20.109(S14):Data analysis (Day7)
Introduction
We hope that you’ll leave lab today with a sense of accomplishment, after inspecting your raw flow cytometry data and then calculating NHEJ repair values. Unfortunately or excitingly – depending on your perspective – it turns out in scientific research that the hard work is just beginning once the data is quantified! Interpreting the data and drawing (sometimes tentative) conclusions requires deep reading and thinking – a process that shouldn’t be rushed. To help get you started, let’s review a few key elements of DNA repair and of the NHEJ pathway in particular.
Our discussions of NHEJ have primarily been focused on Ku80 and DNA-PKcs, along with a few other key players such as Ligase IV, and will remain so here. However, the schematic below from the review by Grundy et al. highlights that many accessory molecules have their own roles to play. Recall that Ku80, with its dimer partner Ku70, completes the first step of NHEJ repair by binding to DNA double-strand breaks (DSBs). Keep this function in mind as you consider how a Ku80 knockout strain might respond to DNA damage. The Ku 80/70 dimer quickly recruits DNA-PKcs, the catalytic subunit of DNA-dependent protein kinase, to form the DNA-PK complex. Although DNA-PKcs has some binding affinity for DSBs, this affinity is increased by two orders of magnitude by the presence of the Ku dimer, and the kinase activity itself cannot proceed in the absence of Ku. Interestingly, although the kinase activity of DNA-PK is known to be important, it is not certain precisely which phosphorylation events are absolutely required for NHEJ.
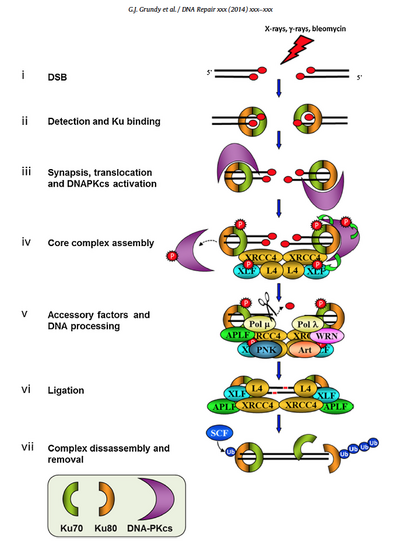
To better understand how DNA-PKcs may differ from Ku80 as a target for changing a cell’s DNA damage response, let’s take a closer look at Compound 401. A number of small molecule inhibitors of DNA-PK were discovered by Griffin et al. in the early 2000s. While both Griffin and the commercial vendor Tocris/R&D Systems refer to the inhibitors generically as DNA-PK rather than DNA-PKcs inhibitors per se, it is clear that the "cs" is implied and kinase activity is what is being inhibited. The inhibitor is described as “ATP-competitive,” indicating that it competes for the ATP binding pocket in the kinase domain of DNA-PK. Moreover, the Griffin inhibition assay used p53 phosphorylation as a readout, rather than phosphorylation after binding to Ku 70/80. Finally, the inhibitor also acts on the unrelated kinase mammalian target of rapamycin (mTOR).
The one piece of wet lab work that you will do today is completing the C401 validation assay. Clonogenic assays of mammalian cells have over a 50 year history, as mentioned in the methods paper by Franken et al. They are useful for assessing the reproductive capacity of cells after irradiation and other types of damage. We will diverge somewhat from the Nature Protocols paper, but it is useful for introducing terms such as the plating efficiency and the surviving fraction. Specifically, we do not need to fix our cells in an independent step, because the stain that we will use contains methanol. (Correction! Our stain contains very little methanol, so fixing does not appear crucial for short-term staining.) Second, we will not use the crystal violet stain, which binds DNA, but instead a Coomassie derivative, which targets proteins. In fact, you may recognize Coomassie as the go-to stain for SDS-PAGE. Protein binding by the dye occurs primarily via arginine, as well as other basic and aromatic residues, as described here. We will use a variant of the original Coomassie Brilliant Blue stain called BioSafe Coomassie.
Most of your time today will be spent at the computer, quantifying flow cytometry data. Recall from the M2D5 introduction that we will proceed in three main steps.
First, reporter expression for GFP and BFP alike will be calculated by multiplying percentage of positive cells by fluorescence intensity (FI). Here we have a choice of whether to use mean, geometric mean, or median fluorescence intensity. Median fluorescence is least susceptible to being influenced by a few outliers, while geometric mean is generally more appropriate for log scale data than arithmetic mean. For normally distributed populations, all three values should be pretty similar. In practice, we have found that while mean and median FI are very different values, after normalization the ultimate NHEJ repair values are quite similar.
