20.109(S14): Choose system conditions and paper discussion (Day3)
Introduction
Today you will become intimately familiar with the plasmid reporter assay for measuring NHEJ that we will use in Module 2. You will examine the different elements present in the plasmid, and more importantly you will try to reconstruct some of the design choices that were made when different variants of the reporter were conceived. We will also continue learning about Ku80, this time through a class-wide discussion of a recent research article on the role of Ku80 in lung cancer.
A key topic that you will need to understand to complete today’s (re-)design exercise is the function and (sequence-level) structure of restriction enzymes. Restriction endonucleases, also called restriction enzymes, cut (“digest”) DNA at specific sequences of bases. The restriction enzymes are named for the prokaryotic organism from which they were isolated. For example, the restriction endonuclease EcoRI (pronounced “echo-are-one”) was originally isolated from E. coli giving it the “Eco” part of the name. “RI” indicates the particular version on the E. coli strain (RY13) and the fact that it was the first restriction enzyme isolated from this strain.
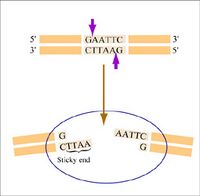
The sequence of DNA that is bound and cleaved by an endonuclease is called the recognition sequence or restriction site. These sequences are usually four or six base pairs long and palindromic, that is, they read the same 5’ to 3’ on the top and bottom strand of DNA. For example, the recognition sequence for EcoRI (see also figure at right) is
5’ GAATTC 3’
3’ CTTAAG 5’

Notice that EcoRI leaves a 5' overhang on either DNA strand; some restriction enzymes leave 3' overhangs instead. Still other restriction enzymes cut precisely in the middle of the palindromic DNA sequence, thus leaving no overhangs after digestion. As you may recall from Module 1, the single-stranded overhangs resulting from DNA digestion by enzymes such as EcoRI are called "sticky" or cohesive ends, while double-stranded ends resulting from digestion by enzymes such as HaeIII are called blunt ends. HaeIII recognizes
5’ GGCC 3’
3’ CCGG 5’
In its simplest description, our reporter assay works as follows: a blue-fluorescent-protein-expressing plasmid is cut by a restriction enzyme(s), then transfected into cells, and repaired at some frequency that we evaluate by measuring the blue fluorescence of said cell population. In fact, the assay design is more subtle than this description encompasses in several respects. The high-level design considerations that we take into account include accuracy of the assay (both technical and biological) and its ease of use. To be confident of the technical accuracy, the assay design includes a second reporter – this time green fluorescent protein – as a transfection control. There is also a control for variable expression of GFP versus BFP, which we will discuss in more detail on Day 5. As another bit of internal validation, one that helps us avoid wasting time with failed products, we have a way to test for successful digestion of the BFP plasmid. Finally, as an example of ease of use, in cases where we cut with multiple restriction enzymes, convenience demands that they are compatible with the same buffer system. We'll dig into some of these considerations further below, and others when we return from spring break.
Protocols
Part 1: Paper discussion
As described in the Day 2 FNT, we will be discussing the paper from Zhang lab in class today.
Technical Background
The main technical topic that may be unfamiliar to (some of) you is the use of short interfering RNA (siRNA). RNA interference, or RNAi, is a post-transcriptional silencing mechanism. You may have heard of gene "knock-outs" before, and in fact xrs6 are essentially Ku80 knock-outs: the DNA sequence is deleted or modified so as to be non-functional. In contrast, RNAi is usually not as potent, and hence is called gene "knock-down." Here the DNA sequence is normal, and mRNA is transcribed – but it is bound up or destroyed before it can be translated. You can read more about the intricacies of RNAi at Scitable.
Discussion Topics
Writing
- How does this abstract style differ from the one you have previously encountered – and also explicitly been taught – in 20.109? How is it the same? What are the pros and cons of each format? As a whole, did the abstract make you want to read the paper?
- This Introduction is short but packs a punch. With the 20.109 guidelines in mind, locate the key elements of an introductory section here. What one sentence best defines the research gap?
- Let's pay special attention to the Methods section, since you will be writing one during Module 2!
- What best practices for Methods section writing do the authors follow? For example, do they write clear topic sentences of appropriate scope?
- What is the authors' strategy for sub-section groupings? What is the role of the first sub-section?
- What methods do the authors seem to assume that most readers will be familiar with?
- What methods do the authors seem to assume need more detailed definition or citation?
- What is the purpose of a passage beginning "Briefly, …"?
- Are there any changes you would suggest the authors make?
- We'll talk about the writing style in the Results and Discussion sections in tandem with our conversation about the technical content. Remember to keep in mind
Content
The following questions will guide our in-class discussion; consider them as a starting point rather than a check-list.
- Figure 1 (Note: caption mis-states which letter label corresponds to which image)
- Describe the primary finding(s) in this Figure and associated Results text.
- What do you think the purpose is of showing both qualitative and quantitative data?
