20.109(S09): Luciferase assays and RNA prep (Day6)
Introduction
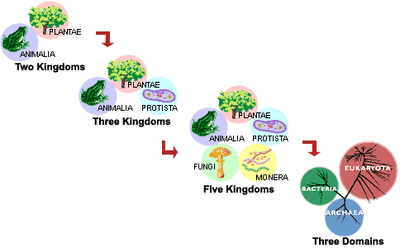
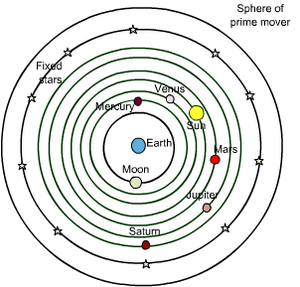
Facts vs Data...plus a measure of measurement
Three basic questions drive all scientific efforts: what’s here, how did it get here, and how does it work…three simple questions that have given rise to a mind-numbing number of textbooks, many of which are themselves mind-numbing. How sad. It’s too easy to open a science textbook and read statements that seem either obvious (e.g. all living things are made of cells) or unbelievably detailed (e.g. poliovirus is a positive-stranded RNA virus so its genome can serve as mRNA). Scientific ideas, even the “no duh” ones, rely on evidence that has been extensively examined and logically interpreted. That’s what makes it science. There may be other ideas and explanations for how things work but when the preponderance of evidence supports an idea, it becomes the science. The process of discovery, driven by data collection and interpretation, is such an integral part of science that it’s surprising how rarely it gets taught. It’s easy to forget that science is a human endeavor, advanced by curious and hardworking individuals and communities, all hoping to improve our understanding of the natural world.
Aristotle (~300 BC) observed nature directly and since then science has presumed that we can learn about the world by collecting evidence. Gathering data has become so integral to science that it seems an obvious part. However when Aristotle started looking at the world, he concluded that
- all organisms were either animals or plants
- plants were less complex than animals and so ranked “lower” on some life ladder
- living things spontaneously arise from non-living things
Read that last point again. Spontaneous generation seemed a sensible idea to wise old Aristotle based on what was observable. And amazingly this completely wrong theory was in place for nearly 2000 years. Rudolph Virchow didn’t state that “where a cell exists, there must have been a preexisting cell” until 1858…not even 200 years ago!
Coupled with data collection is instrumentation and technology. Advances in technology often lead to scientific progress. (e.g., microscopes that heralded the cell theory) and technology also depends on improved scientific descriptions (e.g., the wave and particle theories of light leading to the development of new microscopes). Instruments serve as extensions of our five senses to gather data. As such, new instruments can be like new ways of testing and seeing the world. Combining instruments that can perform high-throuhput assays with bioinformatics tools to interpret the large quantity of resulting data can be especially powerful.
Scientific work generates evidence-based, internally consistent, and well-tested explanations. Gathering data is a big part of the fun and a big part of what you’ll do over the next few lab sessions. In today’s lab we will collect protein-level evidence about the ability of the siRNA you’ve designed to reduce the expression of the Renilla luciferase gene in MES cells. This experiment will test whether the RNAi machinery is functioning in our MES, and also teach us about the varying efficacies of different siRNAs against the same target. Next time you will test for direct knockdown as well as indirect effects of your validated mouse gene siRNA at the message level. The first step in the process will be extracting RNA from the MES.
You will isolate RNA by using a silica (SiO2) column. The column is packed with a silica resin (i.e., beads). The beads have a high ratio of surface area to volume and contain small pores, both of which qualities allow them to interact with specific molecules. When nucleic acids are diluted in a high concentration of a chaotropic salt buffer, they will tend to bind to the silica. This is because chaotropic salts (such as guanidine isothiocyanate) disrupt hydrogen-bond organization between water and macromolecules, essentially dehydrating the nucleic acids and causing them to bind to the resin. Ethanol further precipitates the nucleic acids. The column-bound acids are washed with various buffers to remove salts and other contaminants before finally eluting in pure water, in which nucleic acids are highly soluble. The exact pore size and surface chemistry of the silica beads determine what sizes and kinds of nucleic acid will be bound versus washed away. In our case RNA longer than 200 bp should be isolated, thus enriching for mRNA. Note that prior to isolating RNA, the cells must be lysed and homogenized. Lysis releases the contents of the cells, while homogenization reduces the viscosity of the cell solution by shearing large macromolecules that make the solution difficult to work with.