The second step is to calculate the ratio of BFP to GFP reporter expression for each sample. Whereas during pilots this ratio tended to be greater than 1.0, a preliminary look at our class-wide data suggests that it is now very close to 1.0. This shift could easily be caused by periodic adjustments to and calibration of the flow cytometry equipment, including the lasers. In fact, during set-up with the instructor samples, the voltage for the FSC as well as a more complex scatter parameter that we haven’t discussed both had to be adjusted from pilot values. This outcome highlights why it’s so important – perhaps when taking your own flow data some day – to perform both flow controls (negative and single color) and experimental controls (here dual intact) every single time one does an experiment!
The final step is divide the damaged-BFP:GFP ratio by the maximal possible “repair,” namely the intact-BFP:GFP ratio. Convince yourself that this parameter essentially provides the fraction of BFP plasmids repaired.
We should share with you one final "behind-the-scenes" complication that we know about this newly developed assay, just in case it is illuminating as you interpret your results. Two of the seven damage topologies in the originally designed pMax-BFP-MCS plasmid showed no fluorescence in K1 cells. Yikes, we were worried! The amazing Zac Nagel then found a paper suggesting that the G/C content (our old friend from Module 1) of an MCS can influence downstream gene expression. Moreover, depending on the distance from the gene, G/C rich regions can either activate or suppress expression. We decided that it was safest to design an MCS with lower overall G/C content, to make it most likely that any differences you all see in fluorescence recovery of different damage topologies reflect differences in efficiency of repair rather than efficiency of gene expression. (Keep in mind that using different cut sites means that slightly different MCS remnants are left behind on the plasmid for each distinct topology.) That’s right, you’ve been using a second generation pMax-BFP-MCS all along! Isn’t it funny how biology can interfere with engineering?
Protocols
Part 1: Stain irradiated cell colonies
Note: You may perform the staining protocol either today (M2D7) or next time (M3D1) as you prefer. Typical growth times for clonogenic assays are at least one week, and if you stain next time your counting task may be made easier by having bigger colonies. On the other hand, you have built-in class time today for analysis, and may want to work on your dose response curve after you complete the FC analysis. The decision is totally up to you and your partner!
- Briefly observe your irradiated cells on the TC microscopes. Are you able to find some colonies? About how many cells are in some of these small dispersed clusters?
- Take your plate to the main lab for the remaining steps. First, aspirate the media. You don't need to change the yellow tip between samples if you move from highest [C401] dose to lowest, as that should be the order of least to most colonies.
- Rinse each well with about 2 mL of pre-warmed PBS.
- After removing the PBS, use a serological pipet to consistently add 2 mL of Coomassie to each well.
- Place your plate on the fume hood shaker at 80 rpm for about 1 hour.
- Repeat the PBS rinse, this time with room temperature buffer.
- Let the well plate dry for a short time after aspirating the PBS.
- If you wait a very long time the stain will begin to fade.
- Finally, count the colonies in each well and document these on today's Talk page.
- Do your best to apply a consistent standard for threshold colony size and threshold staining intensity. As long as you are consistent, there is not one right answer as to what constitutes a colony.
- One of your instructors finds it easiest to count colonies by making a dot with a lab marker as she counts each colony, right on the underside of the 6-well plate at the colony location, and then writing down each decade (10, 20, etc.) on the plate as she reaches that number so she doesn't lose her place.
- Whether today or on your own, you should plot your data by surviving fraction (as described in the Nature Protocols paper linked above) for the Module 2 report. However, you don’t need to perform a curve fit as we are essentially looking for a yes/no answer to the question of whether the inhibitor worked.
Part 2: Flow cytometry analysis
Overview:
- You will begin by looking at images from the instructor samples to learn how to read the flow cytometry plots and summary statistics.
- Next you will peek at your own images and form preliminary expectations about your data set.
- Finally, you will work in Excel to precisely calculate the NHEJ repair value for each of your three conditions (two replicates each).
Protocol:
- On one of the lab computers, double-click on the FACS server shortcut.
- Alternatively, on your own computer access 18.159.2.11 directly. Ask your instructors for the username and password.
- Go to the April 2014 folder, then to Agi Stachowiak. Copy over both the T/R and W/F image sets to your laptop: the filenames begin "analysis-images" and only the dates differ.
- Copy over just your own day of statistics, unless you really want access to all of the raw data in your back pocket: the .csv filenames begin "analysis-statistics" and only the dates differ.
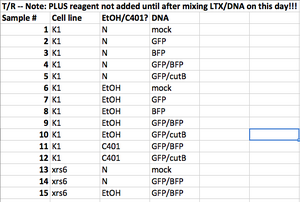
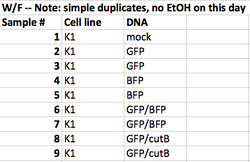
- The instructor samples are listed in the table below. From this table, and from the T/R and W/F image sets, try to address the questions below.
