gel. For example, we might cut with a sticky enzyme in the left-hand MCS (hereafter MCS1), and a blunt enzyme in the right-hand MCS (hereafter MCS2), creating a hybrid break site. Afterward, the DNA could be run on a gel, both to validate cutting (by observing release of the nonsense fragment), and to isolate/purify the cut plasmid reporter. But how should we deal with a more simple cut topology, such as sticky alone or blunt alone? Aha! The same restriction site should be present in both MCS1 and MCS2.
- With the above design strategy in mind, download the MCS file and open it in ApE.
- The very end of each MCS (5' of MCS1 and 3' of MCS2) was used to clone the dual-MCS construct into pMax-BFP. That is, the MCS file represents the insert and the pMax-BFP file represents the vector.
- What enzyme recognizes this end sequence that was used for cloning?
- Hint: just try Googling the first 6 bp of the MCS.
- Given the information you encountered in the design considerations above, what do you think the purpose of the "cut cap" is?
- What enzyme recognizes this end sequence that was used for cloning?
- Manually "clone" the MCS construct into the pMax-BFP file, and save it with a new name, such as pMax-BFP-MCS. What is the total size of the new plasmid?
- With your mouse/trackpad, highlight from the end of MCS1 to the end of MCS2, encompassing the whole insert. Under the Enzymes file menu, choose Selection Only.
- Next, go to Enzymes → Enzyme Selector. Choose "unique" from the lower drop-down menu, and press "Select." Next, choose "equal to 2" and again press "Select." Finally, press "Graphic Map" (not "Graphic Map + U," as "U" will override your previous selections and show only unique enzymes).
- If you wish to, print this graphic for your reference. Note that because of the handy menus in ApE, we are viewing only the insert part of pMax-BFP-MCS, but we are being told (in parentheses) the number of restriction sites for a given enzyme that exist in the entire plasmid. Otherwise, you would need to cross-check against the zero-cutters list to cross these out.
- Paying attention only to the true single- and dual-site enzymes, design a digest to prepare each of the following topologies, using a single enzyme or pair of enzymes for each:
- sticky ends, 5' overhang
- sticky ends, 3' overhang
- blunt ends
- 5' sticky end on upstream side, blunt end on downstream side
- 3' sticky end on upstream side, blunt end on downstream side
- sticky ends that are topologically compatible, but that have one or more sequence mismatches
- sticky ends that are topologically incompatible (overhang is on top strand for both, for example).
- For the double digests, try to choose enzymes that are compatible with the same buffer. The NEB buffer chart can be found linked here. Sometimes using high fidelity (HF) enzymes will be a better bet than the originals, and sometimes a worse bet.
Homework
Due M2D3
- Complete the activity in Part 5 of M2D2. Choose what cut type you will make by signing up on the M2D2 Talk page.
- The following calculations are required for your M2D3 lab day. We will not grade this assignment, but we will check that you have done them before class.
- Using the NEB website, perhaps starting with the enzyme finder or another tool, plan your digest for next time. You should aim to meet the following conditions:
- Digesting in the most optimal buffer for the enzyme or set of enzymes, at the optimal temperature
- Note that all NEB buffers are supplied as 10X concentrates
- Digesting with 2.5 U of each enzyme per μg of DNA
- Digesting 7.0 μg of DNA
- The DNA is at 907 ng/μL
- Preparing a 25 μL total reaction volume
- Pipetting no less than 1 μL of enzyme at a time.
- Note that you may need to prepare an intermediate dilution of some enzymes. To determine stock concentration, note that the "S" size was always bought.
- Alternatively, you can prepare a double-size (or greater) "master mix" for your reaction with excess enzyme, buffer, and water, and then mix a fraction of said mix with the DNA.
- In sum, you should name the reaction buffer, the reaction temperature, and the volumes of DNA, of buffer, of enzyme, and of water that you will use for your reaction.
- Digesting in the most optimal buffer for the enzyme or set of enzymes, at the optimal temperature
2. Finally, recall that your primer design memo is due by 10 PM on the day of your next lab session (Thursday or Friday depending on section).
Reagent list
From Boston Bioproducts unless otherwise noted:
- RIPA Lysis Buffer
- 50 mM Tris-HCL, pH 7.4
- 150 mM NaCL
- 1% NP-40
- 0.5% Sodium deoxycholate
- 0.1% SDS
- 100X Protease Inhibitor cocktail
- AEBSF
- Aprotinin
- E-64 Besstain Leupeptin
- EDTA
- Precision Red Advanced Protein Assay (Cytoskeleton, Inc.)
- 6x Reducing Laemlli Sample Buffer
- 375 mM Tris HCL, pH 6.8
- 9% SDS
- 50% Glycerol
- 9% Betamercaptoethanol
- 0.03% Bromophenol blue
From Bio-Rad
- 4-20% Mini-PROTEAN TGX gel
- TGS Buffer: 25 mM Tris, 192 mM glycine, 0.1% (w/v) SDS, pH 8.3
- Dual Color marker sizes here
- Transfer Buffer
- 25 mM Tris
- 192 mM Glycine
- 20% v/v Methanol
Navigation Links
Next Day: Complete Western and prepare damaged DNA Previous Day: Introduction to cell strains and plating