20.109(S13):Context-setting and primer design (Day1)
Introduction
In this module, you'll perform two investigations of pathogens that can infect birds. Through bird stool, these pathogens can be transferred to the environment –– and in some cases infect other animals. Perhaps the most well-known avian pathogen with zoonotic potential (i.e., inter-species transmission) is the flu virus. For your safety, all the samples we will work with have been screened to exclude those carrying human–pathogenic flu strains. However, we will be able to mine much of the same intellectual content that we could were we studying flu directly.
Prof. Runstadler, the lecturer for this module, studies phylogenetic relationships among avian flus (see for example this paper and this one), a research area that can provide information useful for predicting the next flu pandemic and commencing vaccine production in time. Researchers track viral mutations/evolution, infection of different bird species (including co-infection by multiple strains), and the trafficking patterns of those birds. Your own phylogenetic analysis will consist of comparing bacterial communities in two distinct bird populations. (We admit, not as flashy as studying the flu!) More about the significance of that research in the coming week. See also the Module 1 home page for a visual summary of the relevant concepts and applications for both studies.
Your other research project will pertain to an unusual fungus called microsporidia. Specifically, today you will attempt to design PCR primers that improve upon the sensitivity and/or specificity of the state-of-the-art in microsporidia identification. This experience will give you a sense of how successful scientific investigation hinges on the quality of the methods used.
Microsporidia are parasites that require a eukaryotic host cell for propagation; only in the spore stage of its life cycle can this pathogen independently survive. Of the more than 1000 species in the phylum microsporidia, about a dozen can infect humans, and many more infect other animals – including animals used by humans for production of food and materials. To infect their hosts, microsporidia use a unique projection called a polar tube.
Based on the apparent absence or simplification of many canonical eukaryotic features, it was once proposed that microsporidia must be primitive (as in ancient) eukaryotes whose evolution pre-dated mitochondria. It turns out that microsporidia do have “mitosomes,” and rather than being primitive (except in the sense of simple), these mitochondria-like organelles evolved to a reduced form because mitochondrial function has largely been outsourced to the host. Phylogenetic analysis was the linchpin that resolved the competing hypotheses about microsporidial origin! Relational trees similar to the ones that you will make to describe bacterial populations in seagulls revealed that microsporidia evolved later than initially thought and are a fungus rather than protist. If you're interested, you can read more about the fascinating microporidian genome in this linked review by Keeling and Slamovits, or about microsporidia in general at this PLOS Pathogens feature.
A robust diagnostic for identifying infection by microsporidia – microsporidiosis – is of interest for several reasons related to both prevention and treatment. With respect to treatment, identifying the disease-causing agent can sometimes suggest an appropriate course of action, for example if the drug resistance profile of that pathogen is known. With respect to prevention, it is useful to think about potential sources of human infection. Environmental sources associated with microsporidiosis risk include bodies of water. Testing drinking sources and swimming areas for the presence of microsporidia can provide guidance to at-risk populations (here primarily immunocompromised individuals), whether it be to drink bottled water or to avoid traveling to a certain country. It is also speculated that some strains of microsporidia have zoonotic potential; in this case, at-risk populations may need to test their pets or avoid certain types of pets. Here, molecular diagnostics are particularly useful due to existing evidence of strain-dependent infectivity. For example, some genotypes of E. bieneusi infect only humans, others only a non-human animal, and still others will accept either host -- suggesting zoonotic potential. Finally, infections can of course be transmitted from person to person. Here, sensitive diagnostics are especially important because in some cases infections can be asymptomatic but potentially transmissible. Recent reviews about zoonotic potential and microsporidiosis are linked here should you want to learn more.