- Background. The scatter data is used – in three steps – to make gate P3, which should consist primarily of live, single cells. From the cells gated in P3, two sub-gates are made that capture all GFP-positive cells ("Green cells" gate) and all BFP-positive cells ("Blue cells" gate). Both singly and doubly positive cells are included in each gate. It is important to read the "% Parent" statistics: these indicate XFP-positive cells as a percentage of all the cells in P3. The "% Total" statistics include debris, aggregates, and clearly dead cells!
- What percent Green cells are in the mock sample on each day? What about Blue cells?
- What percent of singly-transfected cells express GFP? Do within-day and cross-day replicates agree well or not?
- What percent of singly-transfected cells express BFP? Do within-day and cross-day replicates agree well or not?
- What percent of co-transfected cells express GFP? Express BFP? Comparing the Green and Blue gates to Q1 and Q4, about what percent of cells seem co-transfected, versus expressing just GFP, and expressing just BFP?
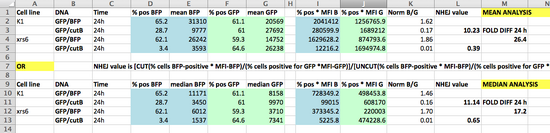
- How is within-day and cross-day replicate agreement for the co-transfected samples? Do the tables below suggest an explanation for why?
- Does ethanol appear to affect scatter profiles? What about affecting GFP, BFP, or co-expression?
- What NHEJ repair value do you calculate for Zac's original BFP plasmid, using the first replicate in the W/F instructor data? Try this calculation by hand, using the mean fluorescence intensity. Later, you can include this data as a check on your Excel worksheet. The value you should calculate is 12.8%.Update: Your instructor picked off GFP mean fluorescence instead of BFP mean fluorescence for the intact case! Here is where computers definitely beat manual picking off of data. The correct number is 8.7%.
- After you understand the instructor data, skim over your 12 sample plots. Can you see apparent differences between K1, K1+401, and xrs6?
- Now that you have a good conceptual understanding of the data, it's time to crunch some numbers. Open the .csv file and save it as a newly named .xlsx file.
- Begin by deleting all of the rows except the twelve containing your own dataset.
- Next delete all of the columns except the few that interest you. Keep in mind that you need to know Green cell and Blue cell gating as a % of the parent gate, P3. Class-wide, you are only required to do your calculations based on mean fluorescence intensity (MFI), to be consistent with Samson lab data. However, you may find it interesting to see whether using median fluorescence intensity gives you the same trends or not. Just a few extra copy-pastes to do both calculations!
- We recommend that you prepare a new Excel file with your NHEJ equations, and just copy-paste in the appropriate % and MFI data; this approach is a versatile one. Your final worksheet might look similar to the screenshot below.
- Remember that for each of the twelve wells you should calculate raw reporter expressions and a BFP/GFP normalized value. Then, for each intact/cut pair you can calculate an NHEJ value. In this way, we should have quadruplicate NHEJ values for most repair topology/cell population conditions, which will allow us to do statistical comparisons.
Reference information:
You must email your Excel sheet to 20109 DOT submit AT gmail DOT com before leaving lab today. We instructors will post a summary file for ease of class-wide data analysis by Wednesday evening or Thursday morning.
Part 3: Statistics practice
You may find averages, standard deviations, and t-tests useful when you report on class results. You will also revisit these topics during Module 3.
You can practice the steps below using the male and female heights that we collect during pre-lab lecture.
- Begin by downloading the following Excel file as a framework to carry out the basic statistical manipulations we discussed in pre-lab lecture. The file is modified from one originally written by Professor Bevin Engelward.
- Find and plot 95% confidence intervals for the male and female heights.
- Compare the means of these two populations. At what confidence level (if any) are they different?
- Would a one-tailed or two-tailed test be more appropriate for this comparison?
For next time
Recall from last time:
- Revise your earlier draft of the Methods section, just through M2D2, applying the feedback you received.
- Prepare the rest of your Methods section (through M2D7) in outline form. Start by considering what methods may be logically grouped together. At a minimum, you should turn in
- sub-section titles,
- topic sentences for each sub-section,
- and a few short phrases indicating what content will be included in that sub-section.
- The phrases do not need to include every single material/concentration/etc. that you will use, but they should convey the scope of that information in very abbreviated form.
- For example, phrases for the first half of a Western sub-section could look like: cell lysis method (RIPA and inhibitors from BBP, scraping, ice 15 min and spin 15 min); Precision Red to measure protein in supernatant; preparation step (add Laemlli, boil equal amounts of protein). The second half would include similar types of phrases to cover the PAGE and transfer steps.
Reagent list
- PBS
- Bio-Safe Coomassie Stain (Bio-Rad)
- Mostly your brains!
Navigation Links
Previous Day: DNA repair assays