- Would you have expected, a priori, for A and B to look identical? Why or why not?
- How do you interpret the y axes in figures C and D?
- Figure 2
- Describe the primary finding(s) in this Figure and associated Results text.
- Looking back at the Methods, what did the authors do to obtain as credible data as possible?
- Table 1
- Describe the primary finding(s) in this Table and associated Results text.
- How do these findings relate to the central question of the study?
- Figure 3
- Describe the primary finding(s) in this Figure and associated Results text.
- What is the unit on the x axes?
- Looking back at the methods, define the survival parameters, and explain how the authors dealt with potentially confounding data.
- Table 2
- Describe the primary finding(s) in this Table and associated Results text.
- Would you want to modify this Table to include the information in the main body text? If so, how?
- Figure 4
- Describe the primary finding(s) in this Figure and associated Results text.
- What is the role of the lower strip in each of A and B? How is this information used in panels C and D?
- What is the purpose of switching from investigating primary tumors to investigating a cell line?
- Relatedly, what is the purpose of obtaining the data for Figure 4 before moving on to the experiments in Figure 5?
- Figure 5
- Describe the primary finding(s) in this Figure and associated Results text. How do the purposes of the different panels differ? Which ones might you group together?
- What is the meaning and the role of the "scramble" siRNA?
- There is a one-word mistake in the associated text – where, and what should the word be instead?
- Looking back at the Methods, how do you interpret the different quadrants of the flow cytometry data plot?
- Discussion
- What new information do we learn here? What types of information tend to come up?
Part 2: Reverse engineering pMax-BFP-MCS
Understanding existing pMax-BFP plasmid
- Begin by downloading the pMax-BFP file and opening it in ApE. This plasmid is a recent iteration of the BFP component in the NHEJ assay developed in Samson lab.
- Also open the product page for the original pMax cloning vector from Lonza, found here.
- You will use the above two resources to answer several questions below.
- What three components are related to propagating the plasmid in bacteria? Why is propagation in bacteria useful?
- What three components are related to plasmid expression in mammalian cells? What is the purpose of each?
- Look at the product page MCS, and compare it to the MCS that you view in ApE. How do they differ? Let's break this question down further.
- What restriction site(s) have been deleted? (See hint below!)
- What restriction site(s) have been added? (Ditto.)
- Hint: choose "Graphic Map + U" under the Enzymes file menu to see single-cutters. Use base-pair numbers and feature labels, along with the Lonza MCS sequence, to orient yourself.
- Recall that mammalian cells require a Kozak sequence for translation. Can you find one here?
- So, by now you should see that much of the original MCS has been deleted. We will save a deeper discussion of MCS considerations for after spring break. For now, know that "extra" sequences in a plasmid may sometimes affect expression of a downstream coding sequence.
- This plasmid was prepared from pMax in two steps. First, the MCS deletion was done. Next, the insertion of BFP and an additional restriction site was done. Of the sites remaining in the MCS, which appear to have been used to clone in BFP?
- What's special about the cut topology of the newly introduced restriction site that is present in the BFP insert? How does it differ from those remaining in the truncated pMax MCS?
- To answer this question, you will need to look up the individual restriction enzymes on the New England Biolabs website. For example, the page for KpnI is linked here: https://www.neb.com/products/r0142-kpni.
- Note that NEB indicates cut sites for the restriction recognition sequence with small triangles.
(Re)-designing pMax-BFP-MCS plasmid
By now you should see that pMax-BFP has a restriction site that produces blunt ends upon cutting. With respect to assay ease of use, blunt ends are convenient because the associated DNA requires minimal or even no purification after digestion. In contrast, after digestion that produces cohesive ends, it is wise to get rid of the tiny fragments, lest they fill back in on the plasmid and change re-ligation efficiency.
For our study, however, we are interested in comparing many different kinds of digested ends. Given that we are performing extra steps anyway, wouldn't it be nice to be confident that we successfully cut our DNA? Based on techniques you learned in Module 1, think about how you might accomplish this validation. We'll come to the answer a little later.
In order to create various digested ends, we will design an extended MCS. It is important that the restriction enzymes that we pick don't occur somewhere else in pMax or in the BFP code-determining sequence. Why? To find these restriction enzymes, we will use another tool from NEB, namely NEBcutter.
- Copy the pMax-BFP sequence into the query box. You can keep most of the default parameters, but one should stand out to you as "wrong." Adjust it, name your project, and then submit the sequence.
- View the "0 cutter" enzymes, i.e., the enzymes that don't cut this plasmid. How many restriction sites (of those recognized by NEB-available enzymes) are absent from pMax-BFP? How many of these are cut in a blunt fashion?
- Assuming that you will need about ten restriction sites in your MCS, how would you possibly go about narrowing this zero-cutter list down? Brainstorm a few ideas with your partner before reading ahead.