Traditionally, many pathogens were identified by imaging techniques for examining morphology and culturing techniques for probing phenotype (as described for microsporidia specifically in the review by Garcia. Recently, molecular identification has emerged as a powerful and perhaps less subjective technique. Specific sequences in the pathogens' DNA can be targeted by unique primers and amplified in a polymerase chain reaction (PCR). Notably, there are conflicting reports as to whether sequence-based identification is any more sensitive than imaging when it comes to microsporidia (compare these 2004 and 1999 papers). At the very least, molecular diagnostics are necessary for species-level identification .
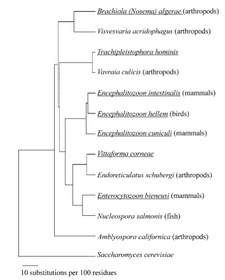
Identification according to sequence requires a highly conserved gene. Ribosomal RNA is an excellent candidate for this strategy: it is essential for life and thus the organism is unlikely to survive rRNA gene mutation. Depending on the pathogen, the small or large subunit or the internal transcribed sequence (ITS) might be the most reliable sequence for identification. The clinical potential of 16S rRNA-based sequencing for bacterial infections is described in the linked review by Dr. Jill Clarridge. For microsporidia, we will start with primers that target the 5' end of the small subunit rRNA. You will have the option of targeting a different portion of the rRNA or the polar tube protein unique to microsporidia, as described in the protocols below. The partial phylogenetic tree of microsporidia shown above is based on rRNA sequences.

Gene amplification by PCR -- whether of 16S rRNA or any other gene -- comprises three main steps: 1) template DNA that contains a sequence of interest is melted, 2) primers anneal to specific locations on the now melted (i.e., single-stranded) DNA, and 3) the primers are extended by a polymerase to synthesize the desired product. Extension occurs at ~70 °C, melting at ~95°C, and annealing at a temperature ~5 °C below the primer melting temperature; thus, the repetition of these steps is called thermal cycling. After each cycle, the newly formed products themselves become templates, causing exponential amplification of the selected sequence. (Note that the early rounds of PCR will not produce the desired product - we will see why in today's pre-lab lecture.)
Based on the numerous applications of PCR, it may seem that the technique has been around forever. In fact it is less than 30 years old. In 1984, Kary Mullis described this technique for amplifying DNA of known or unknown sequence, realizing immediately the significance of his insight.

"Dear Thor!," I exclaimed. I had solved the most annoying problems in DNA chemistry in a single lightening bolt. Abundance and distinction. With two oligonucleotides, DNA polymerase, and the four nucleosidetriphosphates I could make as much of a DNA sequence as I wanted and I could make it on a fragment of a specific size that I could distinguish easily. Somehow, I thought, it had to be an illusion. Otherwise it would change DNA chemistry forever. Otherwise it would make me famous. It was too easy. Someone else would have done it and I would surely have heard of it. We would be doing it all the time. What was I failing to see? "Jennifer, wake up. I've thought of something incredible." --Kary Mullis from his Nobel lecture; December 8, 1983
Protocols
Part 1: Lab practical
You and your partner may work together on the lab practical. (Note: this will not be the case for future quizzes.) You are of course welcome to give different answers should you disagree.
Part 2: Explore existing microsporidia-specific primers
The reference primer set that you will use to amplify microsporidia DNA comes from a 1999 paper from Franzen lab (link). These primers anneal to the 5' end of the small subunit (SSU) rRNA gene.
| Primer name | Direction | Sequence (5'-->3') |
|---|---|---|
| V1 | Forward | CACCAGGTTGATTCTGCCTGAC |
| PMP2 | Reverse | CCTCTCCGGAACCAAACCCTG |
First, let's assess some basic properties of these primers. Then we'll calculate expected product sizes for several different species of microsporidia.
Primer properties
Integrated DNA Technologies, or IDT, sells DNA primers and offers free tools for analyzing them.
You'll also access the National Center for Biotechnology Information (NCBI), a resource that we have used in many different 20.109 modules. Here, you will rely on it for sequence information.
- At the IDT website, under the heading SciTools, choose OligoAnalyzer.