Protocols
Part 1: Preparing cell lysates
Half the class will begin in the TC room, and the other half will have to wait a short while. If you are in the second group, in the meantime skip ahead to Part 3 and begin preparing your bench for RNA work.
- Bring your memory cards for the digital cameras to the TC room.
- Retrieve your cells from the incubator and observe them on one of the microscopes.
- Take digital photographs of the cells in wells transfected with plasmid and siRNAs. Make sure to note the magnification you are using.
- When you get back to lab, email these photographs to you and your lab partner or post them to your userpage.
- Aspirate the media from the cells of your six-well dishes and wash 2X with 2 ml PBS.
- Lyse the cells in each well by adding 500 ul PLB, a reagent sold by Promega that will lyse the cells into a buffer compatible with the luciferase assay reagents. Incubate on the orbital shaker in the main teaching lab for 15 minutes at 150 rpm.
- Move the lysates to RNase-free eppendorf tubes with clean pipet tips. Proceed with Part 2 or Part 3 first, depending on equipment availability. Keep the samples for RNA extraction on ice.
Part 2: Luciferase assays
These assays should be performed as you did last time, on all 10 samples for which you have duplicates. Refer to that protocol for any particulars of the assay that you may have forgotten. The luminometers will be set up at the teaching bench.
Part 3: Isolation of total RNA
Samples
For your microarray experiment, you will only be able to test 2 of your samples. One will be the experimental sample that received siRNA against a mouse gene, and the other will be the scrambled siRNA control (neither of which were transfected with the luciferase plasmid).
Preparation for working with RNA
RNA is strikingly different from DNA in its stability. Consequently it is more difficult to work with RNA in the lab. It is not the techniques themselves that are difficult; indeed, many of the manipulations will seem identical to those used for DNA. However, RNA is rapidly and easily degraded by RNases that exist everywhere. There are several rules for working with RNA. They will improve your chances of success. Please follow them all.
- Use warm water on a paper towel to wash lab equipment, like microfuges, before you begin your experiment. Then wipe them down with “RNase-away” solution.
- Wear gloves when you are touching anything that will touch your RNA.
- Change your gloves often.
- Before you begin your experiment clean your work area, removing all clutter. Wipe down the benchtop with warm water then “RNase-away,” and then lay down a fresh piece of benchpaper.
- Use RNA-dedicated solutions and if possible RNA-dedicated pipetmen.
- Start a new box of pipet tips and label their lid “RNase FREE.”
Qiagen sells a kit for isolating RNA and we will be using their protocol and reagents.
Protocol
- Add 400 ul of Buffer RTL with BME to the two experimental samples you will study by microarray. This should be done in the fume hood since the reagent contains β-mercaptoethanol which smells awful.
- Collect two Qiashredder columns from one of the teaching faculty. The lysates must be passed through these columns to remove particulate matter. Load the top of each Qiashredder column with the cell lysates. The remainder of today’s experiment can be performed in the main teaching lab, but remember to wear gloves throughout to protect your RNA samples from RNases on your hands.
- Microfuge the Qiashredder columns for 2 minutes. Move the flow-through into two properly-labeled eppendorf tubes.
- Add 1 volume (approximately 600 ul) of 70% ethanol to each of the cleared lysates. Invert 3 or 4 times to mix the contents. Do not vortex. A precipitate may form at this stage but it will not effect your RNA isolation.
- Collect two Rneasy minicolumns from the teaching faculty. Apply 700 ul of each sample to the columns and microfuge for 15 seconds. Discard the flow-through into a conical tube labeled something like "temporary waste stream." Keep the collection tube.
- Apply any remaining sample to the columns. Microfuge and discard the flow-through as before.
- Add 700 ul Buffer RW1 to each column. Microfuge for 15 seconds. Discard both the flow-through and the collection tube.
- Transfer each column to a fresh 2 ml collection tube and add 500 ul Buffer RPE to each column. Microfuge for 15 seconds and discard the flow-through into your waste stream conical tube.