- In fact, the following considerations entered into our design decisions
- enzymes that are all or mostly all compatible in the same buffer
- enzymes that are relatively common (i.e., have uses in other lab modules)
- enzymes that we already had in the lab
- enzymes that avoid long stretches of AT or GC
- enzymes that don't require many base-pairs on either side of the recognition site for successful cutting (see https://www.neb.com/tools-and-resources/usage-guidelines/cleavage-close-to-the-end-of-dna-fragments)
We eventually jettisoned the last consideration, when in a planning meeting we came up with an idea for validating DNA cutting. Did you think of any ideas yet? Take a moment to brainstorm… We decided that instead of making one MCS, we would make two, and that these would be separated by a nonsense DNA fragment large enough to see on a gel. For example, we might cut with a sticky enzyme in the left-hand MCS (hereafter MCS1), and a blunt enzyme in the right-hand MCS (hereafter MCS2), creating a hybrid break site. Afterward, the DNA could be run on a gel, both to validate cutting (by observing release of the nonsense fragment), and to isolate/purify the cut plasmid reporter. But how should we deal with a more simple cut topology, such as sticky alone or blunt alone? Aha! The same restriction site should be present in both MCS1 and MCS2.
- With the above design strategy in mind, download the MCS file and open it in ApE.
- The very end of each MCS (5' of MCS1 and 3' of MCS2) was used to clone the dual-MCS construct into pMax-BFP. That is, the MCS file represents the insert and the pMax-BFP file represents the vector.
- What enzyme recognizes this end sequence that was used for cloning?
- Hint: just try Googling the first 6 bp of the MCS.
- Given the information you encountered in the design considerations above, what do you think the purpose of the "cut cap" is?
- What enzyme recognizes this end sequence that was used for cloning?
- Manually "clone" the MCS construct into the pMax-BFP file, and save it with a new name, such as pMax-BFP-MCS. What is the total size of the new plasmid?
- With your mouse/trackpad, highlight from the end of MCS1 to the end of MCS2, encompassing the whole insert. Under the Enzymes file menu, choose Selection Only.
- Next, go to Enzymes → Enzyme Selector. Choose "unique" from the lower drop-down menu, and press "Select." Next, choose "equal to 2" and again press "Select." Finally, press "Graphic Map" (not "Graphic Map + U," as "U" will override your previous selections and show only unique enzymes).
- If you wish to, print this graphic for your reference. Note that because of the handy menus in ApE, we are viewing only the insert part of pMax-BFP-MCS, but we are being told (in parentheses) the number of restriction sites for a given enzyme that exist in the entire plasmid. Otherwise, you would need to cross-check against the zero-cutters list to cross these out.
- Paying attention only to the true single- and dual-site enzymes, design a digest to prepare each of the following topologies, using a single enzyme or pair of enzymes for each:
- sticky ends, 5' overhang
- sticky ends, 3' overhang
- blunt ends
- 5' sticky end on upstream side, blunt end on downstream side
- 3' sticky end on upstream side, blunt end on downstream side
- sticky ends that are topologically compatible, but that have one or more sequence mismatches
- sticky ends that are topologically incompatible (overhang is on top strand for both, for example).
- For the double digests, try to choose enzymes that are compatible with the same buffer. The NEB buffer chart can be found linked here. Sometimes using high fidelity (HF) enzymes will be a better bet than the originals, and sometimes a worse bet. Note that due to an instructor (ahem!) forgetting about a change in NEB's buffer system, a couple of enzyme combinations were ill thought out in the original design, and will require the use of a buffer that has only 50% activity for one of the enzymes.
Part 3: Choose system conditions
On today's Talk page, please sign up for one of the seven cut topologies that are available. We will confirm which enzymes result in which topologies soon, so you can complete your FNTs, but for now don't worry about this detail.
For next time
- Using the NEB website, perhaps starting with the enzyme finder or another tool, plan your digest for next time. You should aim to meet the following conditions:
- Digesting in the most optimal buffer for the enzyme or set of enzymes, at the optimal temperature
- Note that all NEB buffers are supplied as 10X concentrates
- Digesting with 2.5 U of each enzyme per μg of DNA
- Digesting 3.5 μg of DNA
- The DNA is at 907 ng/μL
- Preparing a 25 μL total reaction volume
- Pipetting no less than 1 μL of enzyme at a time.
- Note that you may need to prepare an intermediate dilution of some enzymes. To determine stock concentration, note that the "S" size was always bought.
- Alternatively, you can prepare a double-size (or greater) "master mix" for your reaction with excess enzyme, buffer, and water, and then mix a fraction of said mix with the DNA.
- In sum, you should name the reaction buffer, the reaction temperature, and the volumes of DNA, of buffer, of enzyme, and of water that you will use for your reaction.
- Digesting in the most optimal buffer for the enzyme or set of enzymes, at the optimal temperature
Reagent list
Just your brains!
Navigation Links
Next Day: Complete Western and prepare damaged DNA Previous Day: Begin Western protein analysis