- Keeping the default settings, press the Analyze button and note the resulting length, G/C content, and melting temperature (Tm) of each primer.
- Based on our discussion so far of PCR, are the primer lengths sufficient for a reasonable expectation of sequence uniqueness?
- Are the G/C contents in the appropriate range?
- Now measure and note the Tm with the following settings: [oligo] = 0.2 μM, monovalent salt = 10 mM, divalent salt = 2 mM.
- For this amplification, the annealing temperature (Ta) we are using is 58 °C. Which salt conditions best match this Ta and why?
- Using the appropriate salt conditions, do a self-dimer analysis on each primer and a hetero-dimer analysis of the two primers together.
- It should not be energetically favorable for the primer to dimerize. The delta G for dimers should at least be greater than -9 , and ideally be greater than -5 kcal/mol.
- Annealing of one primer to the 3' end of another primer (or itself) is particularly bad. Why?
- Having a G or C (a "GC clamp") at the 3' end of each primer helps promote complete binding due to the relatively greater hydrogen bonding for GC versus AT. Note that 1-2 bases is best, while 3 or more consecutive GC bases might promote non-specific binding. Do the primers above have such clamps?
- It's best to avoid long stretches of a single base, or even a single type of base (purine or pyrimidine). Are the primers above relatively random or do they break this rule?
- Finally, you would like to test whether these primers bind to a unique site in the genome or instead are promiscuous binders.
- Head over to nucleotide BLAST and copy your first primer into the "Query" box.
- Under Choose Search Set → Organism, start typing Vittaforma corneae; the name should auto-complete as you type.
- Under Algorithm Parameters, be sure "Automatically adjust parameters for short input sequences" is checked.
- Click BLAST and wait for your results.
- The forward primer should identically match the gene that codes for small subunit RNA for its entire length. Find the first gene that is not for SSU rRNA. What is the name of the gene? How many basepairs does the match encompass? Do you expect this annealing site to be a major competitor in your PCR?
- Repeat for the reverse primer.
- When you design your own primers below, you can perform all these same checks on them.
Product sizes
- One microsporidia species that can infect humans is called Encephalitozoon hellem, or E hellem. Find the NCBI accession number AF118143, which shows the SSU rRNA gene sequence for one strain of E hellem.
- To continue with the exercise below, you might find it convenient to paste this sequence intoApE (A plasmid Editor, created by M. Wayne Davis at the University of Utah), which is found on your desktop.
- Look for the forward primer in this sequence. At what basepair do you find it?
- Within ApE, you can use command-F to find matches to sequence fragments.
- Note: V1-PMP2 are designed to broadly target many different species of microsporidia. As you plan your search strategy, keep in mind that any particular strain might have some mismatches to the primer sequence.
- Look for the reverse primer. (What will you have to do to the primer sequence to make it match the coding strand DNA?) At what basepair do you find it?
- Does the sequence have any mismatches to the primer? How many? What about E hellem accession number AF039229?
- What is the total product size expected for each E hellem strain? Are they the same?
- Look for the forward primer in this sequence. At what basepair do you find it?
- Another species of interest is Vittaforma corneae, or V corneae. Repeat the exercise above for accession number JX049115.
- Note: The above accession number will lead you to a partial sequence! For design purposes later, you may need to find the complete sequence.
- Is the product size the same as for E hellem? If different, by how much?
- On a typical DNA gel, do you expect to be able to tell apart fragments of this size? If you have no experience with such gels, talk with the teaching faculty.
- Search on your own for the rRNA gene sequence of the species cuniculi, another member of the genus Encephalitozoon. What product size do you find here?
Part 3: Design novel microsporidia-specific primers
Sign up for the specificity or sensitivity challenge in the table on today's Talk page. Half of the class should try each challenge. You and your partner are designing one primer set together. You may design primers either to the SSU rRNA gene or the polar tube protein. An accession number for the latter to help get you started is AY171240. Shoot for a Tm of 58 ± 1 if possible.