- Add another 500 ul Buffer RPE to each column and microfuge for 2 minutes. Discard the flow-through and the collection tube.
- Transfer each column to a fresh collection tube and microfuge for 1 minute. This step is important since it removes any residual ethanol from the membrane, which could otherwise interfere with later assays.
- Trim the cap off two new 1.5 ml eppendorf tubes (save the caps!) and label the sides of the tubes with your team color, the date and a name for the sample. Transfer the columns into the trimmed eppendorf tubes and elute the RNA from the columns by adding 50 ul of RNase-free water to each. Microfuge for 1 minute then cap and store the samples on ice.
- Empty your waste stream conical tube into the chemical waste in the fume hood.
Part 4: Measure RNA concentration
- Measure the concentration of each RNA sample by adding 10 μl to 990 μl sterile water in a labeled eppendorf tube.
- The water does not have to be RNase-free since the RNA can be degraded and still give legitimate readings in the spectrophotometer.
- SAVE THE REST OF YOUR RNA. Give it to the teaching faculty to be frozen until next time.
- Take two disposable "UVettes" and fill them with 1 mL of water each. These are made of a polymer that does not itself absorb in the UV range, unlike the cuvettes that we used in Module 1. Thus, these cuvettes are much more expensive, and you should take extra care not to drop them!
- For each RNA sample, blank the spectrophotometer in wavelength scan mode with one of the water-filled cuvettes, then dump out the water and replace it with your RNA solution. After measuring in scan mode, use the arrows to select and write down the values at 260 and 280 nm.
- To determine the concentration of RNA in your sample, use the fact that 40 μg/ml of RNA will give a reading of 1 A260. Ideally, next time you will use 1-5 μg of RNA in each reverse transcription reaction. However, you also want both reactions to start with an equal amount of RNA template. Moreover, you cannot add more than 10 μL of RNA per reaction. If you can use 4 μg per reaction within these contraints, do so. Otherwise, figure out which one of your samples is limiting (has the least RNA), and scale the other sample amount that you add so they are equal. The table below may be helpful. You should turn it in as part of your homework assignment next time.
- Calculate the A260/A280 ratios of each RNA, which indicates the relative purity of your sample. This value should ideally be between ~1.9-2.1; however, the ratio is somewhat less reliable for samples tested in water rather than buffer.
| Sample | A260 value | RNA conc., diluted (μg/mL) |
RNA conc., original (μg/mL) |
Max RNA (# μg in 10 μL) | Volume RNA needed |
|---|---|---|---|---|---|
| control/scrambled siRNA sample | |||||
| mouse gene siRNA sample |
DONE!
For next time
- Post your raw luciferase data to the Talk page associated with today's lab. This will be important so others in the lab can consider your results in addition to their own and so other teams who used the same siRNA can consider the reproducibility of their work. Some multi-group comparative analysis should be included in your written report for this module. (You will get participation rather than homework points for this part.)
- Analyze your luciferase data as you did last time during lab, namely with a bar graph comparing the average Renilla/firefly ratios for each sample, and statistical information.
- This time, you can also perform a background subtraction of the data, before taking ratios. Background may come from light leaks into the luminometer, from autoluminescence of the reagents you used, or from other proteins in the cell lysates that might produce some light. As part of your homework, explicitly state which sample that you transfected last time can be used to take these considerations into account.
- Calculate confidence intervals for the averaged replicates (just your two, or better yet your two plus the two from the other lab section), and confidence levels for samples that you think are interesting to compare. Write a figure caption.
- Now write some initial accompanying Results text (~1-2 paragraphs) for your luciferase data. You should explicitly state what can you conclude about the efficacy of each siRNA that you personally used. You might also start thinking about information for the Discussion section, now considering the class data as a whole.
- Calculate the amount of RNA needed for your two reverse transcription reactions, if you have not already done so. Do hand this in as part of your homework.
Reagents list
- PLB
- Promega reagent (“Passive Lysis Buffer”)
- Buffer RTL/BME
- Qiagen reagent with BME
- Buffer RW1
- Qiagen reagent
- Buffer RPE
- Qiagen reagent with ethanol