In diagnostics, a two-stage identification system is often used. First, clinicians look for a quick yes/no -- is the organism there? In this stage, sensitivity is prioritized: one would like as few false negatives as possible, even if that means some false positives come up, because the latter can always be excluded in the second stage. Next, clinicians would like to know precisely what species has infected the patient, for both acute treatment of that patient and long-term tracking of infection prevalence across patients. In this stage, specificity is prioritized.
Professor Runstadler takes a very similar approach to flu screening in his lab. All bird samples are broadly checked for flu infection using primers against highly conserved regions of the virus. Samples that come up positive are screened for human-pathogenic strains such as H2, specifically against the "2" version of hemagglutinin. These samples must be handled with extra precautions to avoid infection of lab personnel.
Important note for both challenges
As you work, remember to keep in mind that the forward and reverse primer bind to different strands of DNA. You may find it easiest to work with only the coding strand sequence at first, and then take the complement (for one of the primers -- which one?) during the very last step of your design. When you are happy with your primers, please add them to the table on the "Talk" page, with each primer written from 5' to 3' -- this direction is the standard way that primers are written on an order form.
Feel free to use free software to help guide your design such as that from NCBI or the P3 program.
Specificity challenge
Assume that the V1/PMP2 primer set is appropriate for a first-stage diagnostic. Design a second-stage diagnostic that can specifically identify V corneae from a complex mixture. In reality, your complex mixture will include only one other strain of microsporidia, E hellem, which you want to avoid amplifying. You will test your primer set against each organism separately and a mixture of the two, and compare your results with the default primer set.
Occasionally, co-infection with multiple strains of an organism may occur. If you are able to design primers that better resolve V corneae and E hellem when they coexist in one solution -- that is, the expected products differ by more than 40 bp -- this could also be an interesting approach to the challenge!
Sensitivity challenge
This challenge will require somewhat more luck than the one above. Your goal is to design primers that can detect a lower DNA concentration than V1-PMP2 can. The default primer set reliably amplifies less than 1 ng in a 50 μL reaction, which is near the lower limit expected for basic PCR. You might be able to improve the amplification somewhat by designing primers that vary according to the parameters you explored in Part 2. These days, folks often ask computers to design several primers for them, and then empirically test which primer set works best. Today we ask you to take the role of the computer and see how you stack up.
For the purposes of this exercise, you should design broadly functional primers that can target either E hellem or V corneae. Note that the latter species produces a substantial amount of off-target PCR product (incorrect size) in addition to the expected PCR product when using V1-PMP2. Thus, the primers you design might end up improving a specificity of sorts -- not for targeting a particular species, but for targeting a particular site in the genome (of one or more species).
For next time
- Continue to familiarize yourself with the workings of OpenWetWare. Specifically, you should:
- Create a user page, add some content to it, and add it to the 20.109(S13):People page. Be careful about how you put your e-mail address on your page; make sure to spell out "at" or otherwise break your address up so it can't be scraped up by spambots. It is fine to use the sample OWW user page as a starting point, but be sure to erase or replace information that does not pertain to you (such as someone else's publications!) and erase or replace general content such as "Research Interest 1" with your own interests.
- Complete the student registration/questionnaire from this link to turn in next time. Click on the "edit" tab of this page, then copy the "source code" you see to your user page, fill out the form, and print it. You do not need to keep the information on your user page after printing it.
- The major assessment for this module will be a laboratory report. Start to familiarize yourself with its expected structure and content at this link.
- One week from today (Mod 1, Day 3), we will have an in-class discussion about an article from the primary scientific literature. You might begin reading and thinking about the paper now, rather than trying to do so only in the two days between Day 2 and Day 3. The paper is linked from PubMed here.
- Finish your EHS training if you haven't already.
Reagent list
Nothing yet (except your brains)!